PLINDER: The Protein-Ligand INteraction Dataset and Evaluation Resource
Project description
The Protein Ligand INteractions Dataset and Evaluation Resource
📚 About
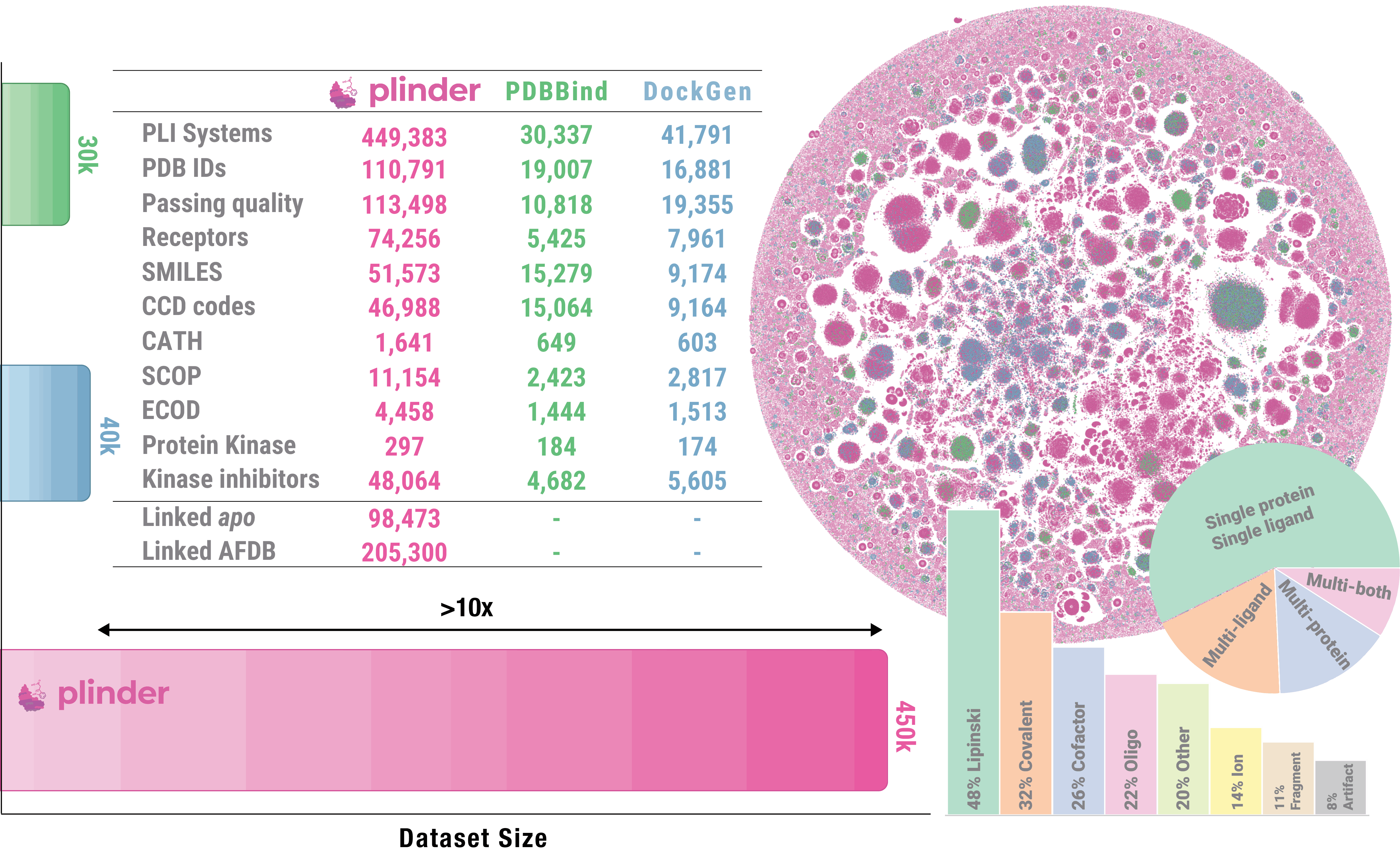
PLINDER, short for protein ligand interactions dataset and evaluation resource, is a comprehensive, annotated, high quality dataset and resource for training and evaluation of protein-ligand docking algorithms:
- > 400k PLI systems across > 11k SCOP domains and > 50k unique small molecules
- 750+ annotations for each system, including protein and ligand properties, quality, matched molecular series and more
- Automated curation pipeline to keep up with the PDB
- 14 PLI metrics and over 20 billion similarity scores
- Unbound (apo) and predicted Alphafold2 structures linked to holo systems
- train-val-test splits and ability to tune splitting based on the learning task
- Robust evaluation harness to simplify and standard performance comparison between models.
