Python client for GBIF
Project description
pygbif
Python client for the GBIF API
Source on GitHub at gbif/pygbif
Other GBIF clients:
R: rgbif, ropensci/rgbif
Ruby: gbifrb, sckott/gbifrb
PHP: php-gbif, restelae/php-gbif
Contributing: CONTRIBUTING.md
Installation
Stable from pypi
pip install pygbifDevelopment version
[sudo] pip install git+git://github.com/gbif/pygbif.git#egg=pygbifpygbif is split up into modules for each of the major groups of API methods.
Registry - Datasets, Nodes, Installations, Networks, Organizations
Species - Taxonomic names
Occurrences - Occurrence data, including the download API
Maps - Maps, get raster maps from GBIF as png or mvt
You can import the entire library, or each module individually as needed.
In addition there is a utils module, currently with one method: wkt_rewind, and a caching method to manage whether HTTP requests are cached or not. See ?pygbif.caching.
Registry module
registry module API:
organizations
nodes
networks
installations
datasets
dataset_metrics
dataset_suggest
dataset_search
Example usage:
from pygbif import registry
registry.dataset_metrics(uuid='3f8a1297-3259-4700-91fc-acc4170b27ce')Species module
species module API:
name_backbone
name_suggest
name_usage
name_lookup
name_parser
Example usage:
from pygbif import species
species.name_suggest(q='Puma concolor')Occurrences module
occurrences module API:
search
get
get_verbatim
get_fragment
count
count_basisofrecord
count_year
count_datasets
count_countries
count_schema
count_publishingcountries
download
download_meta
download_list
download_get
download_citation
download_describe
download_sql
Example usage:
from pygbif import occurrences as occ
occ.search(taxonKey = 3329049)
occ.get(key = 252408386)
occ.count(isGeoreferenced = True)
occ.download('basisOfRecord = PRESERVED_SPECIMEN')
occ.download('taxonKey = 3119195')
occ.download('decimalLatitude > 50')
occ.download_list(user = "sckott", limit = 5)
occ.download_meta(key = "0000099-140929101555934")
occ.download_get("0000066-140928181241064")
occ.download_citation("0002526-241107131044228")
occ.download_describe("simpleCsv")
occ.download_sql("SELECT gbifid,countryCode FROM occurrence WHERE genusKey = 2435098")Maps module
maps module API:
map
Example usage:
from pygbif import maps
out = maps.map(taxonKey = 212, year = 1998, bin = "hex",
hexPerTile = 30, style = "classic-noborder.poly")
out.response
out.path
out.img
out.plot()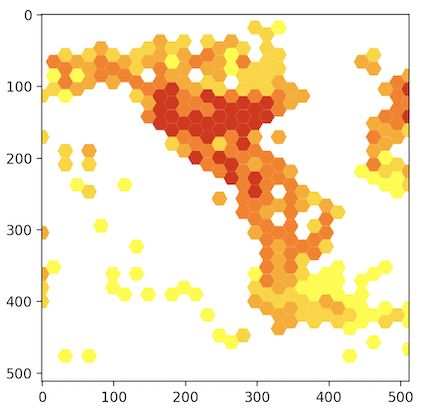
utils module
utils module API:
wkt_rewind
Example usage:
from pygbif import utils
x = 'POLYGON((144.6 13.2, 144.6 13.6, 144.9 13.6, 144.9 13.2, 144.6 13.2))'
utils.wkt_rewind(x)Contributors
Meta
License: MIT, see LICENSE file
Please note that this project is released with a Contributor Code of Conduct. By participating in this project you agree to abide by its terms.
