Circular visualization in Python, plotly implemented
Project description
pyCirclizely: Circular visualization in Python, plotly implemented
Table of contents
Overview
pyCirclizely is a circular visualization python package refactored from pyCirclize to use plotly instead of matplotlib, which in turn was inspired by circlize and pyCircos.
It includes useful genome and phylogenetic tree visualization methods for the bioinformatics field. More detailed documentation is available here.
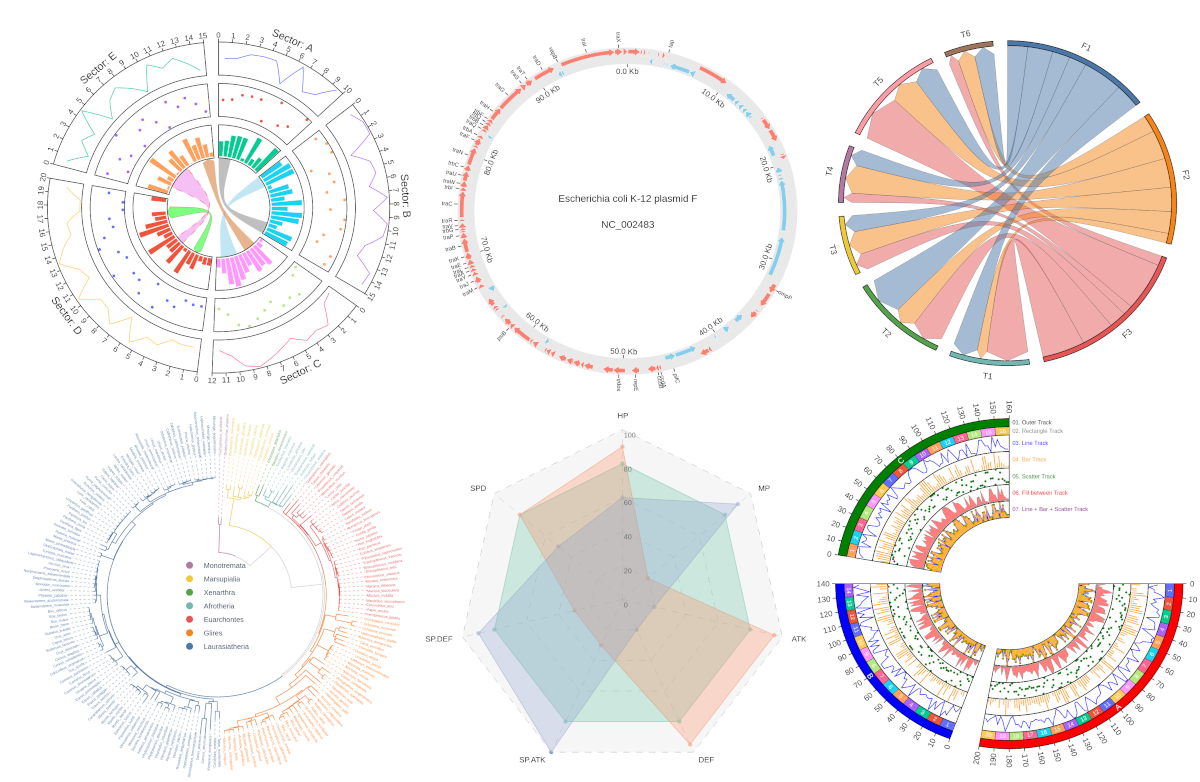
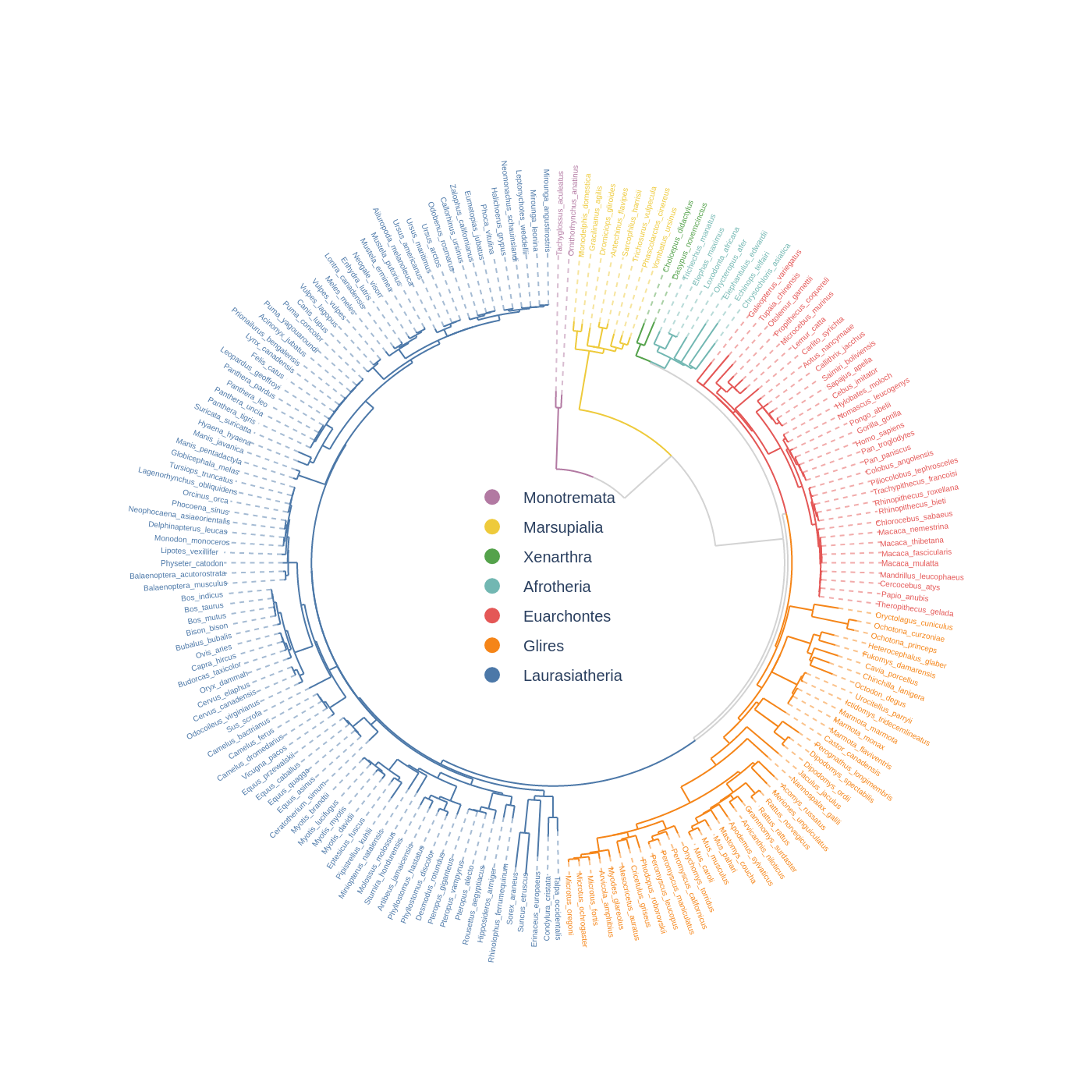
Fig.1 pyCirclizely example plot gallery
Installation
Python 3.10 or later is required for installation.
Install PyPI package:
pip install pycirclizely
Install conda-forge package:
conda install -c conda-forge pycirclizely
API Usage
API usage is described in each of the following sections in the documentation.