The PLINDER project is a community effort, launched by the University of Basel, SIB Swiss Institute of Bioinformatics, Proxima (formerly VantAI), NVIDIA, MIT CSAIL, and will be regularly updated.
To accelerate community adoption, PLINDER will be used as the field’s new Protein-Ligand interaction dataset standard as part of an exciting competition at the upcoming 2024 Machine Learning in Structural Biology (MLSB) Workshop at NeurIPS, one of the field's premiere academic gatherings. More details about the competition and other helpful practical tips can be found at our recent workshop repo: Moving Beyond Memorization.
👋 Join the P(L)INDER user group Discord Server!
🔢 Plinder versions
We version the plinder dataset with two controls:
PLINDER_RELEASE: the month stamp of the last RCSB syncPLINDER_ITERATION: value that enables iterative development within a release
We version the plinder application using an automated semantic
versioning scheme based on the git commit history.
The plinder.data package is responsible for generating a dataset
release and the plinder.core package makes it easy to interact
with the dataset.
🐛🐛🐛 Known bugs:
- Source dataset contains incorrect
entry_release_datedates, please, usequery_indexto get correct dates patched. - Complexes containing nucleic acid receptors may not be saved corectly.
ligand_binding_affinityqueries have been disabled due to a bug found parsing BindingDB
Changelog:
-
2024-06/v2 (Current):
- New systems added based on the 2024-06 RCSB sync
- Updated system definition to be more stable and depend only on ligand distance rather than PLIP
- Added annotations for crystal contacts
- Improved ligand handling and saving to fix some bond order issues
- Improved covalency detection and annotation to reference each bond explicitly
- Added linked apo/pred structures to v2/links and v2/linked_structures
Added binding affinity annotations from BindingDB(see known bugs!)- Added statistics requirement and other changes in the split to enrich test set diversity
