PyAMARES, an Open-Source Python Library for Fitting Magnetic Resonance Spectroscopy Data
Project description

The full documentation for pyAMARES can be found at pyAMARES Documentation.
What is pyAMARES?
The pyAMARES package provides the MRS community with an open-source, easy-to-use MRS fitting method in Python. It imports prior knowledge from Excel or CSV spreadsheets as initial values and constraints for fitting MRS data according to the AMARES model function.
Getting Started
Requirements


Installation
pip install pyAMARESSee the Installation Guide for detailed information.
Run pyAMARES as standard-alone script
amaresFit -f ./pyAMARES/examples/fid.txt -p ./pyAMARES/examples/example_human_brain_31P_7T.csv --MHz 120.0 --sw 10000 --deadtime 300e-6 --ifplot --xlim 10 -20 -o simple_exampleRun pyAMARES in a Jupyter Notebook
Try Jupyter Notebook on Google Colab here
import pyAMARES
# Load FID from a 2-column ASCII file, and set the MR parameters
MHz = 120.0 # 31P nuclei at 7T
sw = 10000 # spectrum width in Hz
deadtime = 300e-6 # 300 us begin time for the FID signal acquisition
fid = pyAMARES.readmrs('./pyAMARES/examples/fid.txt')
# Load Prior Knowledge
FIDobj = pyAMARES.initialize_FID(fid=fid,
priorknowledgefile='./pyAMARES/examples/example_human_brain_31P_7T.csv',
MHz=MHz,
sw=sw,
deadtime=deadtime,
preview=False,
normalize_fid=False,
xlim=(10, -20))# Region of Interest for visualization, -20 to 10 ppm
# Initialize the parameter using Levenberg-Marquard method
out1 = pyAMARES.fitAMARES(fid_parameters=FIDobj,
fitting_parameters=FIDobj.initialParams,
method='leastsq',
ifplot=False)
# Fitting the MRS data using the optimized parameter
out2 = pyAMARES.fitAMARES(fid_parameters=out1,
fitting_parameters=out1.fittedParams, # optimized parameter for last step
method='least_squares',
ifplot=False)
# Save the data
out2.styled_df.to_html('simple_example.html') # Save highlighted table to an HTML page
# Python 3.6 does not support to_html.
out2.result_sum.to_csv('simple_example.csv') # Save table to CSV spreadsheet
out2.plotParameters.lb = 2.0 # Line Broadening factor for visualization
out2.plotParameters.ifphase = True # Phase the spectrum for visualization
pyAMARES.plotAMARES(fid_parameters=out1, filename='simple_example.svg') # Save plot to SVGFitting Result for Example 31P MRS data

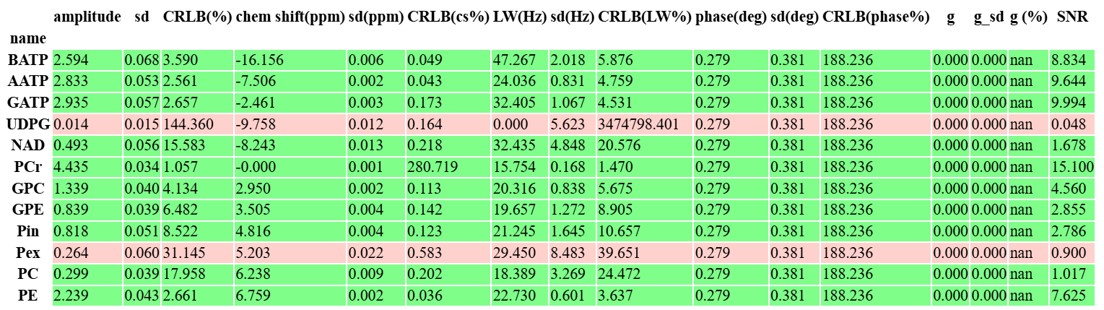
Contributing
PyAMARES is currently in its early stages of development and is actively being improved. We welcome contributions to pyAMARES! Please see our CONTRIBUTING.rst guidelines for more information on how to get started.
How to cite
If you use pyAMARES in your research, please cite:
Xu, J.; Vaeggemose, M.; Schulte, R.F.; Yang, B.; Lee, C.-Y.; Laustsen, C.; Magnotta, V.A. PyAMARES, an Open-Source Python Library for Fitting Magnetic Resonance Spectroscopy Data. Diagnostics 2024, 14, 2668. https://doi.org/10.3390/diagnostics14232668
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pyamares-0.3.28.tar.gz.
File metadata
- Download URL: pyamares-0.3.28.tar.gz
- Upload date:
- Size: 929.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2a6790faf485861bbaad55dbcfb204d7ec7347db2aadfb44d0046cbdab80ed30
|
|
| MD5 |
31ddd7c7469ee54e4051062de9e4b92d
|
|
| BLAKE2b-256 |
97bdc251f129e717ee7f68d58b39b1bb27497f94a8226e49ed76d1ca4c4b89d6
|
File details
Details for the file pyamares-0.3.28-py3-none-any.whl.
File metadata
- Download URL: pyamares-0.3.28-py3-none-any.whl
- Upload date:
- Size: 258.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fc77b2711600e06912d1101c6f6de687677766c805d3b0cc3bec47e76a33940a
|
|
| MD5 |
42c5f34d463f661bbc8b97280baa9a3d
|
|
| BLAKE2b-256 |
d6997083f7ff079927abae51cc55d8c41e8293a3c16476d66343af43f4f4107a
|











