Universal ortholog based Phylogenomic toolkit.
Project description
phyca: phylogeny and collinearity aware assembly evaluation toolkit.
phyca is built around Compleasm utilizing the NCBI Genome database. For a query assembly, phyca improves the precision of BUSCO/Compleasm annotations by up to 7%, makes syntenic comparisons to public reference genomes and rapidly places the assembly on a broad, precomputed phylogeny.
Rationale
BUSCOs are the most conserved genes. Gene duplication and deletion in parallel branches can confound evolutionary genomic analyses. In our article, we explored the extent of BUSCO gene misannotations in major eukaryotic lineages. A misannotated gene is a gene that gets annotated by an annotation software when the original gene copy is lost in a lineage. From our survey of 20,000 plant, fungi and animal species genomes, we found that ~10% of BUSCO genes have significantly greater propensity of being misannotated than others. phyca filters out the misannotation-prone genes and outputs annotaitons and stats for curated BUSCO genes or CUSCOs.
Our original article was based on ODB10 orthologs. Phyca has now been updated to run on ODB12. Please view the updated ortholog stats visualized here.
Installation
pip install phyca
phyca is distributed through PyPI and github. A working installation of Compleasm (including SEPP and pplacer) is necessary to avail all functionality. I recommend creating a conda environment to install Compleasm first and installing phyca in that environment, e.g.,
# create environment
conda create -n phyca python=3.9.25
# install compleasm
conda install bioconda::compleasm=0.2.7
# install phyca
pip install phyca
Note that as of 02/03/2025, there is a known issue with pplacer and SEPP on Debian-based systems. A working solution is provided [here](https://github.com/smirarab/sepp/issues/140).
phyca has the following nonexhaustive dependency structure.
Python (tested with 3.9.25)
↓
│───numpy (tested with 2.0.1)
│───pandas (tested with 2.3.3)
│───matplotlib (tested with 3.9.4)
│───seaborn (tested with 0.13.2)
│───SciPy (tested with 1.13.1)
│───BioNick (tested with 0.0.8)
└───Compleasm (tested with 0.2.7)
│─── hmmer (tested with 3.1b2)
│─── miniprot (tested with 0.13-r248)
│ └─── libgcc (tested with 14.2.0 under conda)
└─── SEPP (tested with 4.4.0)
└─── pplacer and guppy (v1.1.alpha19-0-g807f6f3)
Usage
phyca supports 10 BUSCO lineages: viridiplantae, liliopsida, eudicots, chlorophyta, fungi, ascomycota, basidiomycota, metazoa, arthropoda and vertebrata.
A simple run on a query assembly, would be:
phyca -a <assembly_file> -l <lineage>
The Compleasm output folder can also be used as input if compleasm output was previously generated:
phyca -c <compleasm_direcoty> -l <lineage>
The above run will output BUSCO, CUSCO (Curated USCOs with higher precision) and MUSCO (remaining USCOs) statistics and graphs. It will compare the query to chromosome level genome assemblies from NCBI genome and output a table with a measure of synteny against each genome. It will output a Neighbor-Joining tree based on BUSCO synteny. Finally, it will place the assembly on a large precomputed phylogeny for the lineage and graph the observed decay in BUSCO synteny against inferred phylogenetic distance.
Assembly syntenic comparisons
phyca allows syntenic comparisons between assemblies with compleasm annotations or any set of gene annotations formatted in the same way.
to compute the syntenic distance between two assemblies with the -s flag.
phyca -l <lineage> -s -a <assembly1> -r <assembly2>
The same comparison can be done by pointing to the compleasm output directoreis, if already available.
