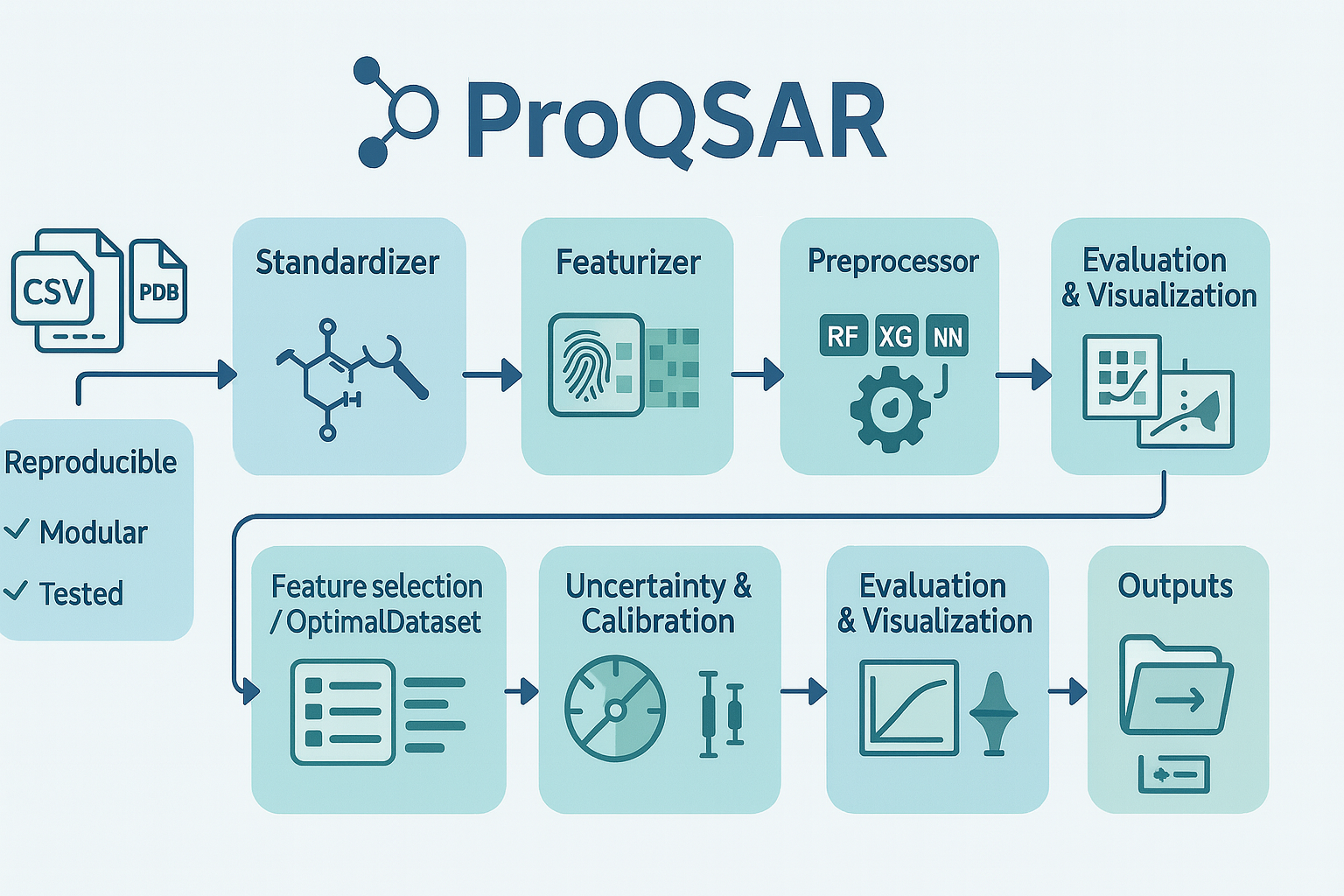
ProQSAR — automation utilities for QSAR workflows (standardization, featurization, model building & evaluation).
Project description
ProQSAR
ProQSAR — automatic pipeline for quantitative structure–activity relationship (QSAR) modeling
A reproducible toolkit for end-to-end QSAR: data standardization, featurization, splitting, model training, uncertainty estimation, and evaluation. Designed for reproducible experiments, continuous integration, and easy integration into ML/CADD pipelines. Full documentation for ProQSAR is available at ReadTheDocs.
Key features
- Data standardization and sanitization (SMILES normalization, valence checks, tautomer/charge handling).
- Modular featurizers: fingerprints, descriptors, learned featurizers (pluggable API).
