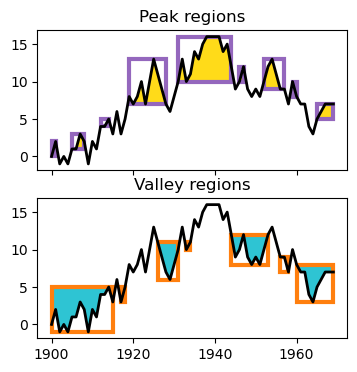
Data analysis of peak and valley regions
Project description
Peakoscope
Peakoscope is a python package for hierarchical analysis of peak and valley regions in numeric data.
- Peak regions can be nested, for example, when a large peak region contains smaller subpeak regions.
- A one-pass algorithm that finds all nested regions and orders them into a tree.
- Trees that can filtered, traversed, partitioned, pruned, grafted and serialized.
- Optional interfaces: matplotlib, pandas, polars, CLI and object-oriented.
Usage examples
Compute the tree of nested peak regions in a data set:
>>> import peakoscope
>>> data = [10, 30, 40, 30, 10, 50, 70, 70, 50, 80]
>>> peaktree = peakoscope.tree(data)
>>> print(peaktree)
0:10
├─5:10
│ ├─9:10
│ └─6:8
└─1:4
└─2:3
Print the tree minus its root:
>>> peaktree = peakoscope.tree(data, forest=True)
>>> print(peaktree - peaktree.roots())
5:10
├─9:10
└─6:8
1:4
└─2:3
Print the leaf nodes (which are local maxima in data) and corresponding slices of data:
>>> for leaf in peaktree.leaf_nodes():
... print(leaf, leaf.subarray(data))
...
9:10 [80]
6:8 [70, 70]
2:3 [40]
Documentation
Demo, tutorials and a glossary:
Authors
- Eivind Tøstesen, contact@tostesen.no
License
Copyright (C) 2021-2025 Eivind Tøstesen. This software is licensed under GPL-3.0-or-later
Citation
Citation can include one or more of:
-
Peakoscope v1.2.0
-
Github URL: https://github.com/eivindtostesen/hierarchical_peak_finding
-
PyPI URL: https://pypi.org/project/peakoscope/
-
The open-access article:
Tøstesen, E. A stitch in time: Efficient computation of genomic DNA melting bubbles. Algorithms for Molecular Biology, 3, 10 (2008). DOI: 10.1186/1748-7188-3-10
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file peakoscope-1.2.0.tar.gz.
File metadata
- Download URL: peakoscope-1.2.0.tar.gz
- Upload date:
- Size: 39.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9a00a44605b3e4dde73aea4639c7dd30829ba02cbf0c99bc72d1018ddaa3bd3c
|
|
| MD5 |
a4f5753573b1320ef646a52a1650a58c
|
|
| BLAKE2b-256 |
33b2a2b11f6e7e7abd41029026dc2c63673e94c4deb84fe73e2b9b9744e9fe08
|
File details
Details for the file peakoscope-1.2.0-py3-none-any.whl.
File metadata
- Download URL: peakoscope-1.2.0-py3-none-any.whl
- Upload date:
- Size: 40.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1f67a6049b7f3e24bc0504b2c8cd99db24eb7c2b70714d3517416e7b1a7e5ed1
|
|
| MD5 |
413a45c34855333254595d7650606d8b
|
|
| BLAKE2b-256 |
665645d92b3047545d5e9881030395541a95a0def7ce93998b8ca94edb174337
|