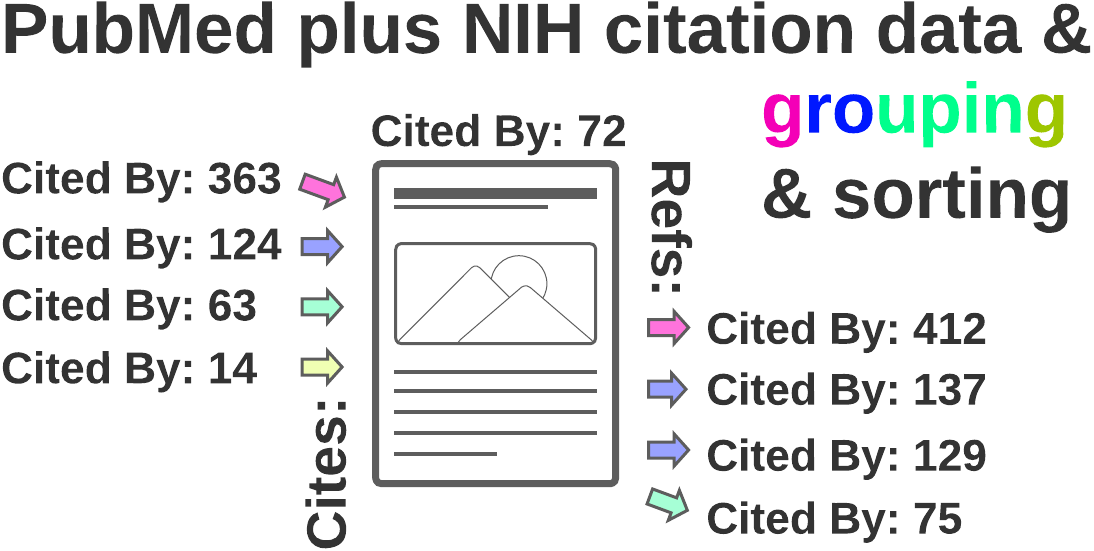
Turbocharge a PubMed literature search using citation data from the NIH
Project description
PubMed ID (PMID) Cite
Turbocharge a PubMed literature search with the command, icite, rather than clicking and clicking and clicking on Google Scholar "Cited by N" links.
This open-source project is part of a peer-reviewed commentary that was invited by the editors of Research Synthesis Methods. Please Cite and star on GitHub if you use pmidcite in your research or literature search.
Contact: dvklopfenstein@protonmail.com
PubMed and NIH Citation data
PubMed contains peer-reviewed research papers in biomedicine, biochemistry, chemistry, behavioral science, and other life sciences.
Freely available Citation data
from the National Institutes of Health (NIH)
is freshly downloaded
when icite is run and
provides much more than just citation counts, including:
- Specifies citing clinical papers
- Performance of the paper among its peers
- Presence of MeSH terms for the human, animal, and molecular/cellular categories
Table of Contents
- Installation
- Quickstart