-
2024-04/v1: Version described in the preprint, with updated redundancy removal by protein pocket and ligand similarity.
-
2024-04/v0: Version used to re-train DiffDock in the paper, with redundancy removal based on <pdbid>_<ligand ccd codes>
🏅 Gold standard benchmark sets
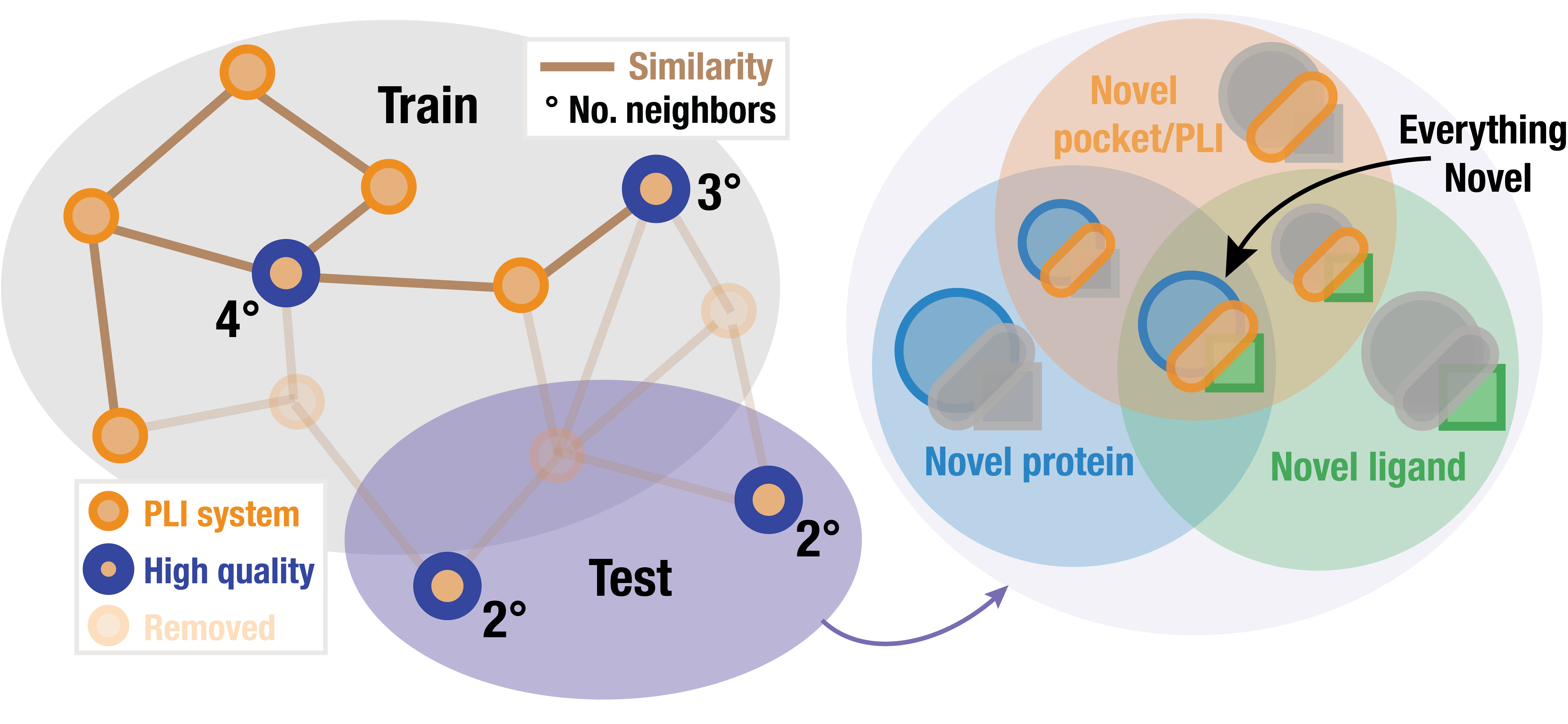
As part of PLINDER resource we provide train, validation and test splits that are
curated to minimize the information leakage based on protein-ligand interaction
similarity.
In addition, we have prioritized the systems that has a linked experimental apo
structure or matched molecular series to support realistic inference scenarios for hit
discovery and optimization.
Finally, a particular care is taken for test set that is further prioritized to contain
high quality structures to provide unambiguous ground-truths for performance
benchmarking.
Moreover, as we enticipate this resource to be used for benchmarking a wide range of methods, including those simultaneously predicting protein structure (aka. co-folding) or those generating novel ligand structures, we further stratified test (by novel ligand, pocket, protein or all) to cover a wide range of tasks.
👨💻 Getting Started
The PLINDER dataset is provided in two ways:
- You can either use the files from the dataset directly using your preferred tooling by downloading the data from the public bucket,
- or you can utilize the dedicated
plinderPython package for interfacing the data.
Downloading the dataset
The dataset can be downloaded from the bucket with gsutil.
$ export PLINDER_RELEASE=2024-06 # Current release
$ export PLINDER_ITERATION=v2 # Current iteration
$ mkdir -p ~/.local/share/plinder/${PLINDER_RELEASE}/${PLINDER_ITERATION}/
$ gsutil -m cp -r "gs://plinder/${PLINDER_RELEASE}/${PLINDER_ITERATION}/*" ~/.local/share/plinder/${PLINDER_RELEASE}/${PLINDER_ITERATION}/
For details on the sub-directories, see Documentation.