- Getting Started
- Plot API Example
- Chord Diagram
- Radar Chart
- Circos Genomics Plot (Prokaryotes)
- Circos Genomics Plot (Eukaryotes)
- Comparative Genomics
- Phylogenetic Tree
- Plot Tips
Code Example
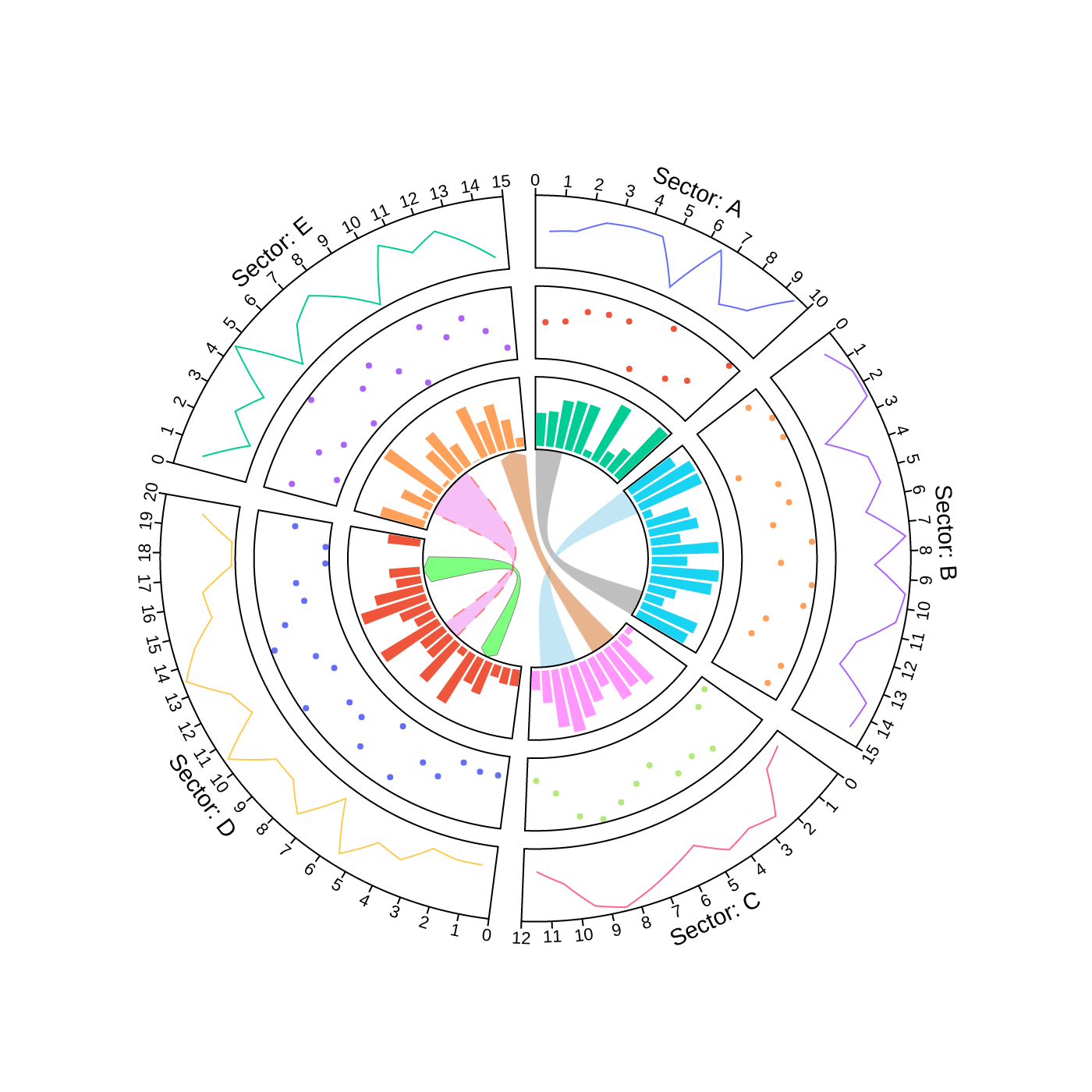
1. Circos Plot
from pycirclizely import Circos
import numpy as np
np.random.seed(0)
# Initialize Circos sectors
sectors = {"A": 10, "B": 15, "C": 12, "D": 20, "E": 15}
circos = Circos(sectors, space=5)
for sector in circos.sectors:
# Plot sector name
sector.text(f"Sector: {sector.name}", r=110, font=dict(size=15))
# Create x positions & random y values
x = np.arange(sector.start, sector.end) + 0.5
y = np.random.randint(0, 100, len(x))
# Plot lines
track1 = sector.add_track((80, 100), r_pad_ratio=0.1)
track1.xticks_by_interval(interval=1)
track1.axis()
track1.line(x, y)
# Plot points
track2 = sector.add_track((55, 75), r_pad_ratio=0.1)
track2.axis()
track2.scatter(x, y)
# Plot bars
track3 = sector.add_track((30, 50), r_pad_ratio=0.1)
track3.axis()
track3.bar(x, y)
# Plot links
circos.link(("A", 0, 3), ("B", 15, 12))
circos.link(("B", 0, 3), ("C", 7, 11), fillcolor="skyblue")
circos.link(("C", 2, 5), ("E", 15, 12), fillcolor="chocolate", direction=1)
circos.link(
("D", 3, 5), ("D", 18, 15), fillcolor="lime",
line=dict(color="black", width=0.5), direction=2,
)
circos.link(