- How to cite
- Google Scholar vs. PubMed
- Jupyter notebooks
- Contributing
- References
Quickstart
- Quickstart on the command line
- 1) Download citation counts and data for a research paper
- 2) Forward citation search: following a paper's Cited by links or Forward snowballing
- 3) Backward citation search: following the links to a paper's references or Backward snowballing
- 4) Summarize a group of citations
- 5) Download citations for all papers returned from a PubMed search
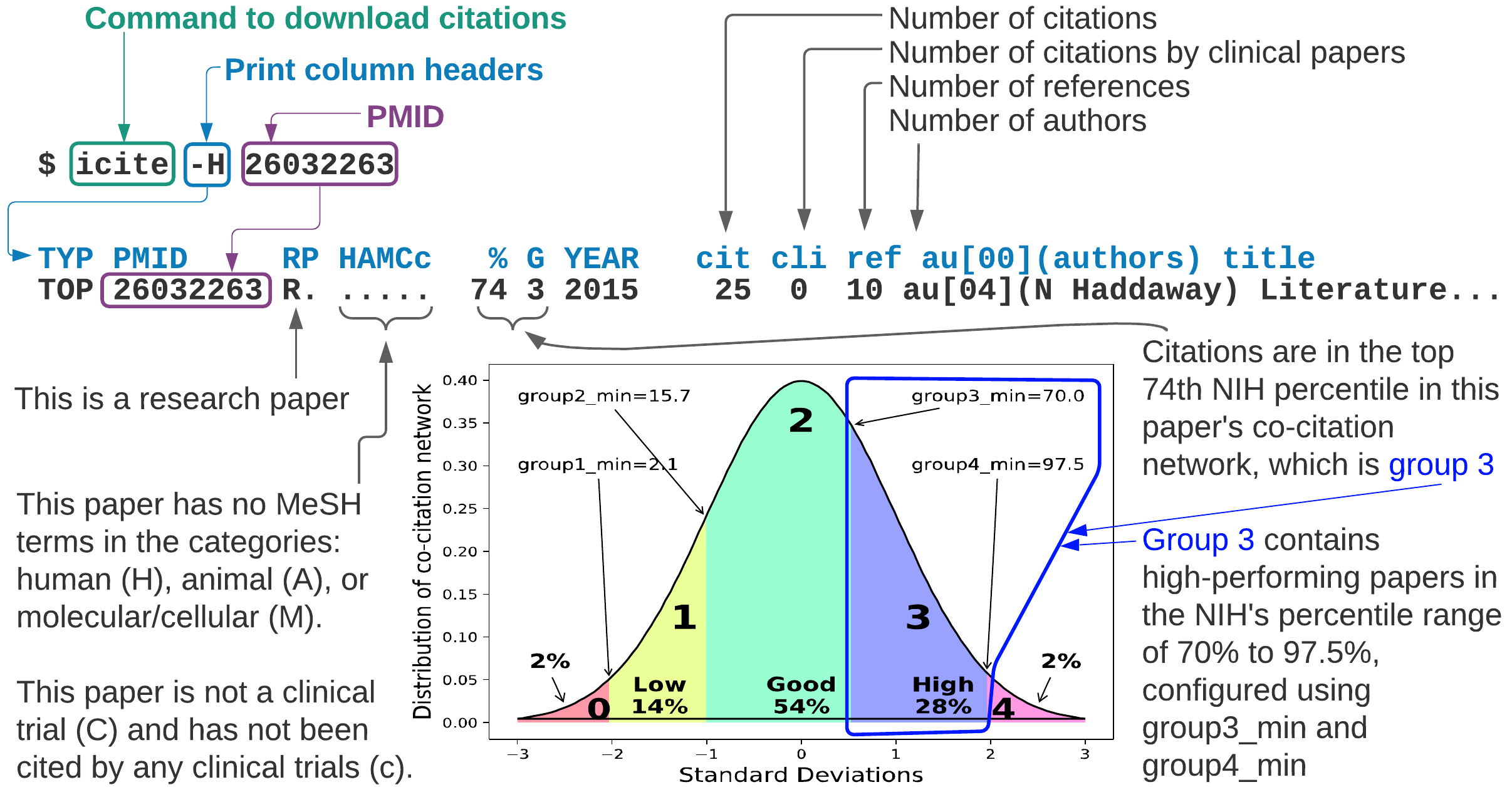
1) Download citation counts and data for a research paper
$ icite -H 26032263
- This paper (PMID 26032263) has
25citations,10references, and4authors. - This paper is performing well (
74th percentile in column%) compared to its peers.
NIH percentile
This paper is performing well (74th percentile) compared to its peers (column %).
The NIH percentile grouping (column G) helps to
highlight the better performing papers in groups 2, 3, and 4 by
sorting the citing papers by group first, then publication year.
The sort places the lower performing papers in groups 0 or 1 at the back.