phyca -l <lineage> -s -c <assembly1_compdir> -m <assembly2_compdir>
Comparisons are done in the following way, adjust for variable query contiguity, and will produce the best results when one of the assemblies is highly contiguous and accurate:
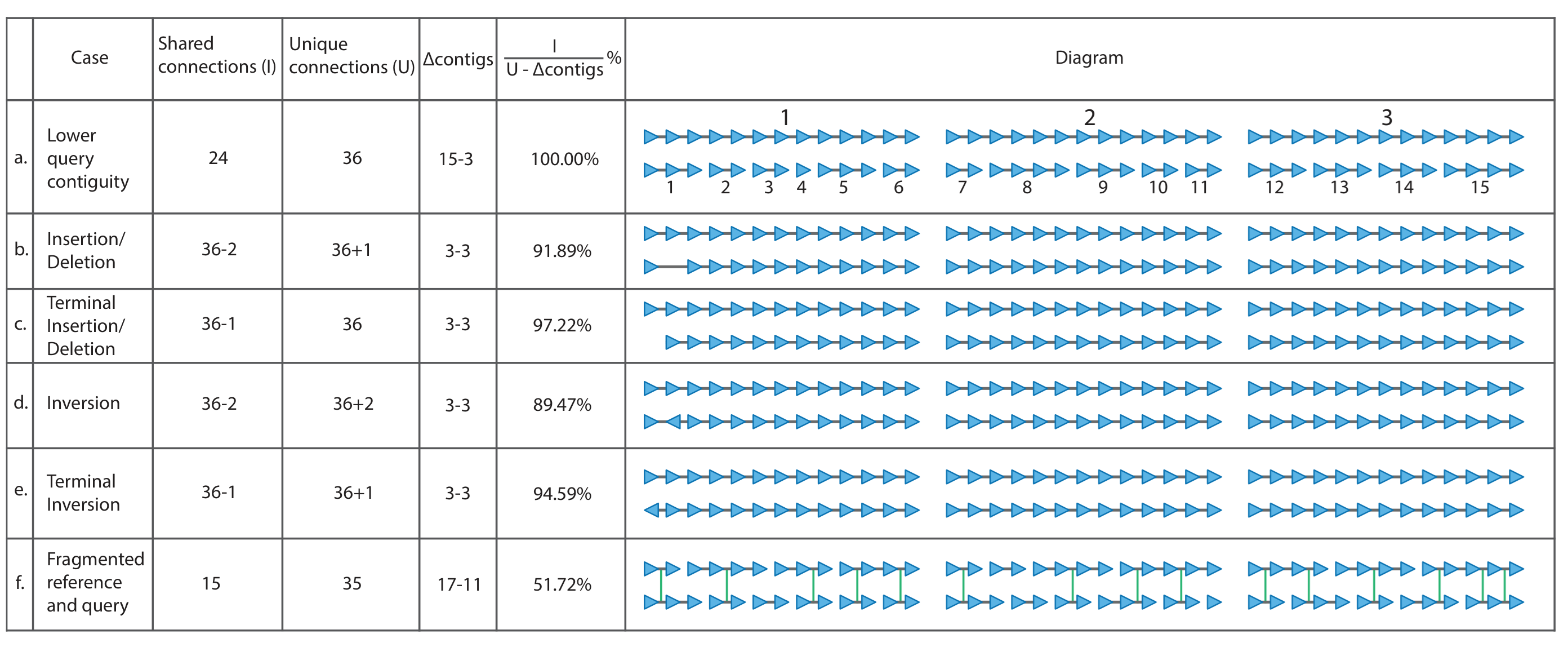
UniPhyDB
The bulk data used by phyca is hosted by AGI's AVA cluster. All alignments, precomputed trees, annotations, metadata and more information is available at phyca.org.
