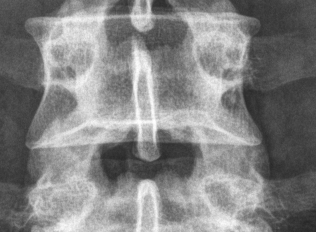
Simple open-source DICOM browser/viewer in Python and Qt4.
Project description
pydiq
Simple open-source multi-platform DICOM browser/viewer in Python and Qt.
NOTE This project has not been updated for a long time. Currently, I have no capacity to improve it. If you feel like contributing, I'll be happy to accept your enhancements / bug fixes. The UI seems not to be entirely working...
Features
- Easy (and fast) viewing of all images in a directory
- Zooming (1:N and N:1)
- Mouse control of window center and width (as in Aeskulap Viewer)
- Proper measurement of Hounsfield units and position by mouse
- PNG image export
To Do
- Better zooming
- Better MRI images support
- RT dose images support
- View in different planes (rectangular + others)
- Coordinate mapping (using translation and rotation matrix)
- Integration of anonymization features (see https://github.com/janpipek/anonymize_dicom )
- Information from the DICOM file in user-friendly display
Dependencies
- Python 3.6+
- qtpy (and therefore PyQt4 / PyQt5 / PySide - not automatically installed by pip!)
- pydicom (1.3)
Tested on Linux and Windows.
Installation
The easiest way is pip install pydiq.
Usage
Usage: pydiq [OPTIONS] [PATH]
Options:
--help Show this message and exit.
Limitations
Currently, the viewer supports only Computed Radiography (CR), Computed Tomography (CT) and Magnetic Resonance Imaging (MRI) images with normal orientation (x, y, z) in one-slice-per-file format.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file pydiq-0.2.2.tar.gz.
File metadata
- Download URL: pydiq-0.2.2.tar.gz
- Upload date:
- Size: 10.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/2.0.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.2.0 requests-toolbelt/0.9.1 tqdm/4.36.1 CPython/3.7.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d589db78ecef1f01c4f8df9f5ae27d6856e3c5dc42fdc6d551c3e73dd42bbfc5
|
|
| MD5 |
6ed4ab8cd6f5a5ec382a915631a9649b
|
|
| BLAKE2b-256 |
96518e88078491bc7fe193da4b58b275196c504bca79d3e8817d4b8031e93c9b
|