New papers appear at the beginning of a sorted list, no matter how many citations they have to better facilitate researchers in finding the latest discoveries.
The grouping of papers by NIH percentile grouping is a novel feature created by dvklopfenstein for this project.
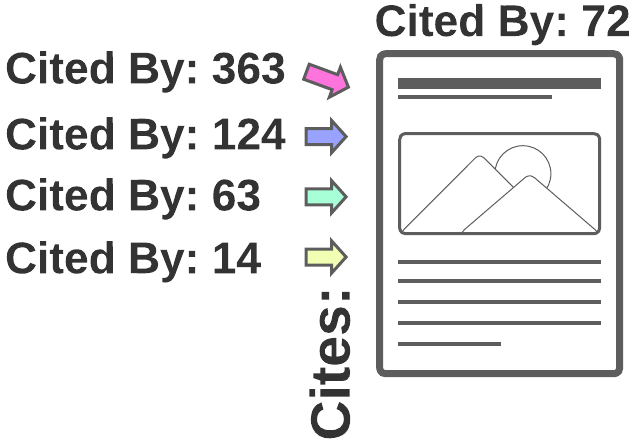
2) Forward citation search
Also known as following a paper's Cited by links or Forward snowballing
icite -H; icite 26032263 --load_citations | sort -k6 -r
or
icite -H; icite 26032263 -c | sort -k6 -r
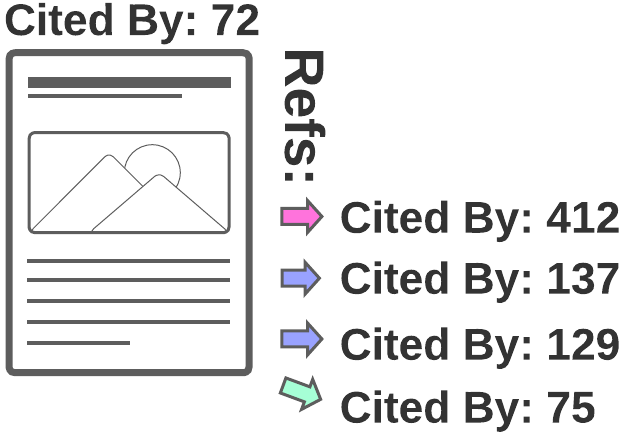
3) Backward citation search
Also known as following links to a paper's references or Backward snowballing
$ icite -H; icite 26032263 --load_references | sort -k6 -r
or
$ icite -H; icite 26032263 -r | sort -k6 -r
4) Summarize a group of citations
Create a file containing numerous PMIDs annotated with icite info
$ icite 30022098 -c -o goatools_cites.txt
WROTE: goatools_cites.txt
Count the number of lines in the file
$ wc -l goatools_cites.txt
468 goatools_cites.txt
Summarize the papers in "goatools_cites.txt"
$ sumpaps goatools_cites.txt
i=026.9% 4=003.0% 3=018.9% 2=028.8% 1=015.9% 0=006.5% 6 years:2018-2024 465 papers goatools_cites.txt
- The output is on one line so many files containing sets of PMIDs may be compared
- The groups are from newest(
i) to top-performing(4), great(3), very good(2), and overlooked(1and0)
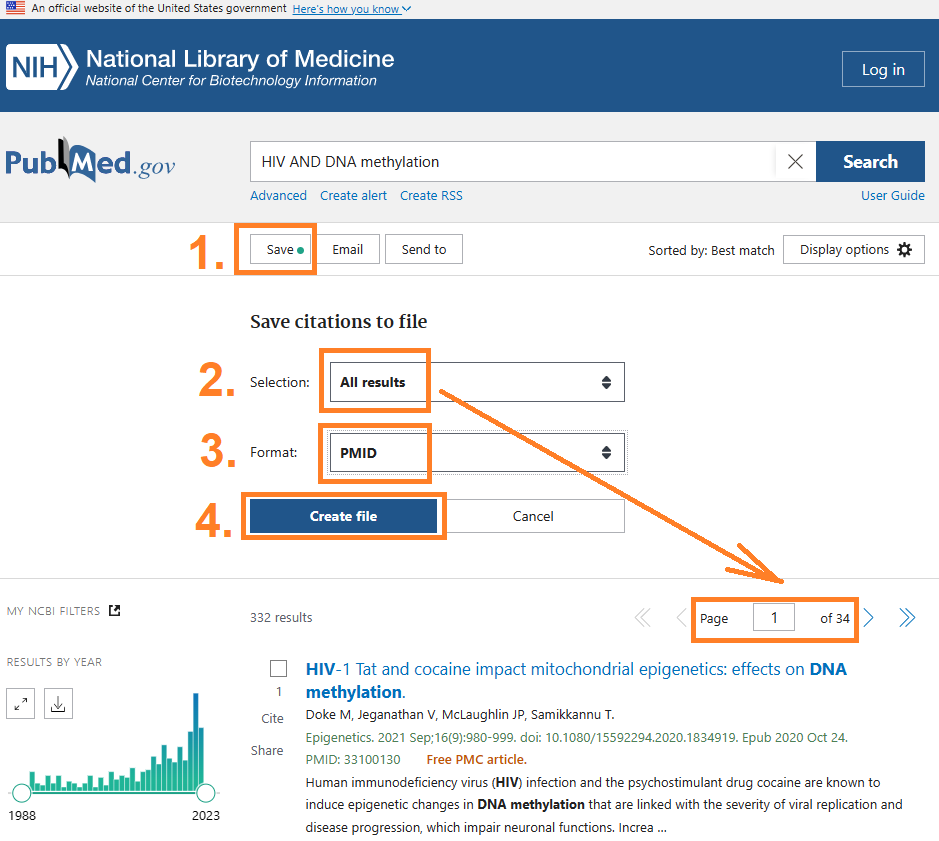
5) Download citations for all papers returned from a PubMed search
- Do a search in PubMed
- Save all results into a file containing all PMIDs found by the search
- Download the list of PMIDs
- Run icite to analyze all the PMIDs
1. Do a search in PubMed
2. Save all results into a list of PMIDs
3. Download the list of PMIDs
4. Run icite to analyze all the PMIDs
$ icite -i pmid-HIVANDDNAm-set.txt -o pmid-HIVANDDNAm-icite.txt
$ grep TOP pmid-HIVANDDNAm-icite.txt | sort -k6
Command Line Interface (CLI)
A Command-Line Interface (CLI) can be preferable to a Graphical User Interface (GUI) because:
- processing can be automated from a script
- time-consuming mouse clicking is reduced
- more data can be seen at once on a text screen than in a browser, giving the researcher a better overall impression of the full set of information [1]
Researchers who use Linux or Mac already work from the command line. Researchers who use Windows can get that Linux-like command line feeling while still running native Windows programs by downloading Cygwin from https://www.cygwin.com/ [1].
