Physiology-Informed ECG Delineator Based on Peak Prominence
Project description
Physiology-Informed ECG Delineation Based on Peak Prominence
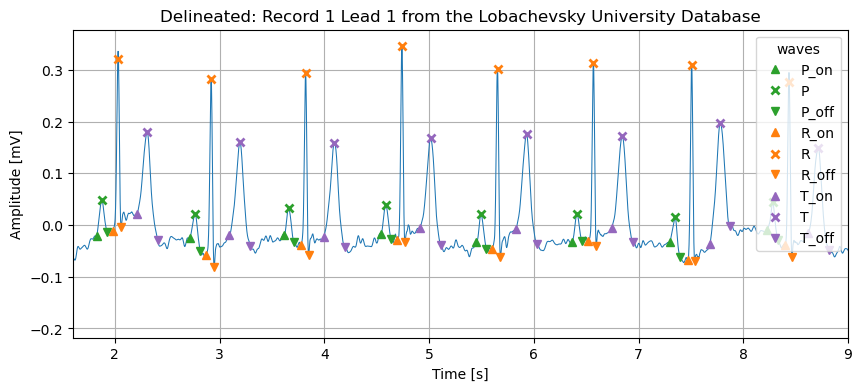
This Python package implements the Peak Prominence ECG Delineator [1] and provides methods for ECG cleaning and R-peak detection [2], resulting in a complete delineation pipeline. This delineator allows for fast and precise detection of the positions, on- and offsets of morphology waves (e.g., P, R, T) in single or multi-lead ECG signals. An optional multi-lead correction procedure can be applied, leveraging information from all leads if available.
Advantages and Limitations
This proposed approach achieves a highly explainable and interpretable wave selection by leveraging prominence information. Hence, wave detection only depends on physiologically motivated parameters chosen so that morphologies of interest are well represented and portrayed, yielding high $F_1$-scores and low errors on established Datasets in comparison to competing methods [1].
NOTE: This approach allows for further customization w.r.t to these parameters so that different parameter choices or other physiologically informed decision rules might result in higher performance or robustness regarding certain morphologies and heartbeat types. Even though the utilized approach for on- and offset detection yielded great performance, it is constrained by typical physiological boundaries. Developing novel prominence computation methods to robustly identify basepoints might therefore yield further improvements.
Limitation on QRS onset/offset: The prominence-based R onset/offset detection may not bracket the complete QRS complex, as the prominence windows around the R peak often don't extend completely to Q/S wave boundaries. Using R onset/R offset as QRS onset/offset estimates is exploratory and should be validated for critical applications. The method is optimized for P and T wave delineation.
Installation
You can install the latest version of the prominence-delineator package from the Python Package Index (PyPI) by running:
pip install prominence-delineator
Complete Working Example
A complete working example is provided in example.ipynb, additionally the basic usage is depicted below.
Basic Usage
The ProminenceDelineator takes the sampling_frequency and optionally several physiological parameters as input.
Then, R-peaks can be detected using any reliable R-peak detector or with the integrated method (applying the FastNVG [2] approach). Before detecting further waves, the required ECG cleaning can easily be performed by using .clean_ecg(ecg).
Finally, morphology waves can be detected using .find_waves() or .find_waves_multilead() for single or multi-lead ECG signals, respectively.
Note: For multi-lead delineation, the ecg and rpeaks input should be in the form of a list or ndarray containing the ECG signals or R-peaks of each lead, respectively. The output will be given as a dictionary with keys denoting the wave type and values lists containing wave positions or lists of wave positions for all leads when processing multi-lead data.
Single Lead ECG Delineation
from prominence_delineator import ProminenceDelineator
# Create an instance of the ProminenceDelineator
PromDelineator = ProminenceDelineator(sampling_frequency=fs)
# Detect the R-peaks in the ECG signal
rpeaks = PromDelineator.find_rpeaks(ecg)
# Clean the ECG signal
ecg = PromDelineator.clean_ecg(ecg)
# Find waves in the ECG signal using the ProminenceDelineator
waves = PromDelineator.find_waves(ecg, rpeaks=rpeaks)
Multi-Lead ECG Delineation
from prominence_delineator import ProminenceDelineator
# Create an instance of the ProminenceDelineator
PromDelineator = ProminenceDelineator(sampling_frequency=fs)
# Detect the R-peaks in the multilead ECG signal
multilead_rpeaks = PromDelineator.find_rpeaks(ecg_multilead)
# Clean the ECG signal
ecg_multilead = PromDelineator.clean_ecg(ecg_multilead)
# Find waves in the ECG signal using the find_waves function
multilead_waves = PromDelineator.find_waves_multilead(ecg_multilead, rpeaks_multilead=multilead_rpeaks)
References
- [1] Emrich, J., Gargano, A., Koka, T., and Muma, M. (2024), Physiology-Informed ECG Delineation Based on Peak Prominence.
- [2] Emrich, J., Koka, T., Wirth, S. and Muma, M. (2023). Accelerated Sample-Accurate R-Peak Detectors Based on Visibility Graphs.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file prominence_delineator-0.0.10.tar.gz.
File metadata
- Download URL: prominence_delineator-0.0.10.tar.gz
- Upload date:
- Size: 46.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c0214a0de4ca04d4619ab8a962c85a6f9c2dfd0c7e6334837062acb144d007a8
|
|
| MD5 |
127e3df3a97ec5ec393a625dc3b71f9a
|
|
| BLAKE2b-256 |
07ac65e96a70e6a72042e50e12260dabd2d701d86d1f81d0b91804bc3e893300
|
File details
Details for the file prominence_delineator-0.0.10-py3-none-any.whl.
File metadata
- Download URL: prominence_delineator-0.0.10-py3-none-any.whl
- Upload date:
- Size: 33.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
206ef885abef6104d7baefdb65ce2dc2ad0ca7fdc84775468b3d0f330bc2b580
|
|
| MD5 |
1060d87deb20548f90eaeee7264dd1b3
|
|
| BLAKE2b-256 |
535a53d0865449a00640d1a41caf05f8ea66d61547a734024f50dd07177eb2c4
|