Changelog
0.6.6 (2025-11-14)
updated matching in occurrences.name_backbone to api v2 176
added checklistKey support to occurrences.search 178
added support for collection.search and institution.search
0.6.5 (2024-10-28)
added support for occurrences.download_citation 158
added support for occurrences.download_sql 157
added support for occurrences.download_describe 142
added support for literature.search 149
0.6.4 (2024-03-12)
fixed a bug in building the documentation with readthedocs 138, 129
tests now run against live GBIF APIs 101, 128
updated caching.py since the remove_expired_responses method is deprecated. 126
0.6.3 (2023-05-25)
added support for predicates: isNull, isNotNull, in and not 92, 102 and 103
added support for nested queries/dictionaries 104
deprecated the add_predicate function and added add_pred_dict to accomodate for newly supported predicates to ensure that the arguments that are sent are added in the payload function 108
added support for multiple download formats 105
updated operators and look-up tables 107
included documentation on newly supported predicates and dictionaries 106
0.6.2 (2023-01-24)
update to fix requesting GBIF downloads
minor documentation updates 95 and 99
0.6.1 (2022-06-23)
update to fix broken dependencies 93
minor documentation updates
0.6.0 (2021-07-08)
Fix for occurrences.download when giving geometry as a string rather than using add_geometry; predicates were being split on whitespace, which doesn’t work for WKT 81 84
Moved to using the logging module instead of print() for giving information on occurrence download methods 78
Clarify that occurrences.count for length 1 inputs only; see occurrences.search for > 1 value 75 77
Improved documentation for species.name_usage method, mostly for the language parameter 68
Gains download method download_cancel for cancelling/deleting a download request 59
0.5.0 (2020-09-29)
occurrences.search now supports recordedByID and identifiedByID search parameters 62
clean up the Contributing file, thanks @niconoe 64
clean up internal imports in the library, thanks @niconoe 65
fix usage of is and ==, was using them inappropriately sometimes (via https://realpython.com/python-is-identity-vs-equality/), 69
remove redundant parameter in a doc string, thanks @faroit 71
make a test for internal fxn gbif_GET_write more general to avoid errors if GBIF changes content type response header slightly 72
0.4.0 (2019-11-20)
changed base url to https for all requests; was already https for maps and downloads in previous versions
occurrences, species, and registry modules gain docstrings with brief summary of each method
pygbif gains ability to cache http requests. caching is off by default. See ?pygbif.caching for all the details 52 56 via @nleguillarme
made note in docs that if you are trying to get the same behavior as the GBIF website for name searching, species.name_backbone is likely what you want 55 thanks @qgroom
for parameters that expect a bool, convert them to lowercase strings internally before doing HTTP requests
0.3.0 (2019-01-25)
pygbif is Python 3 only now 19
Gains maps module with maps.map method for working with the GBIF maps API 41 49
Gains new module utils with one method wkt_rewind 46 thanks @aubreymoore for the inspiration
Fixed bug in registry.installations: typo in one of the parameters identifierTyp instead of identifierType 48 thanks @data-biodiversity-aq
Link to GitHub issues from Changelog 🎉
Fix a occurrence download test 47
Much more thorough docs 25
0.2.0 (2016-10-18)
Download methods much improved 16 27 thanks @jlegind @stijnvanhoey @peterdesmet !
MULTIPOLYGON now supported in geometry parameter 35
Fixed docs for occurrences.get, and occurrences.get_verbatim, occurrences.get_fragment and demo that used occurrence keys that no longer exist in GBIF 39
Added organizations method to registry module 12
Added remainder of datasets methods: registry.dataset_search (including faceting support 37) and registry.dataset_suggest, for the /dataset/search and /dataset/suggest routes, respectively 40
Added remainder of species methods: species.name_lookup (including faceting support 38) and species.name_usage, for the /species/search and /species routes, respectively 18
Added more tests to cover new methods
Changed species.name_suggest to give back data stucture as received from GBIF. We used to parse out the classification data, but for simplicity and speed, that is left up to the user now.
start parameter in species.name_suggest, occurrences.download_list, registry.organizations, registry.nodes, registry.networks, and registry.installations, changed to offset to match GBIF API and match usage throughout remainder of pygbif
0.1.5.4 (2016-10-01)
Added many new occurrence.search parameters, including repatriated, kingdomKey, phylumKey, classKey, orderKey, familyKey, genusKey, subgenusKey, establishmentMeans, facet, facetMincount, facetMultiselect, and support for facet paging via **kwargs 30 34
Fixes to **kwargs in occurrence.search so that facet parameters can be parsed correctly and requests GET request options are collected correctly 36
Added spellCheck parameter to occurrence.search that goes along with the q parameter to optionally spell check full text searches 31
0.1.4 (2016-06-04)
Added variable types throughout docs
Changed default limit value to 300 for occurrences.search method
tox now included, via @xrotwang 20
Added more registry methods 11
Started occurrence download methods 16
Added more names methods 18
All requests now send user-agent headers with requests and pygbif versions 13
Bug fix for occurrences.download_get 23
Fixed bad example for occurrences.get 22
Fixed wheel to be universal for 2 and 3 10
Improved documentation a lot, autodoc methods now
0.1.1 (2015-11-03)
Fixed distribution for pypi
0.1.0 (2015-11-02)
First release
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pygbif-0.6.6.tar.gz.
File metadata
- Download URL: pygbif-0.6.6.tar.gz
- Upload date:
- Size: 64.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9b46a249314c7bc575a511428700108404e0459bd08b9789d4eb793ed90bd276
|
|
| MD5 |
9c81fde78f810620deabb63cc1ea471f
|
|
| BLAKE2b-256 |
11ff41bcdaef9ebbabb7d5a1862ac5fd3cf646fe2dc5c73731613e3a9a7cdeb6
|
File details
Details for the file pygbif-0.6.6-py3-none-any.whl.
File metadata
- Download URL: pygbif-0.6.6-py3-none-any.whl
- Upload date:
- Size: 78.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0e46ad6daba9b4f87d49616e2b8934ed2c8ab6fe21aa75c4bc83298819cf9875
|
|
| MD5 |
dcf91191c860be2de4a178ffe062f10c
|
|
| BLAKE2b-256 |
07d719a39b911bd59576b35d9c61564a4ce6c1af1966bced6422a8c656e4cf09
|

