PubMed vs Google Scholar
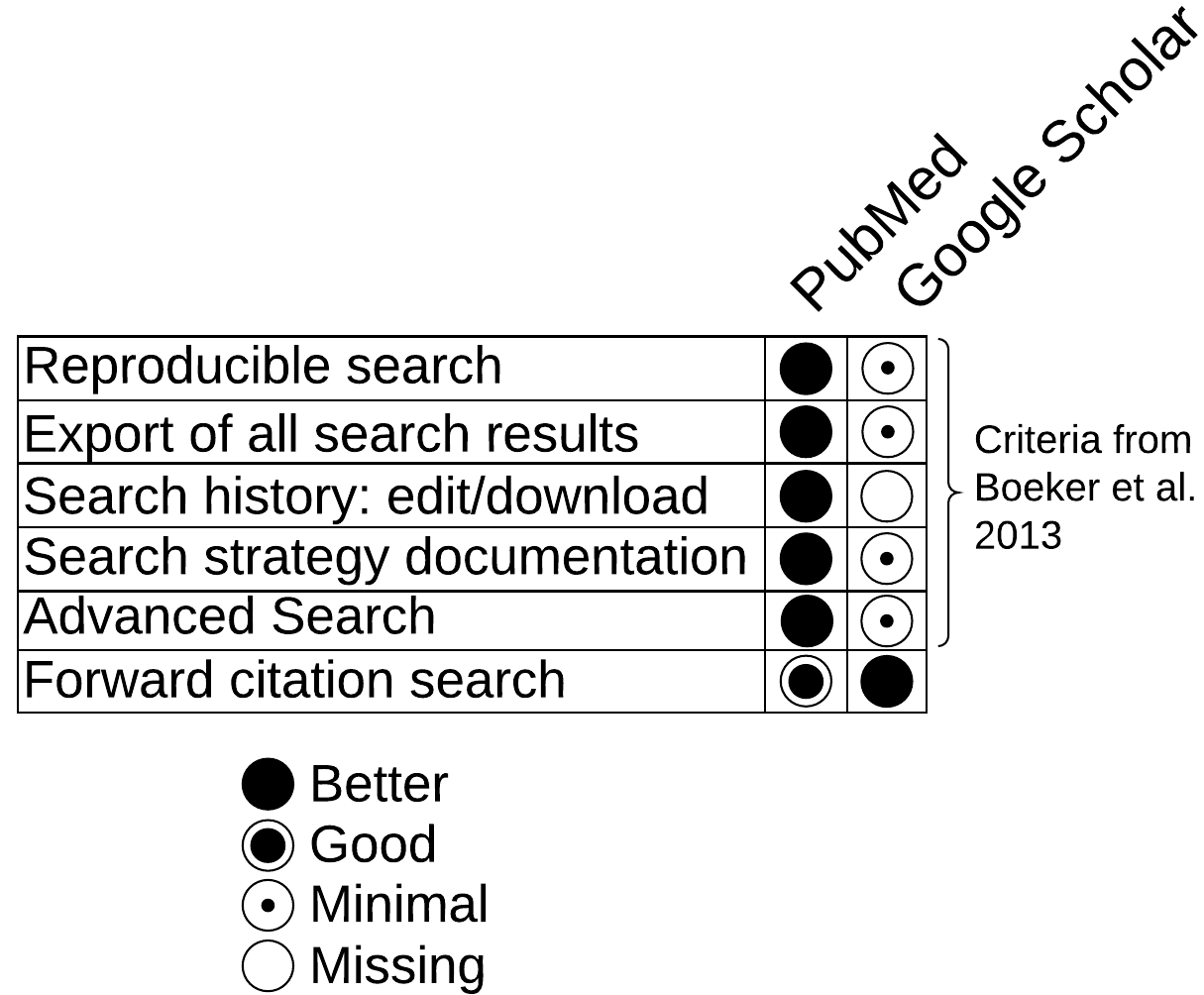
In 2013, Boeker et al. [6] recommended that a scientific search interface contain five integrated search criteria. PubMed implements all five, while Google did not in 2013 or today.
Google's highly popular implementation of the forward citation search through their ubiquitous "Cited by N" links is a "Better" experience than the PubMed's "forward citation search" implementation.
But if your research is in the health sciences and you are amenable to working from the command line, you can use PubMed in your browser plus citation data downloaded from the NIH using the command-line using pmidcite. The NIH's citation data includes a paper's ranking among its co-citation network.
