Tool to preprocess, analyze and visualize DNA methylation data from Illumina micro-arrays
Project description





Tutorials | API documentation | Source code | Release on pip
Pylluminator is a Python package designed to provide an efficient workflow for processing, analyzing, and visualizing DNA methylation data. Pylluminator is inspired from the popular R packages SeSAMe and ChAMP.
Pylluminator supports the following Illumina’s Infinium Beadchip array versions:
human: 27k, 450k, MSA, EPIC, EPIC+, EPICv2
mouse: MM285
mammalian: Mammal40
Main functionalities
idat files parsing
data preprocessing
Type-I probes channel inference
Dye bias correction (3 methods: using normalization control probes / linear scaling / non-linear scaling)
Detection p-value calculation (pOOBAH)
Background correction (NOOB)
Batch effect correction (ComBat)
Missing beta values imputation
Lift over annotation
data analysis and visualisation
beta values (density, PCA, MDS, dendrogram…)
DMPs accounting for replicates / random effects, DMRs
CNV, CNS
pathway analysis with GSEApy (GSEA, ORA)
quality control
Visualization examples:
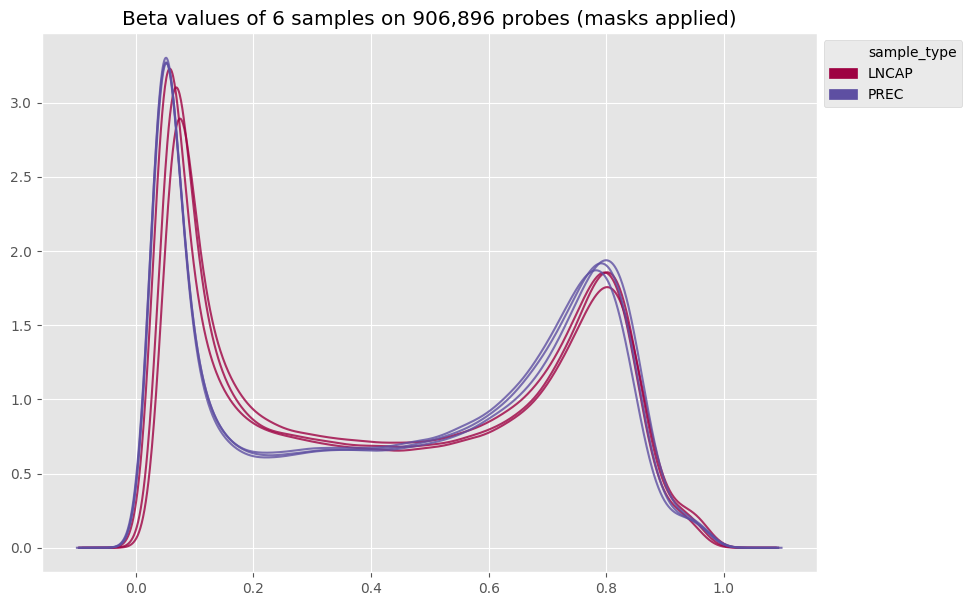
Fig 1. Samples beta values density |
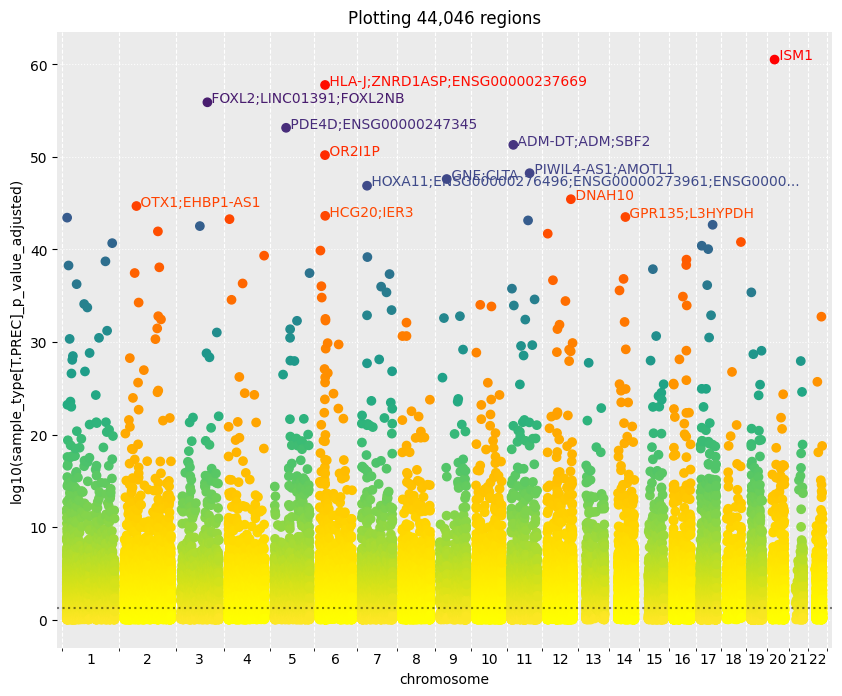
Fig 2. Differentially methylated regions (DMRs) |
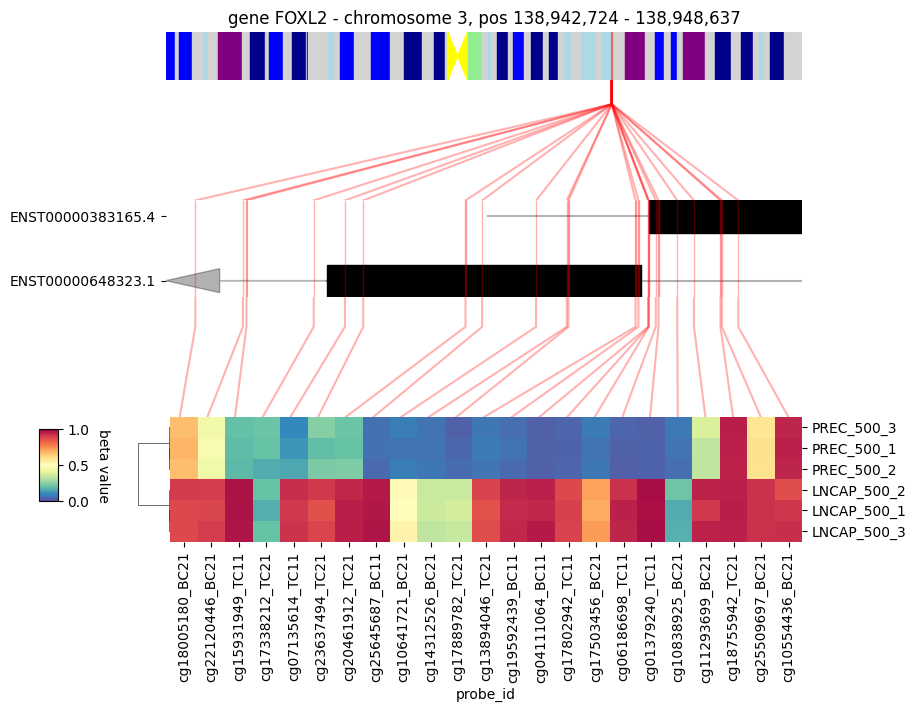
Fig 3. Probes beta values associated with a specific gene |
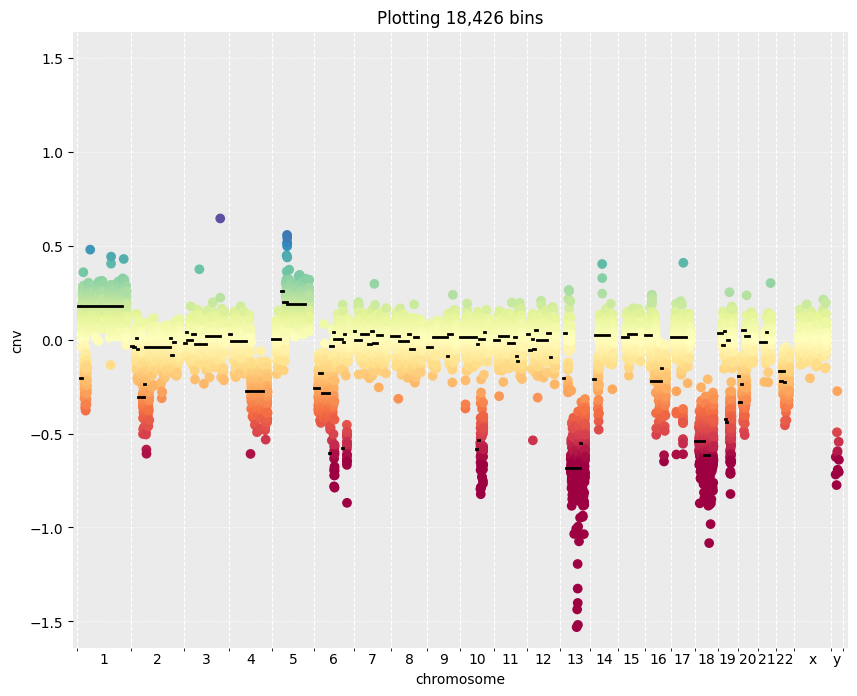
Fig 4. Copy number variations (CNVs) |
Installation
With uv (recommended)
uv is a fast Python package manager. If you don’t have it yet, install it with:
curl -LsSf https://astral.sh/uv/install.sh | shThen install Pylluminator into a uv-managed project:
uv add pylluminatorOr with the optional GSEA extras:
uv add "pylluminator[gsea]"With pip
You can install Pylluminator directly with:
pip install pylluminatorOr, if you want to use the GSEA functionalities, install the additional dependencies with:
pip install pylluminator[gsea]From source
We recommend using uv to build pylluminator from source. The project requires Python 3.12 or later.
Install uv (if needed)
curl -LsSf https://astral.sh/uv/install.sh | shClone and install
git clone https://github.com/eliopato/pylluminator.git
cd pylluminator
uv syncThis creates a virtual environment and installs all dependencies automatically. To include optional extras:
uv sync --extra gsea
uv sync --extra dev
uv sync --extra docsRun scripts or tests within the project environment using uv run:
uv run pytestUsage
Refer to https://pylluminator.readthedocs.io/ for step-by-step tutorials and detailed documentation.
Citing
Pylluminator is described in detail in: Pylluminator: fast and scalable analysis of DNA methylation data in Python, available on BioRxiv
If you use this package in your research, please cite our work.
If you use the updated version of the EPICv2/hg38 annotations, please cite Re-annotating the EPICv2 manifest with genes, intragenic features, and regulatory elements, (BioRxiv link)
Contributing
We welcome contributions! If you’d like to help improve the package, please follow these steps:
Fork the repository.
Create a new branch for your feature or bugfix.
Make your changes and test them.
Submit a pull request describing your changes.
The packages used for development (testing, packaging and building the documentation) can be installed with:
uv sync --extra dev --extra docsBug reports / new features suggestion
If you encounter any bugs, have questions, or feel like the package is missing a very important feature, please open an issue on the GitHub Issues page.
