An eXplainable Cell-specific machine learning method to predict clinical Phenotypes using single-cell multi-omics
Project description

Getting Started...
Here, we introduce CellPhenoX, an eXplainable machine learning method to identify cell-specific phenotypes that influence clinical outcomes for single-cell data. CellPhenoX integrates robust classification models, explainable AI techniques, and a statistical covariate framework to generate interpretable, cell-specific scores that uncover cell populations associated with a clinical phenotype of interest.
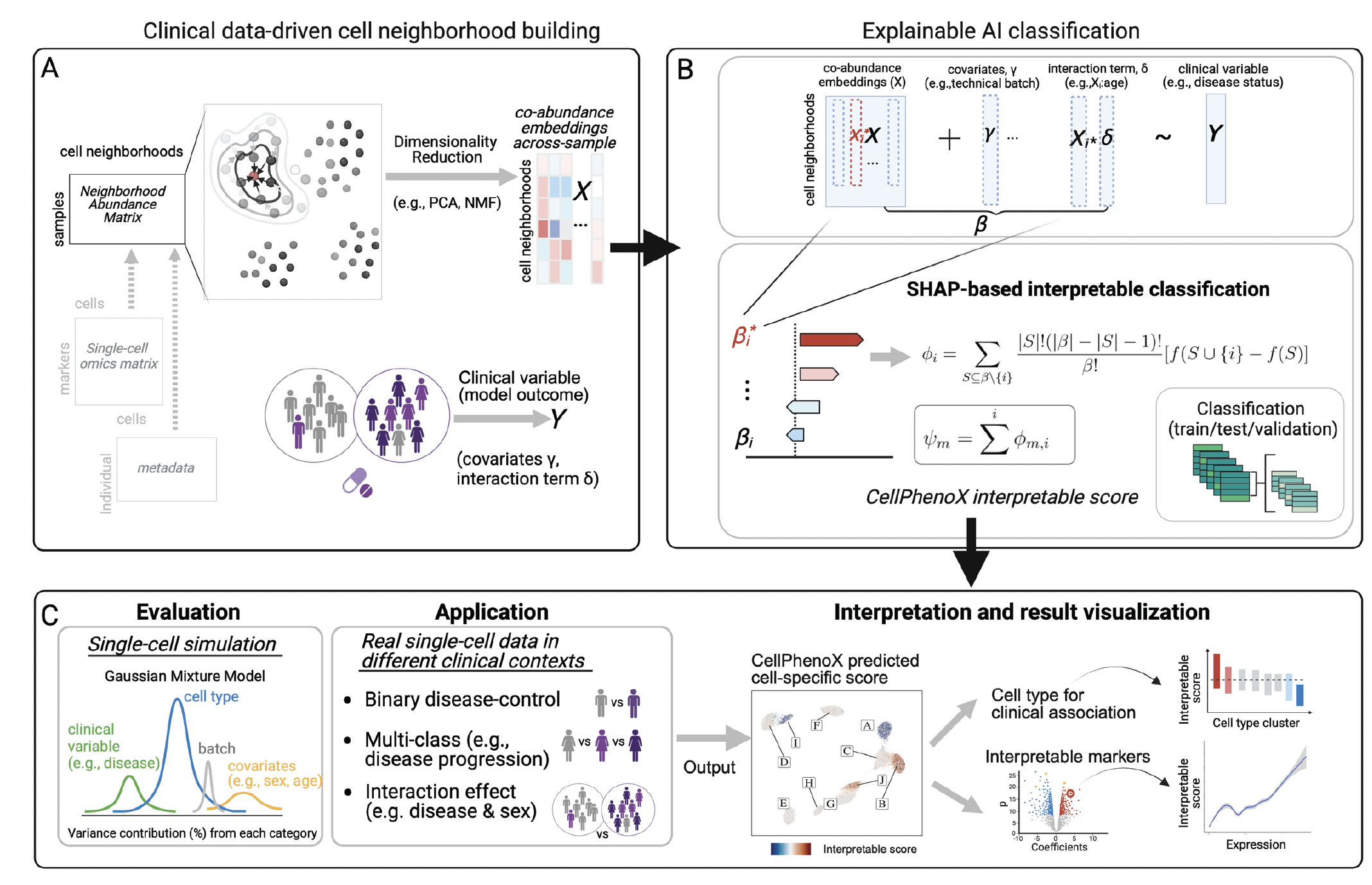
Figure 1. CellPhenoX leverages cell neighborhood co-abundance embeddings, Xi , across samples and clinical variable Y as inputs. By applying an adapted SHAP framework for classification models, CellPhenoX generates Interpretable Scores that quantify the contribution of each feature Xi, along with covariates and interaction term Xi, to the prediction of a clinically relevant phenotype Y. The results are visualized at single-cell level, showcasing Interpretable Scores at low-dimensional space, correlated cell type annotations, and associated marker genes.