What is in PubMed? Take a quick tour
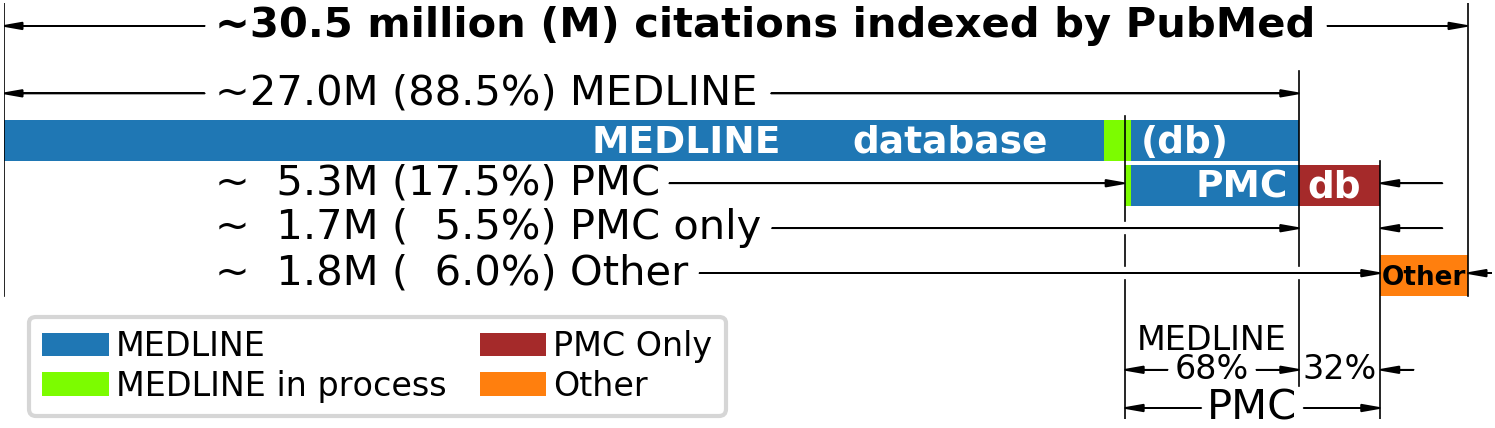

PubMed is a search interface and toolset used to access over 30.5 million article records from databases such as:
- MEDLINE: a highly selective database started in the 1960s
- PubMed Central (PMC): an open-access database for full-text papers that are free of cost
- Additional content such as books and articles published before the 1960s
Jupyter notebook examples
Jupyter notebook examples showing how to use the pmidcite Python library
- 1) Download NIH-OCC citation data
- 2) Download missing or load existing NIH-OCC citation data
- 3) Print a paper's citation and reference data
- 4) Sort NIH iCite entries
- 5) Query PubMed
Installation
To install from PyPI:
$ pip install -U pmidcite
# Or:
$ pip install --upgrade pmidcite
To install using Bioconda
$ conda install -c bioconda pmidcite
To install locally
$ git clone https://github.com/dvklopfenstein/pmidcite.git
$ cd ./pmidcite
$ pip install .
Setup
Save your literature search in a GitHub repo.
1. Add a pmidcite init file
Add a .pmidciterc init file to a non-git managed directory, such as home (~)
$ icite --generate-rcfile | tee ~/.pmidciterc
[pmidcite]
email = myname@email.edu
# To download PubMed search results, get an NCBI API key here:
# https://ncbiinsights.ncbi.nlm.nih.gov/2017/11/02/new-api-keys-for-the-e-utilities
apikey = MY_LONG_HEX_NCBI_API_KEY
tool = my_scripts
$ export PMIDCITECONF=~/.pmidciterc
Do not version manage the .pmidciterc using a tool such as GitHub because it
contains your personal email and your private NCBI API key.
2. NCBI E-Utils API key
To download PubMed abstracts and PubMed search results using NCBI's E-Utils,
get an NCBI API key using these instructions:
https://ncbiinsights.ncbi.nlm.nih.gov/2017/11/02/new-api-keys-for-the-e-utilities
Set the apikey value in the config file: ~/.pmidciterc
Contributing
See the contributing guide for detailed instructions on how to get started contributing to the pmidcite project.