("D", 8, 10), ("E", 2, 8), fillcolor="violet",
line=dict(color="red", width=1.0, dash="dash"),
)
fig = circos.plotfig()
fig.write_image("example01.png", format="png")
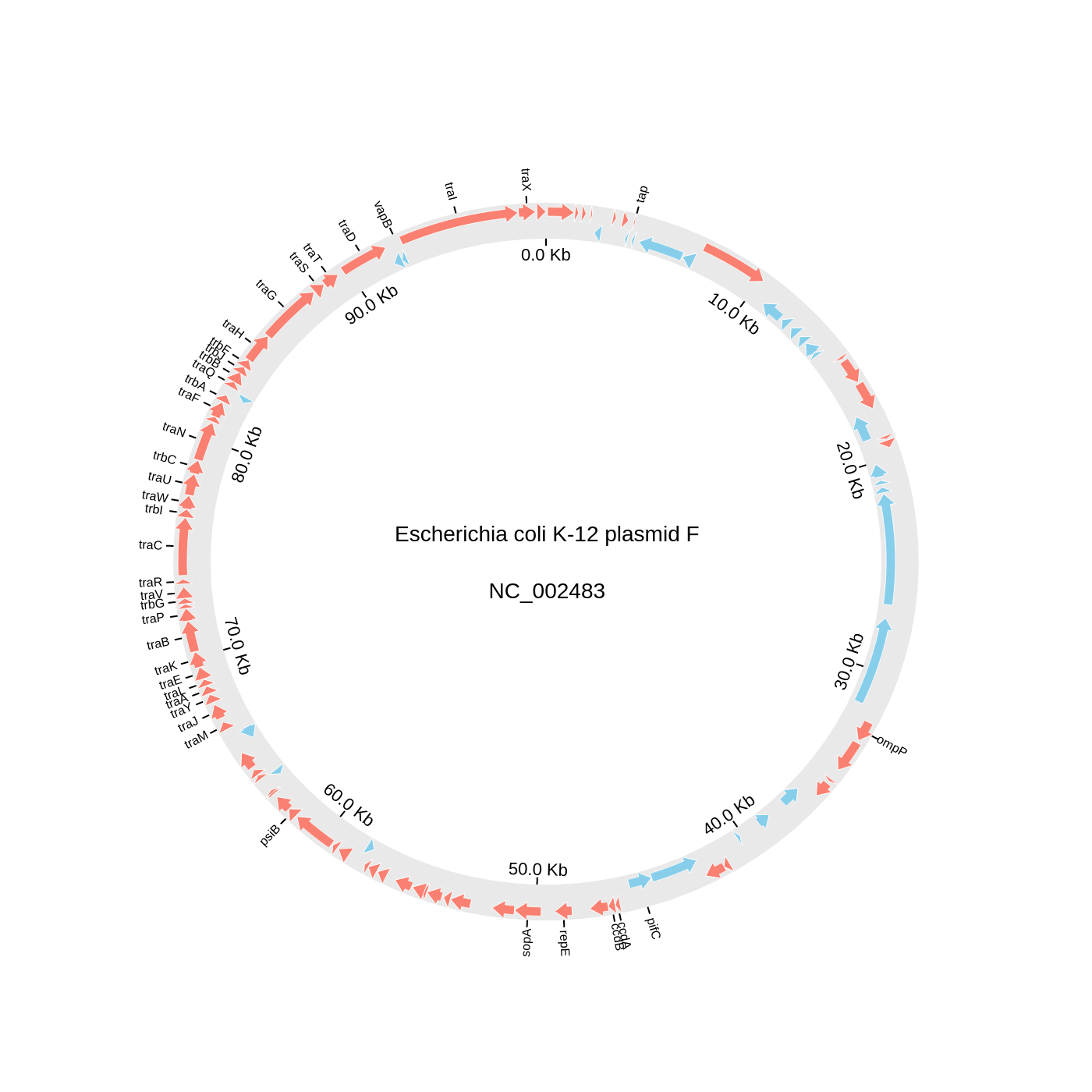
2. Circos Plot (Genomics)
from pycirclizely import Circos
from pycirclizely.utils import fetch_genbank_by_accid
from pycirclizely.parser import Genbank
# Download `NC_002483` E.coli plasmid genbank
gbk_fetch_data = fetch_genbank_by_accid("NC_002483")
gbk = Genbank(gbk_fetch_data)
# Initialize Circos instance with genome size
sectors = gbk.get_seqid2size()
space = 0 if len(sectors) == 1 else 2
circos = Circos(sectors, space=space)
circos.text(f"Escherichia coli K-12 plasmid F<br><br>{gbk.name}", font=dict(size=14))
seqid2features = gbk.get_seqid2features(feature_type="CDS")
for sector in circos.sectors:
# Setup track for features plot
f_cds_track = sector.add_track((95, 100))
f_cds_track.axis(fillcolor="lightgrey", line=dict(width=0), opacity=0.5)
r_cds_track = sector.add_track((90, 95))
r_cds_track.axis(fillcolor="lightgrey", line=dict(width=0), opacity=0.5)
# Plot forward/reverse strand CDS
features = seqid2features[sector.name]
for feature in features:
if feature.location.strand == 1:
f_cds_track.genomic_features(
feature, plotstyle="arrow", fillcolor="salmon", line=dict(width=0.5)
)
else:
r_cds_track.genomic_features(