- Flexible dataset splitting: random, scaffold, stratified.
- Built-in pipelines for training and evaluation with uncertainty estimation.
- Simple CLI and Python API for reproducible experiments and batch processing.
- CI-tested with unit tests and example notebooks.
Installation
Choose the preferred installation method.
From PyPI
pip install proqsar
From conda (anaconda.org/tieulongphan)
conda install -c tieulongphan proqsar
Docker (containerized)
docker pull tieulongphan/proqsar:latest
# run an example container (bind-mount your project directory)
docker run --rm -v $(pwd):/workspace -w /workspace tieulongphan/proqsar:latest proqsar --help
From source (developer)
git clone https://github.com/Medicine-Artificial-Intelligence/proqsar.git
cd proqsar
pip install -e .[dev]
Development & contributing
Thanks for your interest in contributing! A quick checklist:
- Fork the repository and create a feature branch.
- Implement your changes and include unit tests.
- Run linting and tests locally (
pre-commit,flake8,pytest). - Open a Pull Request describing the change and add tests/examples.
Citation / Publication
If you use ProQSAR in research, please cite the project. Example BibTeX placeholder:
@misc{proqsar2025,
title = {ProQSAR: Automatic pipeline for QSAR modeling},
author = {Tuyet-Minh Phan and Tieu-Long Phan and Phuoc-Chung Nguyen Van and contributors},
year = {2025},
howpublished = {\url{https://github.com/Medicine-Artificial-Intelligence/proqsar}}
}
Authors & Contributors
License
This project is licensed under MIT License - see the License file for details.
Acknowledgments
This work has received support from the Korea International Cooperation Agency (KOICA) under the project entitled “Education and Research Capacity Building Project at University of Medicine and Pharmacy at Ho Chi Minh City”, conducted from 2024 to 2025 (Project No. 2021-00020-3).
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file proqsar-0.0.9.tar.gz.
File metadata
- Download URL: proqsar-0.0.9.tar.gz
- Upload date:
- Size: 3.7 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d402491204ea24066ba53ae04c63806d2ad2c032bc355c359920ba34c2297572
|
|
| MD5 |
5219a9f0ffff0ebbc0c521ffd2fcd888
|
|
| BLAKE2b-256 |
efd30c2bbfd501d55d61dad2ef1b9c88fa03bdeb18c14808a143d4b5cac44eff
|
Provenance
The following attestation bundles were made for proqsar-0.0.9.tar.gz:
Publisher:
publish-package.yml on Medicine-Artificial-Intelligence/ProQSAR
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
proqsar-0.0.9.tar.gz -
Subject digest:
d402491204ea24066ba53ae04c63806d2ad2c032bc355c359920ba34c2297572 - Sigstore transparency entry: 563655334
- Sigstore integration time:
-
Permalink:
Medicine-Artificial-Intelligence/ProQSAR@aa5897fae146c23a46fb52a69a5f669cb62a035f -
Branch / Tag:
refs/tags/v0.0.9 - Owner: https://github.com/Medicine-Artificial-Intelligence
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-package.yml@aa5897fae146c23a46fb52a69a5f669cb62a035f -
Trigger Event:
release
-
Statement type:
File details
Details for the file proqsar-0.0.9-py3-none-any.whl.
File metadata
- Download URL: proqsar-0.0.9-py3-none-any.whl
- Upload date:
- Size: 140.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d3c82d7f6c874cdd5e360fc2c2453f0f8f2aa6a39b9fb0d5f5e91fdddcdc73f1
|
|
| MD5 |
4c6c4e394b4413ba9380159e4a2726b7
|
|
| BLAKE2b-256 |
3ed59e32d4d67d7c1499dc9113cc8b78fe827a698702bad792543b7fa6220543
|
Provenance
The following attestation bundles were made for proqsar-0.0.9-py3-none-any.whl:
Publisher:
publish-package.yml on Medicine-Artificial-Intelligence/ProQSAR
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
proqsar-0.0.9-py3-none-any.whl -
Subject digest:
d3c82d7f6c874cdd5e360fc2c2453f0f8f2aa6a39b9fb0d5f5e91fdddcdc73f1 - Sigstore transparency entry: 563655339
- Sigstore integration time:
-
Permalink:
Medicine-Artificial-Intelligence/ProQSAR@aa5897fae146c23a46fb52a69a5f669cb62a035f -
Branch / Tag:
refs/tags/v0.0.9 - Owner: https://github.com/Medicine-Artificial-Intelligence
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-package.yml@aa5897fae146c23a46fb52a69a5f669cb62a035f -
Trigger Event:
release
-
Statement type: