PKSmart: An Open-Source Computational Model to Predict in vivo Pharmacokinetics of Small Molecules
Project description
PKSmart
Drug exposure is a critical determinant of drug safety and efficacy, defined through human phar-macokinetics (PK) parameters such as steady-state volume of distribution (VDss), total body clear-ance (CL), half-life (t½), fraction unbound in plasma (fu), and mean residence time (MRT), which influence a drug's blood concentration profile. In this study, we modelled these human PK parame-ters for 1,283 unique compounds using molecular structural fingerprints, physicochemical proper-ties, and predicted animal PK data. We first predicted animal PK parameters (VDss, CL, fu) for rats, dogs, and monkeys for 371 compounds using molecular structural fingerprints and physico-chemical properties. Next, we employed Morgan fingerprints, Mordred descriptors, and predicted animal PK parameters in a hyperparameter-optimized Random Forest algorithm to predict human PK parameters. Repeated nested cross-validation demonstrated predictive performance for VDss (R² = 0.53 and Geometric Mean Fold Error, GMFE= 2.13), CL (R² = 0.31, GMFE = 2.46), fu (R² = 0.63, GMFE = 2.71), MRT (R² = 0.27, GMFE = 2.50), and t½ (R² = 0.31, GMFE = 2.46). External validation on 315 compounds showed strong performance for VDss (R² = 0.39, GMFE = 2.46) and CL (R² = 0.46, GMFE = 1.95). Comparison with AstraZeneca PK models revealed similar predic-tive performance, with Pearson correlations ranging from 0.46 to 0.73 for animal PK parameters of VDss, CL, and fu. PKSmart models further demonstrated predictive performance for VDss (R² = 0.33, RMSE = 0.58), comparable to AstraZeneca's human VDss models (R² = 0.30, RMSE = 0.60) on an external test set of 51 compounds. To our knowledge, this is the first publicly available set of PK models with performance on par with industry standard models. These models enable early in-tegration of predicted PK properties into workflows such as Design-Make-Test-Analyze (DMTA) cycles using only chemical structures as input. We also developed a web-hosted application, PKSmart (https://broad.io/PKSmart), accessible via browser, with all associated code available for local use.
Install using PyPI
pip install pksmart
Install from source
- Clone this repo
git clone https://github.com/Manas02/pksmart-pip
- Install the
PKSmartPackage
poetry install
poetry build
Usage
Help
Simply run pksmart or pksmart -h or pksmart --help to get helper.
Running PKSmart as CLI
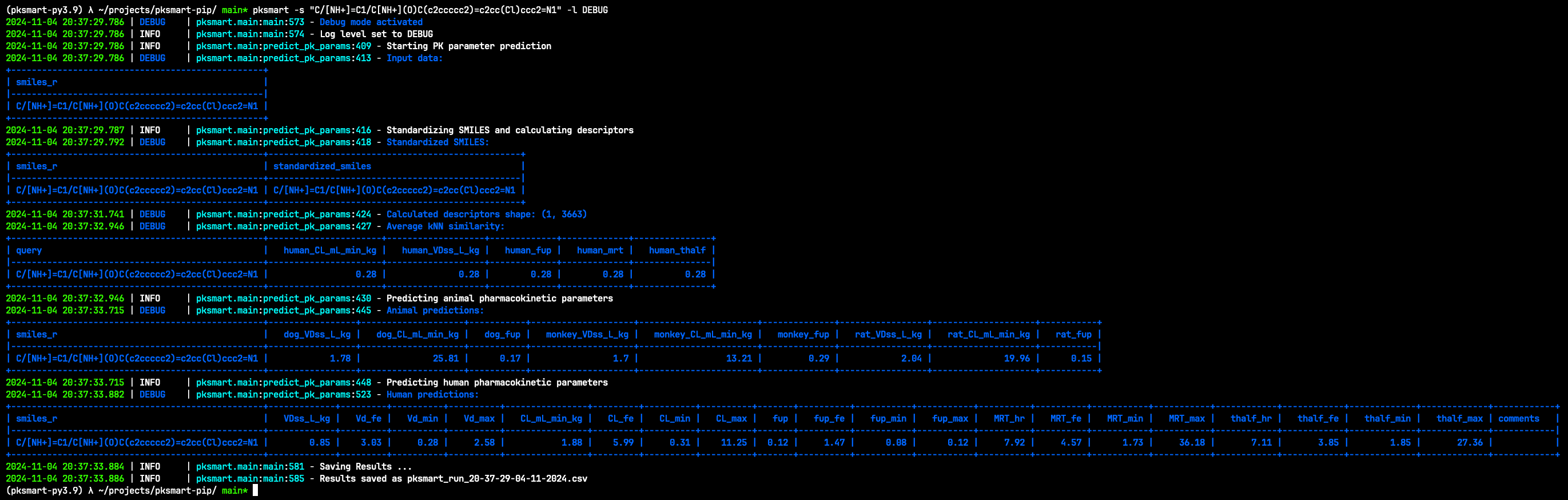
Run pksmart -s or pksmart --smi or pksmart --smiles to run inference on a single SMILES string.
Alternatively, Run pksmart -f or pksmart --file to run inference using a file containing newline separated SMILES strings.
Running PKSmart as Library
import pksmart
if __name__ == "__main__":
smiles = "CCCCCO"
out = pksmart.predict_pk_params(smiles)
print(out)
Cite
If you use PKSmart in your work, please cite:
PKSmart: An Open-Source Computational Model to Predict in vivo Pharmacokinetics of Small Molecules Srijit Seal, Maria-Anna Trapotsi, Vigneshwari Subramanian, Ola Spjuth, Nigel Greene, Andreas Bender bioRxiv 2024.02.02.578658; doi: https://doi.org/10.1101/2024.02.02.578658
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pksmart-3.0.1.tar.gz.
File metadata
- Download URL: pksmart-3.0.1.tar.gz
- Upload date:
- Size: 42.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.2 CPython/3.12.8 Darwin/24.6.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
79b4aba558a830b64ea1280fd6caa843107db65e942da1659e08e024122f87ae
|
|
| MD5 |
45db5ea44b8ee4f119073a0d78339572
|
|
| BLAKE2b-256 |
64293851ed792c69eec6c98d5bc083e391c4dd667e958ad6b141bf19a776e768
|
File details
Details for the file pksmart-3.0.1-py3-none-any.whl.
File metadata
- Download URL: pksmart-3.0.1-py3-none-any.whl
- Upload date:
- Size: 43.0 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.2 CPython/3.12.8 Darwin/24.6.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0226969196cade1954b8355b600132d32a709fc7f12ef71a5db102f52de84743
|
|
| MD5 |
984b26165564b1ae5500344ccf1a694f
|
|
| BLAKE2b-256 |
603cf3b4c8c8848875f69f72f793d37b9f465ea19a6b4ac00d95761393593bbc
|