You can install pyCellPhenoX from PyPI:
pip install pyCellPhenoX
github (link):
# install pyCellPhenoX directly from github
git clone git@github.com:fanzhanglab/pyCellPhenoX.git
Dependencies/ Requirements
When using pyCellPhenoX please ensure you are using the following dependency versions or requirements
python = "^3.9"
pandas = "^2.2.3"
numpy = "^2.1.1"
xgboost = "^2.0"
numba = ">=0.54"
shap = "^0.46.0"
scikit-learn = "^1.5.2"
matplotlib = "^3.9.2"
statsmodels = "^0.14.3"
Tutorials
Please see the Command-line Reference for details. Additonally, please see Vignettes on the documentation page.
API
pyCellPhenoX has four major functions which are apart of the object:
- split_data() - Split the data into training, testing, and validation sets
- model_train_shap_values() - Train the model using nested cross validation strategy and generate shap values for each fold/CV repeat
- get_shap_values() - Aggregate SHAP values for each sample
- get_intepretable_score() - Calculate the interpretable score based on SHAP values.
Additional major functions associated with pyCellPhenoX are:
- marker_discovery() - Identify markers correlated with the discriminatory power of the Interpretable Score.
- nonNegativeMatrixFactorization() - Perform non Negative Matrix Factorization (NMF)
- preprocessing() - Prepare the data to be in the correct format for CellPhenoX
- principleComponentAnalysis() - Perform Principle Component Analysis (PCA)
Each function has uniqure arguments, see our documentation for more information
License
Distributed under the terms of the MIT license, pyCellPhenoX is free and open source software.
Code of Conduct
For more information please see Code of Conduct or Code of Conduct Documentation
Contributing
For more information please see Contributing or Contributing Documentation
Issues
If you encounter any problems, please file an issue along with a detailed description.
Citation
If you have used pyCellPhenoX in your project, please use the citation below:
Young, J., Inamo, J., Caterer, Z., Krishna, R., Zhang, F. CellPhenoX: An eXplainable Cell-specific machine learning method to predict clinical Phenotypes using single-cell multi-omics, in submission, 2025.
Contact
Please contact fanzhanglab@gmail.com for further questions or protential collaborative opportunities!
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pycellphenox-1.0.8.tar.gz.
File metadata
- Download URL: pycellphenox-1.0.8.tar.gz
- Upload date:
- Size: 90.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.5 CPython/3.12.6 Darwin/22.4.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
92f376b742c770e75f0b523899371dd2b1656c9c5fa7bd2bbec457a60c802e81
|
|
| MD5 |
46192d11dc0d1321e7c7585795acb23d
|
|
| BLAKE2b-256 |
ab3ad3862b04d21582129e9fd84eecb3c8fc4dd371f8aa5d814b397d5c827dc4
|
File details
Details for the file pycellphenox-1.0.8-py3-none-any.whl.
File metadata
- Download URL: pycellphenox-1.0.8-py3-none-any.whl
- Upload date:
- Size: 20.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.5 CPython/3.12.6 Darwin/22.4.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7b283886431849aa2954521c8a4a6aff041f2b3aa0048d8fae7482003cb14acb
|
|
| MD5 |
ba2dcfa14b0addf9b93e10e63e5f3b30
|
|
| BLAKE2b-256 |
3bace6a25ccc4c157cca7048158530672055a1b02253c93f552ac542f3f5dffe
|


















