A Pyrodigal extension to predict genes in RNA viruses with the standard and alternative genetic code.
Project description
🔥🦠 Pyrodigal-rv 
A Pyrodigal extension to predict genes in RNA viruses (with standard and alternative genetic code).
🗺️ Overview
Pyrodigal is a Python module that provides
Cython bindings to Prodigal,
an efficient gene finding method for genomes and metagenomes based on dynamic programming.
Additionally, pyrodigal-gv is a small extension module for pyrodigal (both written by Martin Larralde) which distributes additional metagenomic models for giant viruses and viruses that use alternative genetic codes, first provided by Antônio Camargo in prodigal-gv.
Inspired by the additional metagenomic models for giant viruses and bacteriophages in pyrodigal-gv, pyrodigal-rv substitutes those metagenomic models and the bacterial models from pyrodigal for metagenomic models from RNA viruses which mostly use the standard genetic code (translation table 1), but also include RNA virus models with alternative genetic codes.
See below for which viral families and which genetic codes are included. The process of model generation is documented in a separate repo.
Code and instructions below are exactly the same as for pyrodigal-gv.
🔧 Installing
pyrodigal-rv can be installed directly from PyPI
as a universal wheel that contains all required data files:
$ pip install pyrodigal-rv
💡 Example
Just use the provided ViralGeneFinder class instead of the usual GeneFinder
from pyrodigal, and the new viral models will be used automatically in
meta mode:
import Bio.SeqIO
import pyrodigal_rv
record = Bio.SeqIO.read("sequence.gbk", "genbank")
orf_finder = pyrodigal_rv.ViralGeneFinder(meta=True)
for i, pred in enumerate(orf_finder.find_genes(bytes(record.seq))):
print(f">{record.id}_{i+1}")