When opening an issue, please provide as much detail as possible, including:
Steps to reproduce the issue
The version of the package you are using
Any relevant code snippets or error messages
License
This project is licensed under the MIT License - see the LICENSE file for details.
Acknowledgements
This package is strongly inspired from SeSAMe and includes code from methylprep for .idat files parsing.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pylluminator-2.2.tar.gz.
File metadata
- Download URL: pylluminator-2.2.tar.gz
- Upload date:
- Size: 86.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.10.12 {"installer":{"name":"uv","version":"0.10.12","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"Ubuntu","version":"24.04","id":"noble","libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
41733147233aa2930f32d56f755c84af35d6ba54d1a9e45db39f1a13d5776979
|
|
| MD5 |
c87789e8a490540291a9c2b3938fd6e5
|
|
| BLAKE2b-256 |
de35415326ce2f914dca611769fcc4f1f963e92130034fe7b9f0702c942b7f2e
|
File details
Details for the file pylluminator-2.2-py3-none-any.whl.
File metadata
- Download URL: pylluminator-2.2-py3-none-any.whl
- Upload date:
- Size: 92.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.10.12 {"installer":{"name":"uv","version":"0.10.12","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"Ubuntu","version":"24.04","id":"noble","libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":null}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
46ee4f246a37219fe1dff2f988f95bd8e174f969b782a8be82f2fbcca499a0a4
|
|
| MD5 |
eb1039e249a5655914cac876e5472126
|
|
| BLAKE2b-256 |
1ffd1767f7cba15e094e501bc5e12c85058247a7834fa52952b03a4baf7b9c50
|