Installing the Python package
plinder is available on PyPI.
pip install plinder
License
Data curated by PLINDER are made available under the Apache License 2.0. All data curated by BindingDB staff are provided under the Creative Commons Attribution 4.0 License. Data imported from ChEMBL are provided under their Creative Commons Attribution-Share Alike 4.0 Unported License.
📝 Documentation
A more detailed description is available on the documentation website.
📃 Citation
Durairaj, Janani, Yusuf Adeshina, Zhonglin Cao, Xuejin Zhang, Vladas Oleinikovas, Thomas Duignan, Zachary McClure, Xavier Robin, Gabriel Studer, Daniel Kovtun, Emanuele Rossi, Guoqing Zhou, Srimukh Prasad Veccham, Clemens Isert, Yuxing Peng, Prabindh Sundareson, Mehmet Akdel, Gabriele Corso, Hannes Stärk, Gerardo Tauriello, Zachary Wayne Carpenter, Michael M. Bronstein, Emine Kucukbenli, Torsten Schwede, Luca Naef. 2024. “PLINDER: The Protein-Ligand Interactions Dataset and Evaluation Resource.” bioRxiv ICML'24 ML4LMS
Please see the citation file for details.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file plinder-0.2.26.tar.gz.
File metadata
- Download URL: plinder-0.2.26.tar.gz
- Upload date:
- Size: 28.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4f5bc456ca3fd01eb80fee22f8f891e3d9ea060659fbdfe6f93363cabda833f0
|
|
| MD5 |
225758f21f474ffd2f2e539bd4192dd3
|
|
| BLAKE2b-256 |
4efedc17a16e5c71e0870a23a236866ee2880d6e04ce1bc8a568e0212a4cc536
|
Provenance
The following attestation bundles were made for plinder-0.2.26.tar.gz:
Publisher:
main.yaml on plinder-org/plinder
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
plinder-0.2.26.tar.gz -
Subject digest:
4f5bc456ca3fd01eb80fee22f8f891e3d9ea060659fbdfe6f93363cabda833f0 - Sigstore transparency entry: 969124932
- Sigstore integration time:
-
Permalink:
plinder-org/plinder@85b3f1cb1763530a6cfd934f4263a1777c41afa4 -
Branch / Tag:
refs/heads/main - Owner: https://github.com/plinder-org
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
main.yaml@85b3f1cb1763530a6cfd934f4263a1777c41afa4 -
Trigger Event:
push
-
Statement type:
File details
Details for the file plinder-0.2.26-py3-none-any.whl.
File metadata
- Download URL: plinder-0.2.26-py3-none-any.whl
- Upload date:
- Size: 5.0 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
044f183cdacb4a6200f2dd69db6bb445b138c81994fc65e2c5b590722a16c7a8
|
|
| MD5 |
65c42dbdc5c50a0a3de63c62856a7ac2
|
|
| BLAKE2b-256 |
3ba11930c69c19a976cc8ac2aad61ef6703abc3f3eb77124ea4f3bc0fc1e38f0
|
Provenance
The following attestation bundles were made for plinder-0.2.26-py3-none-any.whl:
Publisher:
main.yaml on plinder-org/plinder
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
plinder-0.2.26-py3-none-any.whl -
Subject digest:
044f183cdacb4a6200f2dd69db6bb445b138c81994fc65e2c5b590722a16c7a8 - Sigstore transparency entry: 969124935
- Sigstore integration time:
-
Permalink:
plinder-org/plinder@85b3f1cb1763530a6cfd934f4263a1777c41afa4 -
Branch / Tag:
refs/heads/main - Owner: https://github.com/plinder-org
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
main.yaml@85b3f1cb1763530a6cfd934f4263a1777c41afa4 -
Trigger Event:
push
-
Statement type: