A fast generic spectrum simulator
Project description
PyEchelle
PyEchelle is a simulation tool, to generate realistic 2D spectra, in particular cross-dispersed echelle spectra. However, it is not limited to echelle spectrographs, but allows simulating arbitrary spectra for any fiber-fed or slit spectrograph, where a model file is available. Optical aberrations are treated accurately, the simulated spectra include photon and read-out noise.
PyEchelle uses numba for implementing fast Python-based simulation code. It also comes with CUDA support for major speed improvements.
Example usage
You can use PyEchelle directly from the console:
pyechelle --spectrograph MaroonX --fiber 2-4 --sources Phoenix --phoenix_t_eff 3500 -t 10 --rv 100 -o mdwarf.fit
If you rather script in python, you can do the same as above with the following python script:
from pyechelle.simulator import Simulator
from pyechelle.sources import Phoenix
from pyechelle.spectrograph import ZEMAX
sim = Simulator(ZEMAX("MaroonX"))
sim.set_fibers([2, 3, 4])
sim.set_sources(Phoenix(t_eff=3500))
sim.set_exposure_time(10.)
sim.set_radial_velocities(100.)
sim.set_output('mdwarf.fits', overwrite=True)
sim.run()
Both times, a PHOENIX M-dwarf spectrum with the given stellar parameters, and a RV shift of 100m/s for the MAROON-X spectrograph is simulated.
The output is a 2D raw frame (.fits) and will look similar to:
Check out the Documentation for more examples.
Pyechelle is the successor of Echelle++ which has a similar functionality but was written in C++. This package was rewritten in python for better maintainability, easier package distribution and for smoother cross-platform development.
Installation
As simple as
pip install pyechelle
Check out the Documentation for alternative installation instruction.
Usage
See
pyechelle -h
for all available command line options.
See Documentation for more examples.
Concept:
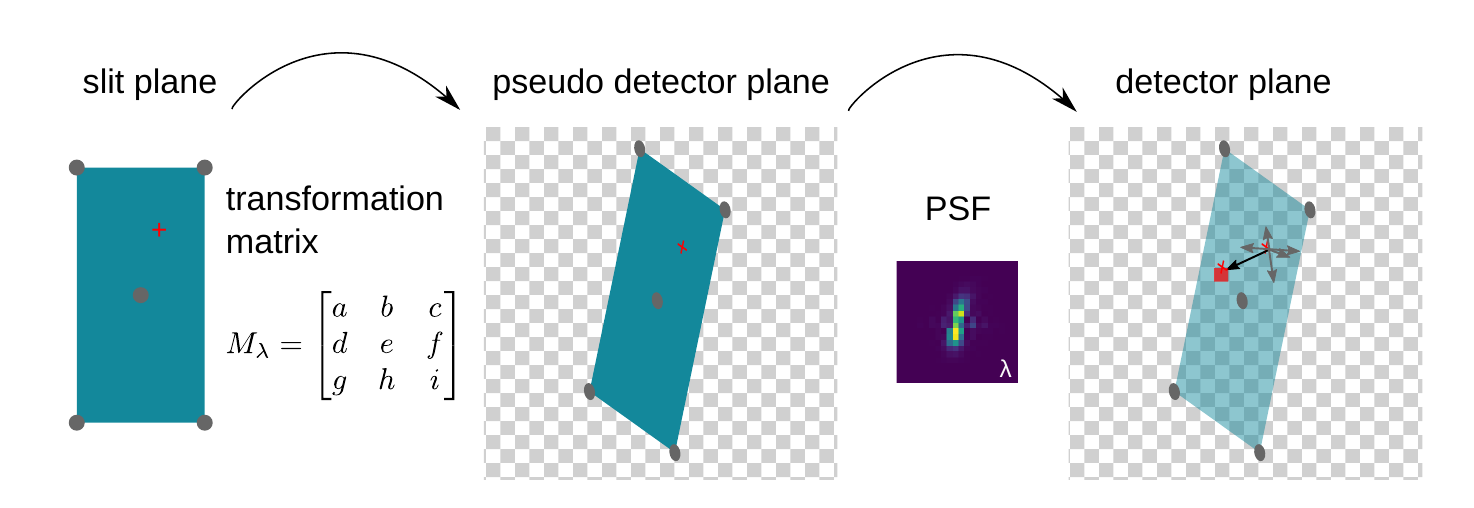
The basic idea is that any spectrograph can be modelled with a set of wavelength-dependent transformation matrices and point spread functions which describe the spectrographs' optics:
First, wavelength-dependent affine transformation matrices are extracted from the ZEMAX model of the spectrograph. As the underlying geometric transformations (scaling, rotation, shearing, translation) vary smoothly across an echelle order, these matrices can be interpolated for any intermediate wavelength.
Second, a wavelength-dependent point spread functions (PSFs) is applied on the transformed slit images to properly account for optical aberrations. Again, the PSF is only slowly varying across an echelle order, allowing for interpolation at intermediate wavelength.
Both, the matrices and the PSFs have to be extracted from ZEMAX only once. It is therefore possible to simulate spectra without access to ZEMAX
Citation
Please cite this paper if you find this work useful in your research.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pyechelle-0.4.0.tar.gz.
File metadata
- Download URL: pyechelle-0.4.0.tar.gz
- Upload date:
- Size: 57.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.6.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2dbcdf041453fdc0571846dabeb7dacb17fef8a61f14572b45f846d467f969f2
|
|
| MD5 |
d3279da3cf97ec7b1b34e7dff50a9b4c
|
|
| BLAKE2b-256 |
d9d051525180e0732a3417ea42da3de32f6809b75f672110d35fe3038d52dd78
|
File details
Details for the file pyechelle-0.4.0-py3-none-any.whl.
File metadata
- Download URL: pyechelle-0.4.0-py3-none-any.whl
- Upload date:
- Size: 64.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.6.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
33c09cb2cf7d89e25edfb4038060d6d05e1ad18903a3701c1ea7855a6a001792
|
|
| MD5 |
fc08cd6540014a4d97764c3048485c51
|
|
| BLAKE2b-256 |
66efae1fadf8b5e625c3125b6883a4959bf6081866afb9efe2f2e3ff29f1aeb3
|