feature, plotstyle="arrow", fillcolor="skyblue", line=dict(width=0.5)
)
# Plot 'gene' qualifier label if exists
labels, label_pos_list = [], []
for feature in features:
start = int(feature.location.start)
end = int(feature.location.end)
label_pos = (start + end) / 2
gene_name = feature.qualifiers.get("gene", [None])[0]
if gene_name is not None:
labels.append(gene_name)
label_pos_list.append(label_pos)
f_cds_track.xticks(
label_pos_list, labels, label_orientation="vertical", text_kws=dict(font=dict(size=8))
)
# Plot xticks (interval = 10 Kb)
r_cds_track.xticks_by_interval(
10000, outer=False, label_formatter=lambda v: f"{v/1000:.1f} Kb"
)
fig = circos.plotfig()
fig.write_image("example02.png", format="png")
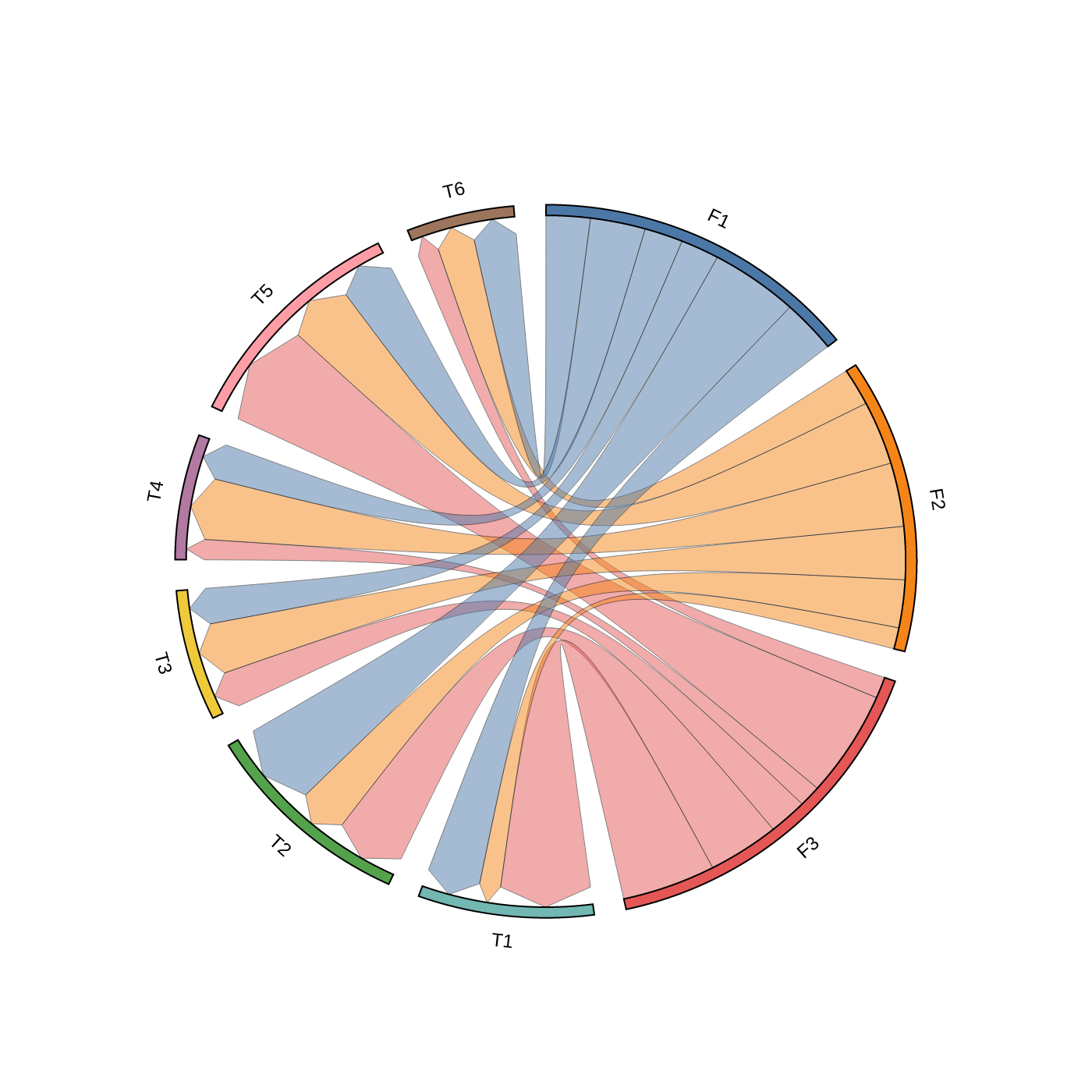
3. Chord Diagram
from pycirclizely import Circos
import pandas as pd
# Create matrix dataframe (3 x 6)
row_names = ["F1", "F2", "F3"]
col_names = ["T1", "T2", "T3", "T4", "T5", "T6"]
matrix_data = [
[10, 16, 7, 7, 10, 8],
[4, 9, 10, 12, 12, 7],
[17, 13, 7, 4, 20, 4],
]
matrix_df = pd.DataFrame(matrix_data, index=row_names, columns=col_names)
# Initialize Circos instance for chord diagram plot
circos = Circos.chord_diagram(
matrix_df,
space=5,
cmap="T10",
label_kws=dict(font=dict(size=12)),
link_kws=dict(line=dict(color="black", width=0.5), direction=1),
)
fig = circos.plotfig()
fig.write_image("example03.png", format="png")
4. Phylogenetic Tree
from plotly import graph_objects as go
from pycirclizely import Circos
from pycirclizely.utils import load_example_tree_file, ColorCycler
# Initialize Circos from phylogenetic tree
tree_file = load_example_tree_file("large_example.nwk")
circos, tv = Circos.initialize_from_tree(
tree_file,
r_lim=(30, 100),
leaf_label_size=5,
line_kws=dict(line=dict(color="lightgrey", width=1.0)),
)
# Define group-species dict for tree annotation
# In this example, set minimum species list to specify group's MRCA node
group_name2species_list = dict(
Monotremata=["Tachyglossus_aculeatus", "Ornithorhynchus_anatinus"],
Marsupialia=["Monodelphis_domestica", "Vombatus_ursinus"],
Xenarthra=["Choloepus_didactylus", "Dasypus_novemcinctus"],
Afrotheria=["Trichechus_manatus", "Chrysochloris_asiatica"],
Euarchontes=["Galeopterus_variegatus", "Theropithecus_gelada"],
Glires=["Oryctolagus_cuniculus", "Microtus_oregoni"],
Laurasiatheria=["Talpa_occidentalis", "Mirounga_leonina"],
)
# Set tree line color & label color
color_cycler = ColorCycler("T10")
groups = list(group_name2species_list.keys())
colors = color_cycler.get_colors(len(groups))[::-1]
group_name2color = {name: color for name, color in zip(groups, colors)}
for group_name, species_list in group_name2species_list.items():
color = group_name2color[group_name]
tv.set_node_line_props(