print(pred.translate())
ViralGeneFinder has an additional keyword argument, viral_only, which can
be set to True to run gene calling using only viral models.
🔨 Command line
pyrodigal-rv comes with a very simple command line similar to Prodigal and pyrodigal:
$ pyrodigal-rv -i <input_file.fasta> -a <gene_translations.fasta> -d <gene_sequences.fasta>
Contrary to prodigal and pyrodigal, the pyrodigal-rv script runs in meta mode
by default! Running in single mode can be done with pyrodigal-rv -p single but
the results will be exactly the same as pyrodigal, so why would you ever do this ⁉️
📊 Benchmarking
The benchmarking is documented in this repo.
Accuracy
To evaluate pyrodigal-rv ORF prediction in RNA viruses, all Riboviria sequences in RefSeq indicated as "complete" by the sequence submission authors and without N's in the sequence were used as a benchmark (n=9,001).
All tools were run in closed mode (-c) and pyrodigal was forced to use genetic code 1 (-g 1) for the benchmarking as this is the most used genetic code by RNA viruses.
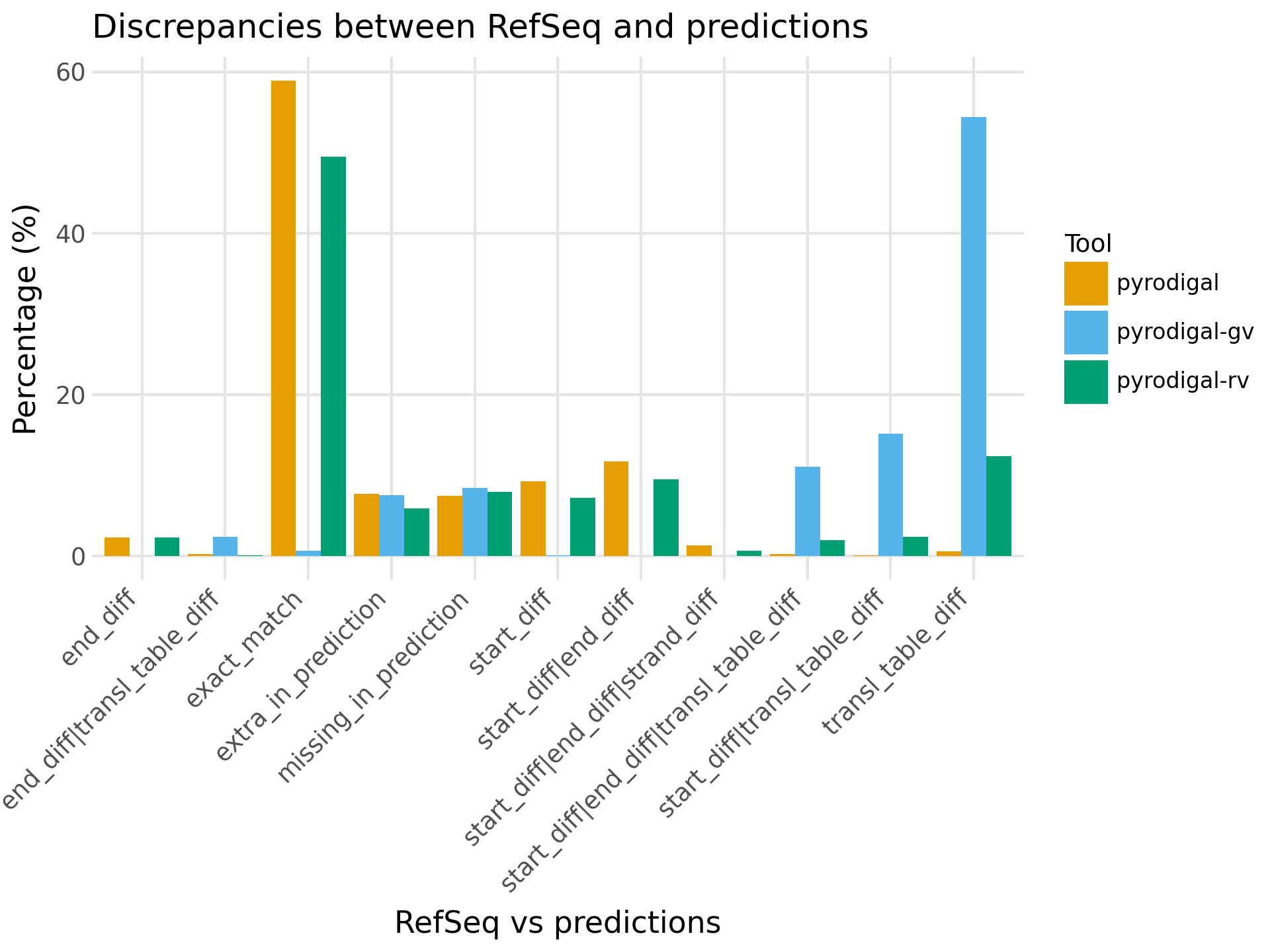
After comparison with the CDS annotations from RefSeq, pyrodigal and pyrodigal-rv give 58.9% and 49.4% exact matches respectively, while both of them also predicted ~25% CDSs with different start and/or stop sites compared to RefSeq.
For pyrodigal-rv another 12.4% was predicted to only have a different translation table.
As expected pyrodigal-gv had almost no exact matches because it contains no metagenomic models with genetic code 1, and it also predicts 28.8% CDSs with different start/stop sites (4.6% higher than pyrodigal-rv).
pyrodigal-rv also performed best in context of extra and missing CDS predictions (considerably lower amount extra predictions and only 0.4% more missing predictions compared to pyrodigal).
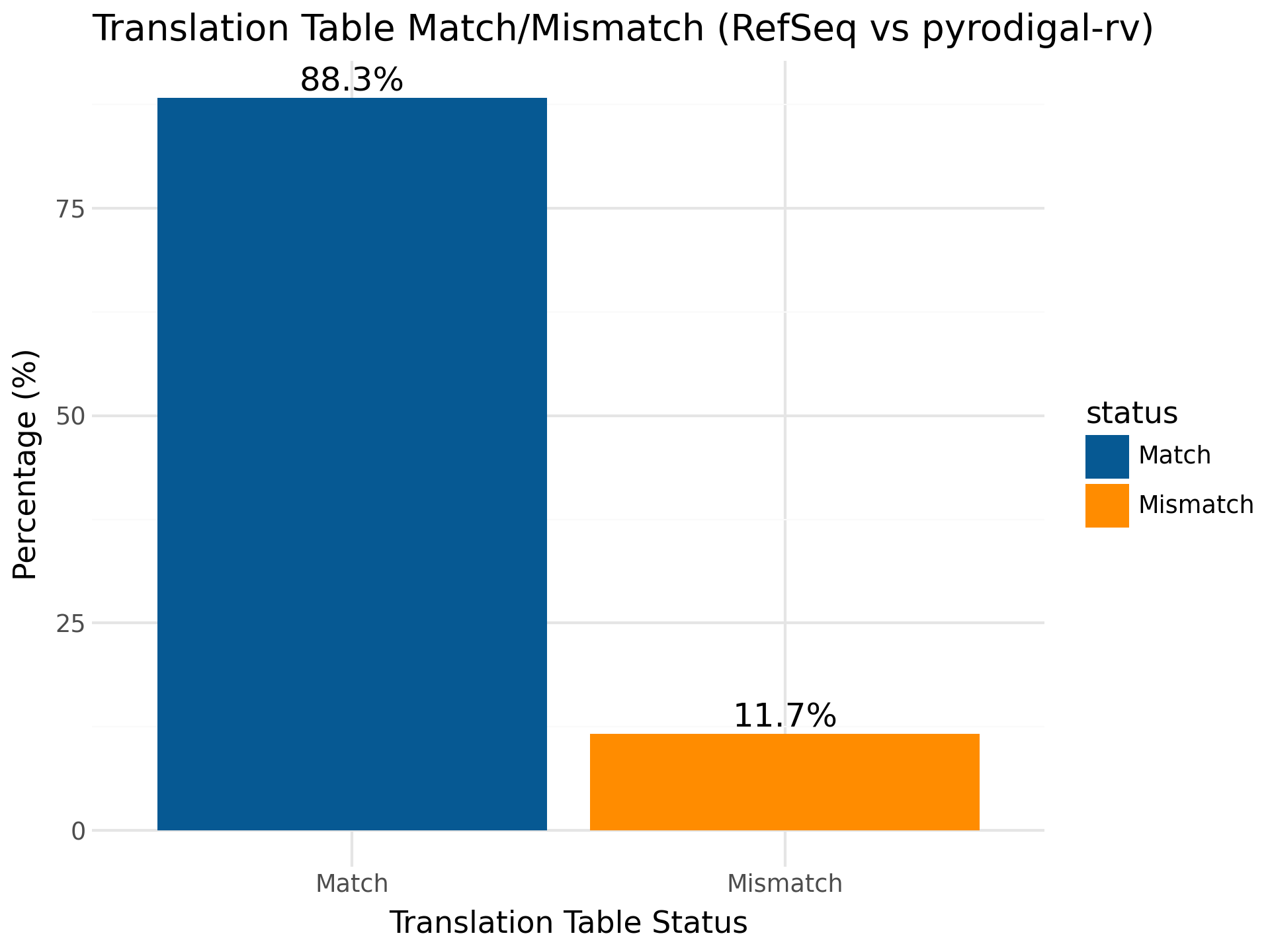
pyrodigal-rv adds the ability to predict the right genetic code for your RNA virus sequence, when comparing to RefSeq, 11.7% of the sequences had a mismatch in genetic code.
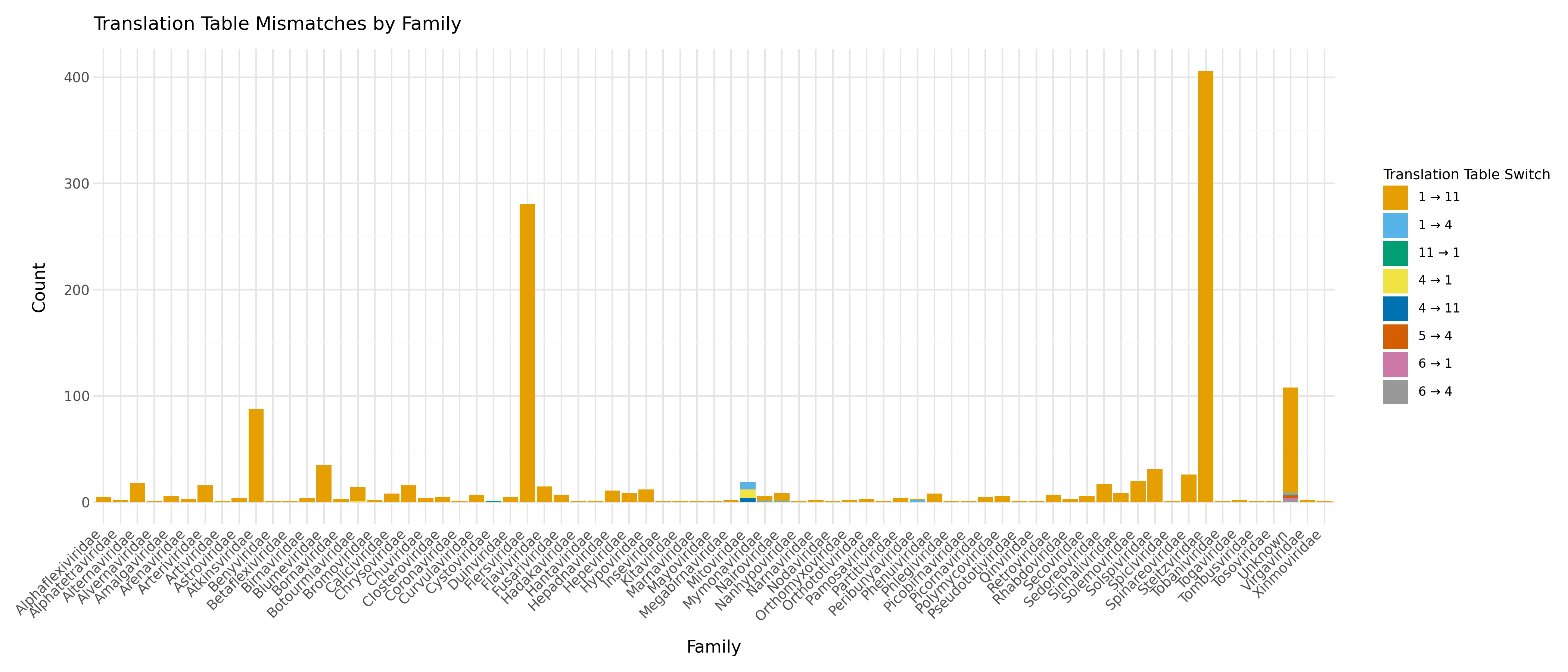
However, when examining more closely the majority of these sequences belong to the Atkinsviridae, Blumeviridae, Fiersviridae, Solspiviridae and Steitzviridae, which are RNA phages and should use the bacterial genetic code 11 (as predicted by pyrodigal-rv).
This shows that not all sequences in RefSeq are annotated with the correct translation table and this benchmark underestimated pyrodigal-rv's accuracy in number of exact matches.
Disclaimer: The training models for pyrodigal-rv contain some RefSeq sequences.
Speed
CLI speed was benchmarked with hyperfine over 10 runs of the same command on 9,000 sequences for each CLI (pyrodigal, pyrodigal-gv and pyrodigal-rv) using 10 processes (-j 10 --pool process).
| Command | Mean [s] | Min [s] | Max [s] | Relative |
|---|---|---|---|---|
pyrodigal |
63.883 ± 0.597 | 63.402 | 65.288 | 2.19 ± 0.03 |
pyrodigal-gv |
29.568 ± 0.563 | 28.250 | 30.286 | 1.01 ± 0.02 |
pyrodigal-rv |
29.150 ± 0.199 | 28.860 | 29.540 | 1.00 |
🔖 Citation
Pyrodigal is scientific software, with a published paper in the Journal of Open-Source Software. Please cite both Pyrodigal and Prodigal if you are using it in an academic work, for instance as:
Pyrodigal (Larralde, 2022), a Python library binding to Prodigal (Hyatt et al., 2010).
Detailed references are available on the Publications page of the online documentation.
💭 Feedback
⚠️ Issue Tracker
Found a bug ? Have an enhancement request ? Head over to the GitHub issue tracker if you need to report or ask something. If you are filing in on a bug, please include as much information as you can about the issue, and try to recreate the same bug in a simple, easily reproducible situation.
🏗️ Contributing
Contributions are more than welcome! See
CONTRIBUTING.md
for more details.
📋 Changelog
This project adheres to Semantic Versioning and provides a changelog in the Keep a Changelog format.
⚖️ License
This library is provided under the GNU General Public License v3.0.
The Prodigal code was written by Doug Hyatt and is distributed under the
terms of the GPLv3 as well. See vendor/Prodigal/LICENSE for more information.
This project is in no way affiliated, sponsored, or otherwise endorsed by the original Prodigal authors. It was developed by Martin Larralde during his PhD project at the European Molecular Biology Laboratory in the Zeller team. RNA virus models were added by Lander De Coninck.
📐 Models
Click to see included models and genetic codes:
| model | parent_family | name | viral | gc_content | genetic_code | uses_sd |
|---|---|---|---|---|---|---|
| 1 | Tymoviridae_1_model | V | 54.5 | 1 | 0 | |
| 2 | Picobirnaviridae_6_model | V | 43.5 | 6 | 0 | |
| 3 | Polymycoviridae_1_model | V | 57.7 | 1 | 0 | |
| 4 | Atkinsviridae_11_model | V | 49.0 | 11 | 1 | |
| 5 | Duinviridae_11_model | V | 43.6 | 11 | 1 | |
| 6 | Aspiviridae_1_model | V | 36.0 | 1 | 0 | |
| 7 | Narnaviridae_1_model | V | 50.5 | 1 | 0 | |
| 8 | Peribunyaviridae_1_model | V | 35.9 | 1 | 0 | |
| 9 | Nodaviridae_1_model | V | 49.5 | 1 | 0 | |
| 10 | Sedoreoviridae_1_model | V | 37.6 | 1 | 0 | |
| 11 | Narnaviridae_6_model | V | 51.3 | 6 | 0 | |
| 12 | Qinviridae_1_model | V | 46.6 | 1 | 0 | |
| 13 | Narnaviridae_4_model | V | 41.3 | 4 | 1 | |
| 14 | Tombusviridae_6_model | V | 51.5 | 6 | 1 | |
| 15 | Orthototiviridae_1_model | V | 48.3 | 1 | 0 | |
| 16 | Tombusviridae_16_model | V | 53.1 | 16 | 1 | |
| 17 | Carmotetraviridae_1_model | V | 50.7 | 1 | 0 | |
| 18 | Steitzviridae_11_model | V | 50.4 | 11 | 1 | |
| 19 | Picobirnaviridae_1_model | V | 42.0 | 1 | 1 | |
| 20 | Dicistroviridae_1_model | V | 40.7 | 1 | 0 | |
| 21 | Astroviridae_1_model | V | 45.9 | 1 | 0 | |
| 22 | Hepadnaviridae_1_model | V | 47.4 | 1 | 0 | |
| 23 | Tombusviridae_1_model | V | 49.8 | 1 | 0 | |
| 24 | Solspiviridae_11_model | V | 49.9 | 11 | 1 | |
| 25 | Cystoviridae_11_model | V | 51.3 | 11 | 1 | |
| 26 | Picobirnaviridae_5_model | V | 36.2 | 5 | 0 | |
| 27 | Blumeviridae_11_model | V | 45.2 | 11 | 1 | |
| 28 | Alphaormycoviridae_1_model | V | 44.6 | 1 | 0 | |
| 29 | Orthomyxoviridae_1_model | V | 40.0 | 1 | 0 | |
| 30 | Fiersviridae_4_model | V | 49.2 | 4 | 1 | |
| 31 | Flaviviridae_1_model | V | 42.1 | 1 | 0 | |
| 32 | Splipalmiviridae_1_model | V | 49.3 | 1 | 0 | |
| 33 | Picobirnaviridae_4_model | V | 43.0 | 4 | 0 | |
| 34 | Betaormycoviridae_1_model | V | 41.6 | 1 | 0 | |
| 35 | Tombusviridae_4_model | V | 48.4 | 4 | 0 | |
| 36 | Pseudototiviridae_1_model | V | 55.2 | 1 | 0 | |
| 37 | Fiersviridae_6_model | V | 48.7 | 6 | 0 | |
| 38 | Fimoviridae_1_model | V | 31.0 | 1 | 0 | |
| 39 | Botourmiaviridae_4_model | V | 44.8 | 4 | 1 | |
| 40 | Fiersviridae_11_model | V | 50.1 | 11 | 1 | |
| 41 | Yueviridae_1_model | V | 41.3 | 1 | 0 | |
| 42 | Dicistroviridae_6_model | V | 35.9 | 6 | 1 | |
| 43 | Spinareoviridae_1_model | V | 43.6 | 1 | 0 | |
| 44 | Matonaviridae_1_model | V | 61.3 | 1 | 0 | |
| 45 | Picornaviridae_1_model | V | 44.2 | 1 | 0 | |
| 46 | Caulimoviridae_1_model | V | 40.7 | 1 | 0 | |
| 47 | Barnaviridae_1_model | V | 50.7 | 1 | 0 | |
| 48 | Chrysoviridae_1_model | V | 49.1 | 1 | 0 | |
| 49 | Mitoviridae_16_model | V | 44.1 | 16 | 0 | |
| 50 | Picornaviridae_4_model | V | 48.3 | 4 | 0 | |
| 51 | Picornaviridae_6_model | V | 43.0 | 6 | 0 | |
| 52 | Partitiviridae_1_model | V | 44.9 | 1 | 0 | |
| 53 | Qinviridae_6_model | V | 47.8 | 6 | 1 | |
| 54 | Botourmiaviridae_1_model | V | 51.6 | 1 | 0 | |
| 55 | Potyviridae_1_model | V | 41.9 | 1 | 0 | |
| 56 | Fiersviridae_16_model | V | 50.0 | 16 | 0 | |
| 57 | Yadokariviridae_1_model | V | 45.6 | 1 | 0 | |
| 58 | Narnaviridae_16_model | V | 45.3 | 16 | 1 | |
| 59 | Flaviviridae | Pestivirus_1_model | V | 45.0 | 1 | 0 |
| 60 | Flaviviridae | Pegivirus_1_model | V | 55.2 | 1 | 0 |
| 61 | Mitoviridae | Unuamitovirus_4_model | V | 35.9 | 4 | 0 |
| 62 | Flaviviridae | Hepacivirus_1_model | V | 55.0 | 1 | 0 |
| 63 | Mitoviridae | Duamitovirus_4_model | V | 41.4 | 4 | 0 |
| 64 | Mitoviridae | Triamitovirus_4_model | V | 39.9 | 4 | 0 |
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pyrodigal_rv-0.1.0.tar.gz.
File metadata
- Download URL: pyrodigal_rv-0.1.0.tar.gz
- Upload date:
- Size: 1.8 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3ccfbdeadda103cd2f17d3d753acb280a2fe35411ac166cd08cdecf317f64d88
|
|
| MD5 |
b1035169474eb21283739ef4569fbe89
|
|
| BLAKE2b-256 |
173755900c64b0db57d3b9ec269de81641b0f885ed7b989efcece575dc89be5e
|
Provenance
The following attestation bundles were made for pyrodigal_rv-0.1.0.tar.gz:
Publisher:
package.yml on LanderDC/pyrodigal-rv
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
pyrodigal_rv-0.1.0.tar.gz -
Subject digest:
3ccfbdeadda103cd2f17d3d753acb280a2fe35411ac166cd08cdecf317f64d88 - Sigstore transparency entry: 725419871
- Sigstore integration time:
-
Permalink:
LanderDC/pyrodigal-rv@f0ee9a067a54a8b402bd5cf4952b6aec719d2435 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/LanderDC
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
package.yml@f0ee9a067a54a8b402bd5cf4952b6aec719d2435 -
Trigger Event:
push
-
Statement type:
File details
Details for the file pyrodigal_rv-0.1.0-py2.py3-none-any.whl.
File metadata
- Download URL: pyrodigal_rv-0.1.0-py2.py3-none-any.whl
- Upload date:
- Size: 1.8 MB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0f852ece25d2c66fd68f326df374ef29db5db7113c3003d370d0d702efbb73a7
|
|
| MD5 |
6af320aecc228c285869c07e18b54279
|
|
| BLAKE2b-256 |
e3ee9f693b001aaa5b262c2603954190d9d23092edb4ad44be474d7413f1afe8
|
Provenance
The following attestation bundles were made for pyrodigal_rv-0.1.0-py2.py3-none-any.whl:
Publisher:
package.yml on LanderDC/pyrodigal-rv
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
pyrodigal_rv-0.1.0-py2.py3-none-any.whl -
Subject digest:
0f852ece25d2c66fd68f326df374ef29db5db7113c3003d370d0d702efbb73a7 - Sigstore transparency entry: 725419873
- Sigstore integration time:
-
Permalink:
LanderDC/pyrodigal-rv@f0ee9a067a54a8b402bd5cf4952b6aec719d2435 -
Branch / Tag:
refs/tags/v0.1.0 - Owner: https://github.com/LanderDC
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
package.yml@f0ee9a067a54a8b402bd5cf4952b6aec719d2435 -
Trigger Event:
push
-
Statement type:





