Contact
email: dvklopfenstein@protonmail.com
https://orcid.org/0000-0003-0161-7603
How to Cite
If you use pmidcite in your research or literature search, please cite paper 1 (pmidcite) and paper 3 (NIH citation data).
Please also consider reading and citing Gusenbauer's response (paper 2) about improving search for all during the information avalanche of these times:
-
The pmidcite paper:
Commentary to Gusenbauer and Haddaway 2020: Evaluating Retrieval Qualities of PubMed and Google Scholar
Klopfenstein DV and Dampier W
2020 | Research Synthesis Methods | PMID: 33031632 | DOI: 10.1002/jrsm.1456 | pdf -
Gusenbauer's response to the pmidcite paper:
What every Researcher should know about Searching – Clarified Concepts, Search Advice, and an Agenda to improve Finding in Academia
Gusenbauer M and Haddaway N
2020 | Research Synthesis Methods | PMID: 33031639 | DOI: 10.1002/jrsm.1457 | pdf -
The NIH citation data used by pmidcite -- Scientific Influence, Translation, and Citation counts:
The NIH Open Citation Collection: A public access, broad coverage resource
Hutchins BI ... Santangelo GM
2019 | PLoS Biology | PMID: 31600197 | DOI: 10.1371/journal.pbio.3000385
References
Please consider reading and citing the paper [4] which inspired the creation of pmidcite [1] and the authors' response to our paper [2]:
- Which Academic Search Systems are Suitable for Systematic Reviews or Meta-Analyses? Evaluating Retrieval Qualities of Google Scholar, PubMed and 26 other Resources
Gusenbauer M and Haddaway N
2019 | Research Synthesis Methods | PMID: 31614060 | DOI: 10.1002/jrsm.1378
Mentioned in this README are also these outstanding contributions:
-
Relative Citation Ratio (RCR): A New Metric That Uses Citation Rates to Measure Influence at the Article Level
Hutchins BI, Xin Yuan, Anderson JM, and Santangelo, George M.
2016 | PLoS Biology | PMID: 27599104 | DOI: 10.1371/journal.pbio.1002541 -
Google Scholar as replacement for systematic literature searches: good relative recall and precision are not enough
Boeker M et al.
2013 | BMC Medical Research Methodology | PMID: 24160679 | DOI: 10.1186/1471-2288-13-131 -
Best Match: New relevance search for PubMed
Fiorini N ... Lu Zhiyong
2018 | PLoS Biology | PMID: 30153250 | DOI: 10.1371/journal.pbio.2005343
PDFs
- PMIDCITE Manuscript with the original text box formatting
- Supplemental Material
- Gusenbauer's Response
Contact
dvklopfenstein@protonmail.com
https://orcid.org/0000-0003-0161-7603
Copyright (C) 2019-present pmidcite, DV Klopfenstein, PhD. All rights reserved.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pmidcite-0.3.1.tar.gz.
File metadata
- Download URL: pmidcite-0.3.1.tar.gz
- Upload date:
- Size: 10.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
16a45509c04ba61802e912ac15aff5b71a5d2a5a0f29abd808ef5602ff37ebae
|
|
| MD5 |
10f77ea97ce9107440fd4b1a1f4434b6
|
|
| BLAKE2b-256 |
4198ecc9ff67993653990b9032a19ed39d9d33b42f11101369a7ca067497bacc
|
File details
Details for the file pmidcite-0.3.1-py3-none-any.whl.
File metadata
- Download URL: pmidcite-0.3.1-py3-none-any.whl
- Upload date:
- Size: 2.6 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e366e0c4439bb718e39650f1d4aac9265387d0cae3a6627976262255e7662f27
|
|
| MD5 |
03348d43179abedb5eed0a851cb9b0e1
|
|
| BLAKE2b-256 |
61c39e8b6cb940badec36ba5a43dea3894003f0262e7940498dd43a92dae1725
|