Example Output
USCO graph:
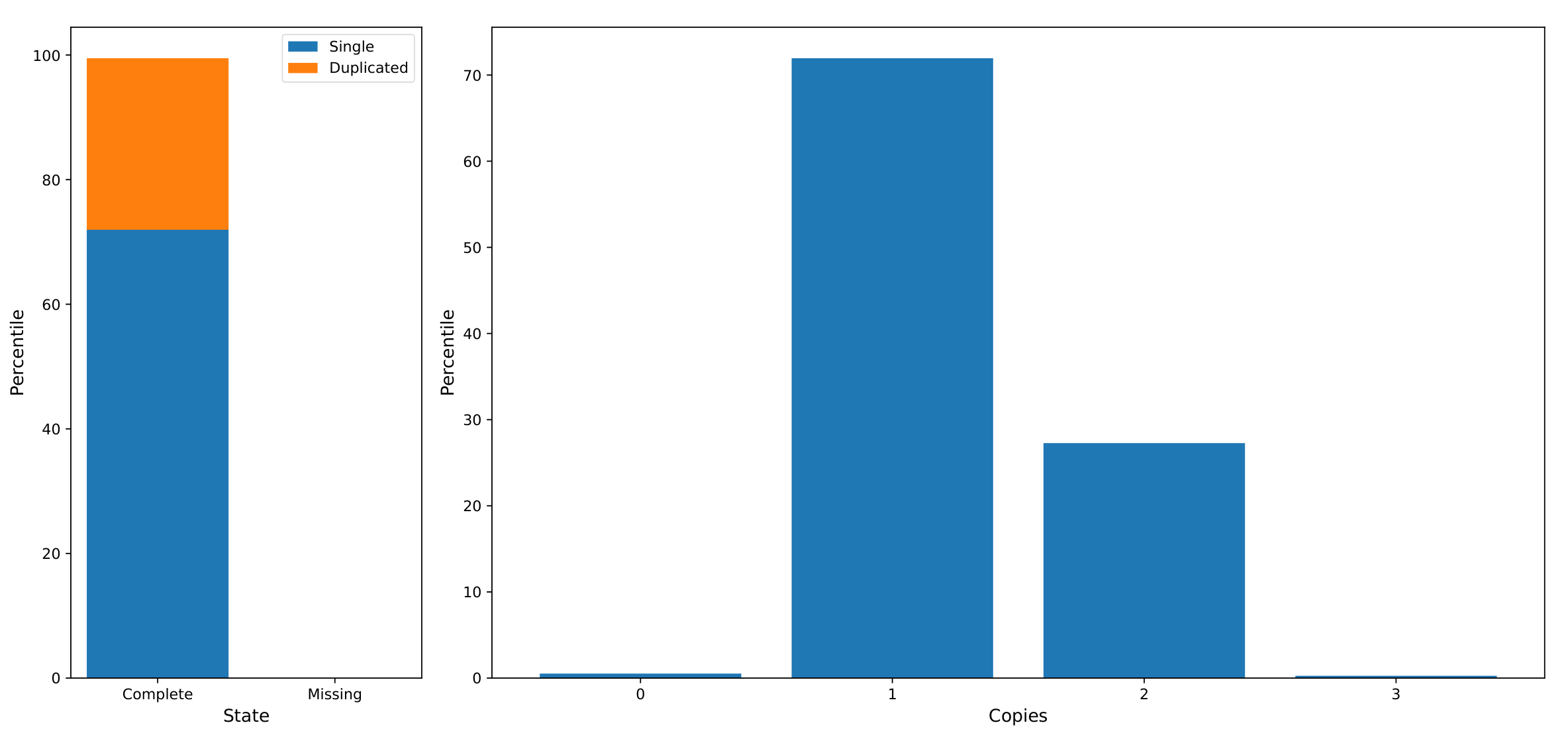
Synteny decay plot:
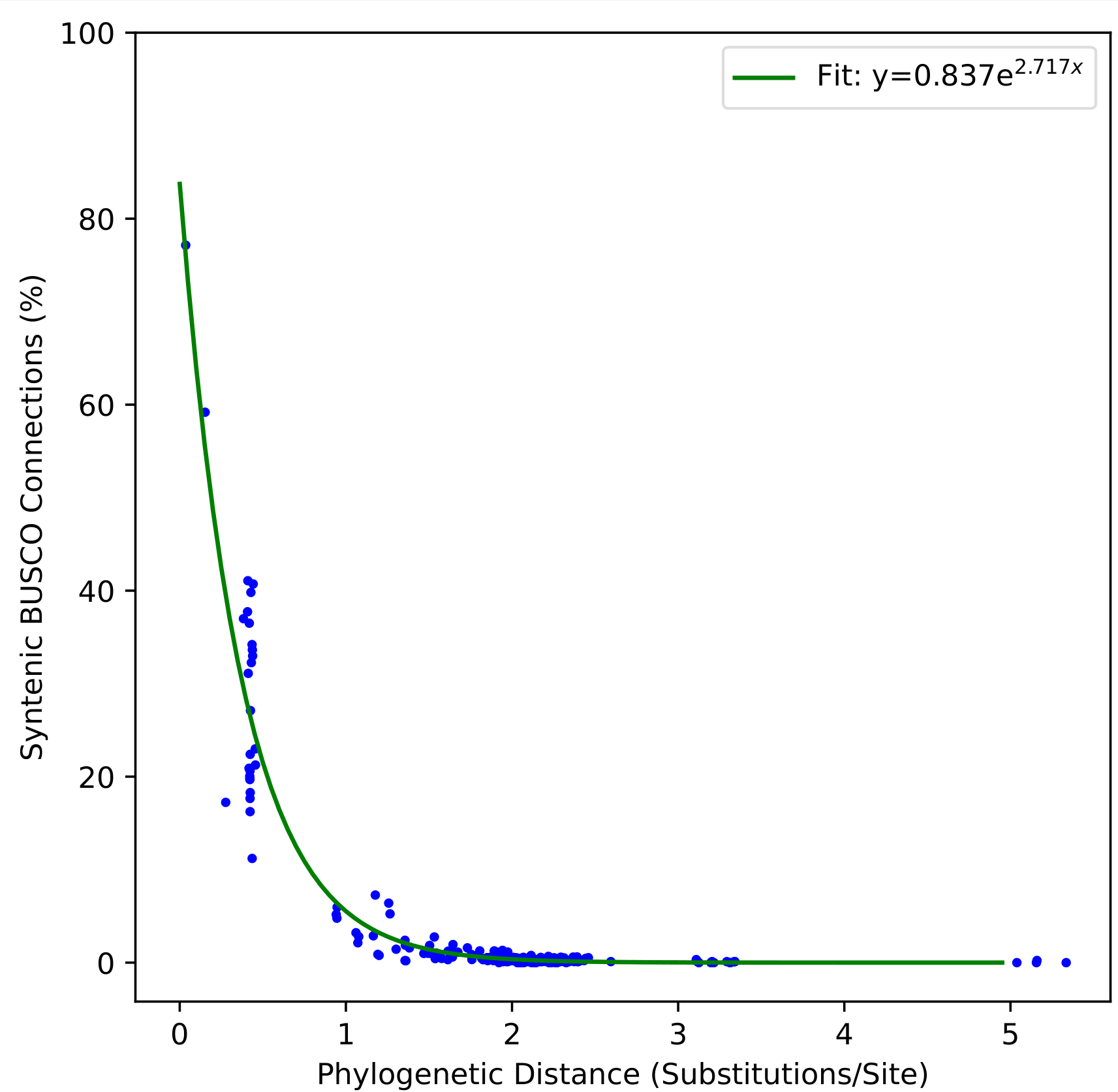
Placement tree snippet:
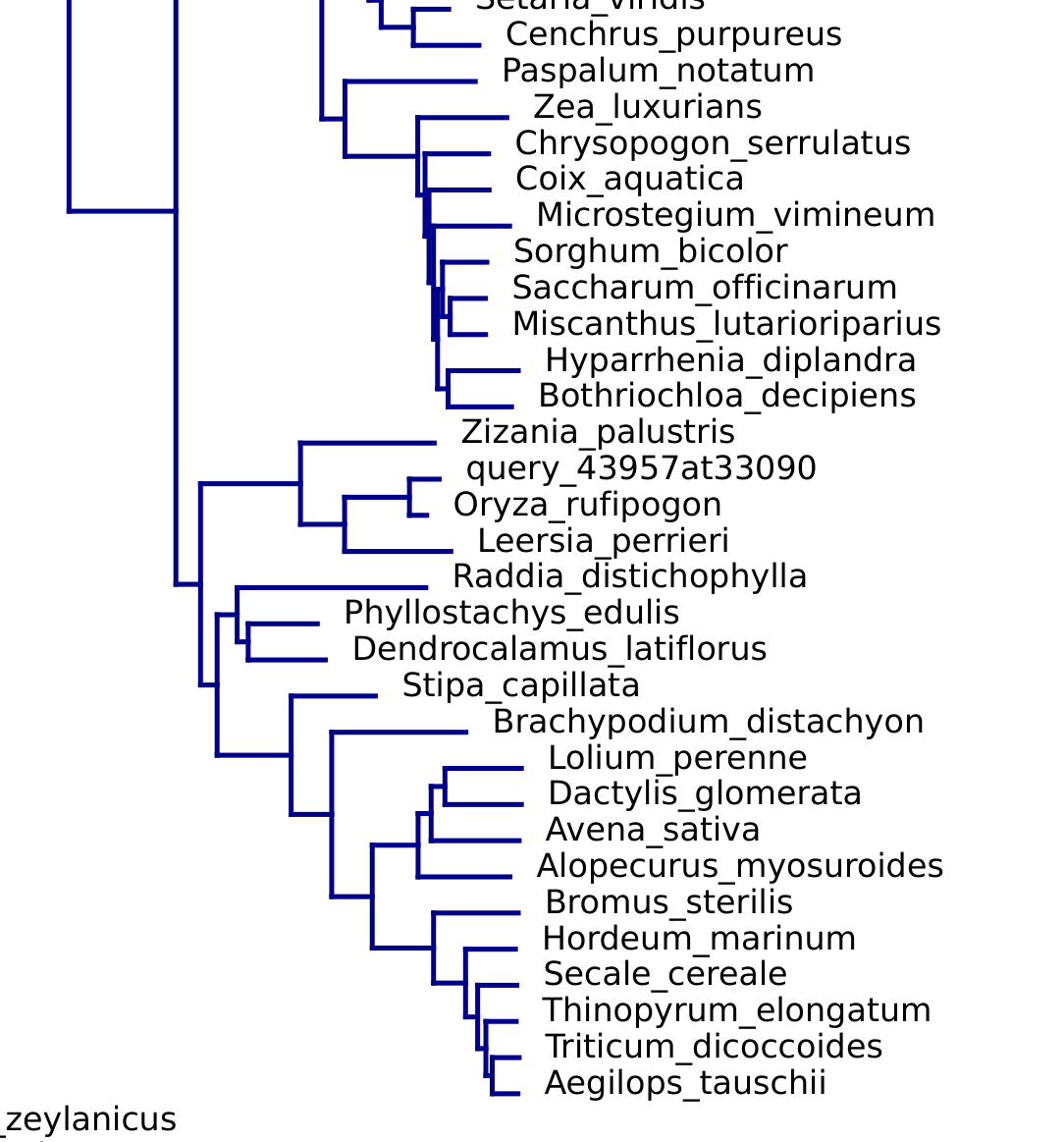
Citation
Alam, M.N.U., Román-Palacios, C., Copetti, D. et al. Universal orthologs infer deep phylogenies and improve genome quality assessments. BMC Biol 23, 224 (2025). https://doi.org/10.1186/s12915-025-02328-2
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file phyca-0.0.3.tar.gz.
File metadata
- Download URL: phyca-0.0.3.tar.gz
- Upload date:
- Size: 131.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6f16f374114ac81a1d72d729a4ca433c8598fe1d3a3dd2593785b36840dfbd93
|
|
| MD5 |
79284655a33f8c008fce6c8b30ff6a82
|
|
| BLAKE2b-256 |
405f16051a8e7d687bd52dc5b01afcafa45f1aff1ffc4eeb52475a74f54f0515
|
File details
Details for the file phyca-0.0.3-py3-none-any.whl.
File metadata
- Download URL: phyca-0.0.3-py3-none-any.whl
- Upload date:
- Size: 184.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
27b0671d68220620467f8b63cf7b22ccfff27bfbcd275a5ccc9bd009050ea0ea
|
|
| MD5 |
98c99d52be02e0c2d4e8bef062734bd5
|
|
| BLAKE2b-256 |
a7a6b30b42bf7f078fd96bcb5278b17b386d037d5121635160bfed54b2fba4e9
|