species_list, line=dict(color=color), apply_label_color=True
)
# Plot figure & set legend on center
fig = circos.plotfig()
for group_name, color in group_name2color.items():
# Use a dummy trace to create a legend entry
fig.add_trace(
go.Scatter(
x=[None],
y=None,
mode="markers",
marker=dict(color=color, size=10),
name=group_name,
showlegend=True,
)
)
fig.update_layout(
legend=dict(
x=0.5,
y=0.47,
xanchor="center",
yanchor="middle",
font=dict(size=10),
bgcolor="rgba(255,255,255,0.0)",
)
)
fig.write_image("example04.png", format="png", scale=2)
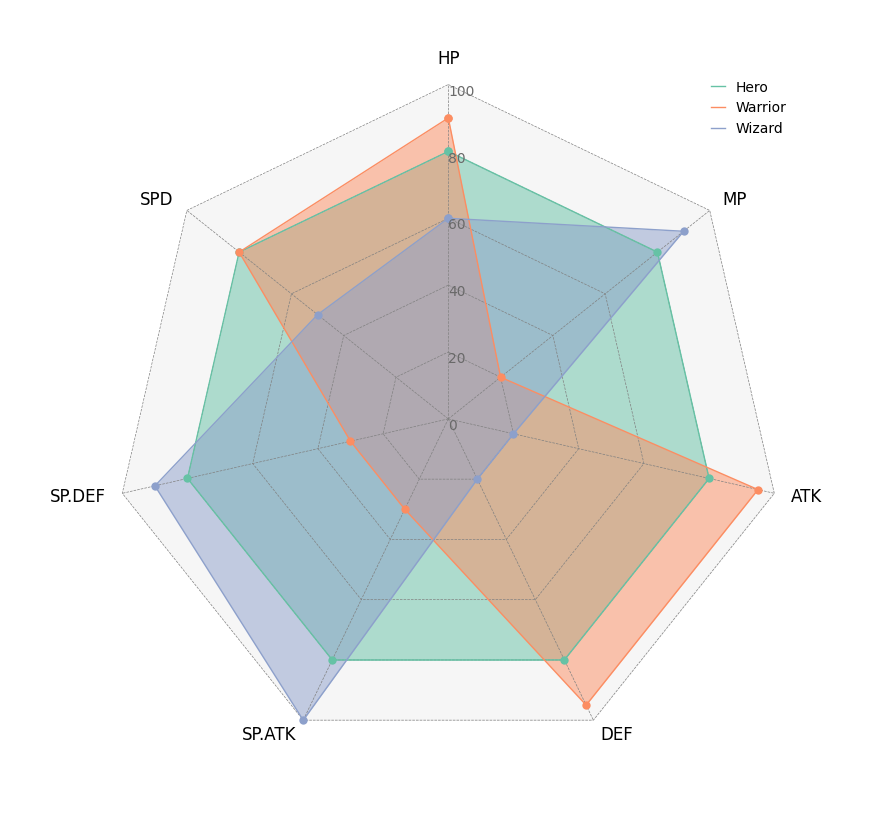
5. Radar Chart
from pycirclizely import Circos
import pandas as pd
# Create RPG jobs parameter dataframe (3 jobs, 7 parameters)
df = pd.DataFrame(
data=[
[80, 80, 80, 80, 80, 80, 80],
[90, 20, 95, 95, 30, 30, 80],
[60, 90, 20, 20, 100, 90, 50],
],
index=["Hero", "Warrior", "Wizard"],
columns=["HP", "MP", "ATK", "DEF", "SP.ATK", "SP.DEF", "SPD"],
)
# Initialize Circos instance for radar chart plot
circos = Circos.radar_chart(
df,
vmax=100,
marker_size=6,
grid_interval_ratio=0.2,
)
# Plot figure & set legend on upper right
fig = circos.plotfig()
fig.write_image("example05.png", format="png", scale=2)
Not Implemented Features
Further features to be implemented are the plotly built-in subplot and legend (complementary to the hover text) framework in the package.
Moreover, support for other types of hover info besides hover text will improve the user-plot interaction.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pycirclizely-1.0.4.tar.gz.
File metadata
- Download URL: pycirclizely-1.0.4.tar.gz
- Upload date:
- Size: 3.5 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.9.25 {"installer":{"name":"uv","version":"0.9.25","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"Ubuntu","version":"25.10","id":"questing","libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5a338bec87d32d7429abadcbd83eab89e4912a37ee9e22488d285b0ab43b661d
|
|
| MD5 |
67141272a04721fb74cc0238dc964391
|
|
| BLAKE2b-256 |
44ec3565456d7fb35bec1d3cb8ad2eac424cabec6cb849e4cd5c75ea13f5a3b7
|
File details
Details for the file pycirclizely-1.0.4-py3-none-any.whl.
File metadata
- Download URL: pycirclizely-1.0.4-py3-none-any.whl
- Upload date:
- Size: 79.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.9.25 {"installer":{"name":"uv","version":"0.9.25","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"Ubuntu","version":"25.10","id":"questing","libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
eca339ae8f9488d7aba45af254e88ed7545a0cebdce5f9772b8cb3259ef711d7
|
|
| MD5 |
99f5237c933a7edc2a995e9e9a98eef9
|
|
| BLAKE2b-256 |
68fa9cc86c6ecc2004bed6ad2ac0a709a2eeb26c7327b5a41f35a28c2cbecac6
|