A semantic reasoning and graph analysis engine for biomedical concepts using the UMLS.
Project description
PyUMLS-Similarity
PyUMLS-Similarity is a semantic similarity engine for biomedical concept analysis, built on the UMLS. It supports transformer-based embeddings, various similarity metrics (e.g. path, LCH, Resnik, Lin), and interactive exploration of concept relationships. It aims to facilitate the integration of these methods into existing NLP workflows.
Features
UMLS Integration
- Connects to local UMLS MySQL databases (2024AB and earlier).
- Parses MRCONSO, MRREL, MRSTY, MRDEF, MRHIER; validates schema and vocabularies.
Graph Construction
- Builds directed/undirected graphs from MeSH or SNOMED using
PAR,RB,RN,CHD, andis_a. - Visualizes paths with PyVis; optional Neo4j export for interactive exploration.
Semantic Similarity
- Implements path-based metrics: Path Length, Wu-Palmer, LCH.
- Supports IC-based metrics: Resnik, Lin, Jiang-Conrath.
- Adds embedding-based methods: Word2Vec, GloVe, Transformer (e.g., PubMedBERT).
- Outputs can be normalized and compatible with heatmap comparison.
Concept Linking & LCS
- Resolves CUIs from terms or accepts direct input.
- Computes shortest paths and lowest common subsumers.
- Extracts STRs, definitions, synonyms, semantic types/groups.
NER and Free Text Input
- Runs NER with Hugging Face models (e.g.,
dslim/bert-base-NER). - Links extracted spans to UMLS CUIs and computes similarity between all pairs.
Output & Visualization
- Saves results to CSV, Excel, or Parquet.
- Plots similarity heatmaps across selected metrics.
- Renders interactive graphs with hoverable concept details.
Requirements and Installation
Installation
You can install pyumls-similarity via pip:
pip install pyumls-similarity
You must also have access to a local UMLS installation and corresponding MySQL database. Additional setup steps are outlined below.
UMLS Setup Requirements
To use this package, you must have a MySQL instance of the UMLS configured in your environment. You can get a UMLS License at: https://www.nlm.nih.gov/research/umls/index.html
You can install the UMLS following the instructions here: https://www.nlm.nih.gov/research/umls/implementation_resources/metamorphosys/help.html
It is recommended that you also follow modify your my.ini file for MySQL using the parameters here for optimal performance (https://www.nlm.nih.gov/research/umls/implementation_resources/scripts/README_RRF_MySQL_Output_Stream.html)
Optional: Neo4j Integration
To explore UMLS concept graphs visually and semantically:
Install Neo4j and run a graph database locally or in the cloud. The engine supports exporting UMLS graphs to Neo4j (AuraDB) with rich concept metadata and relationship types.
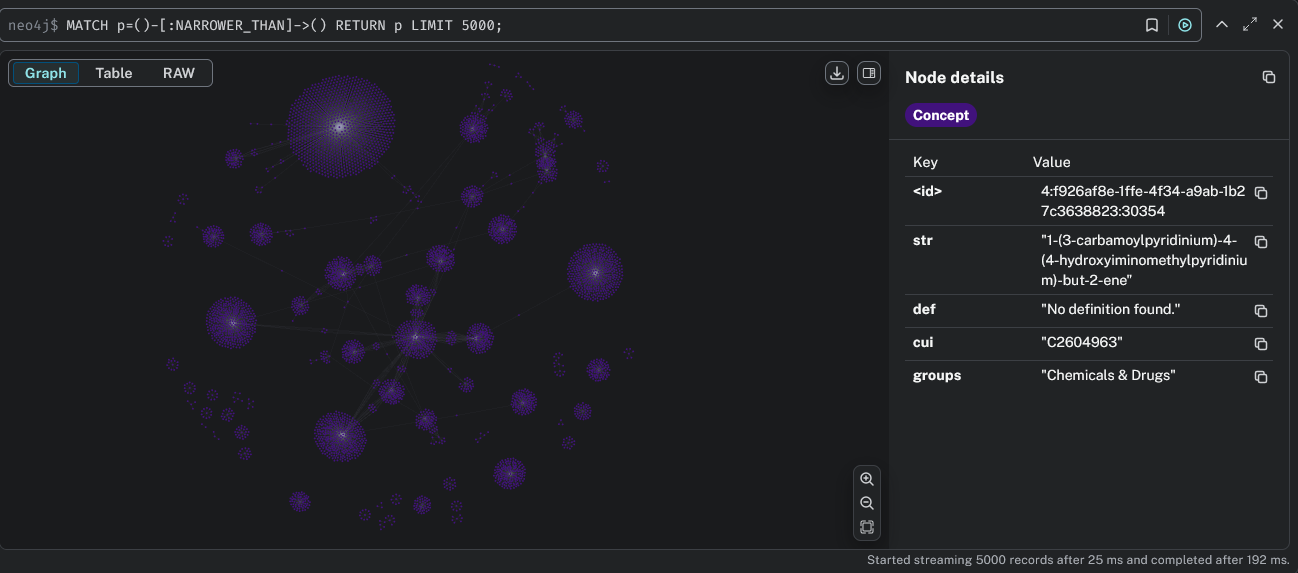
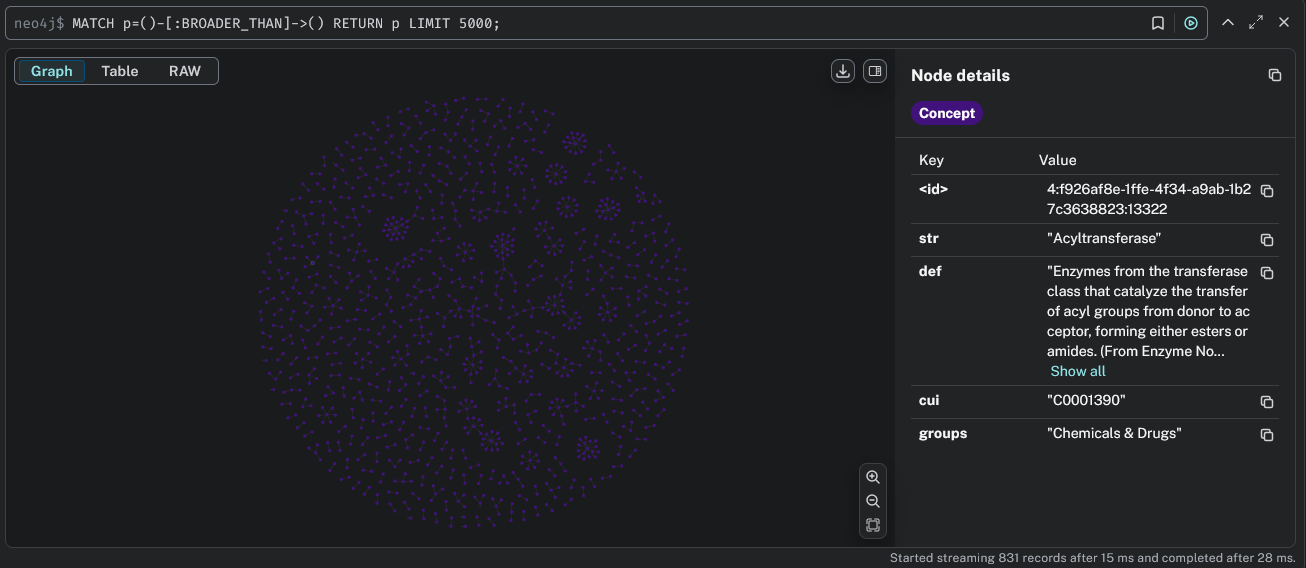
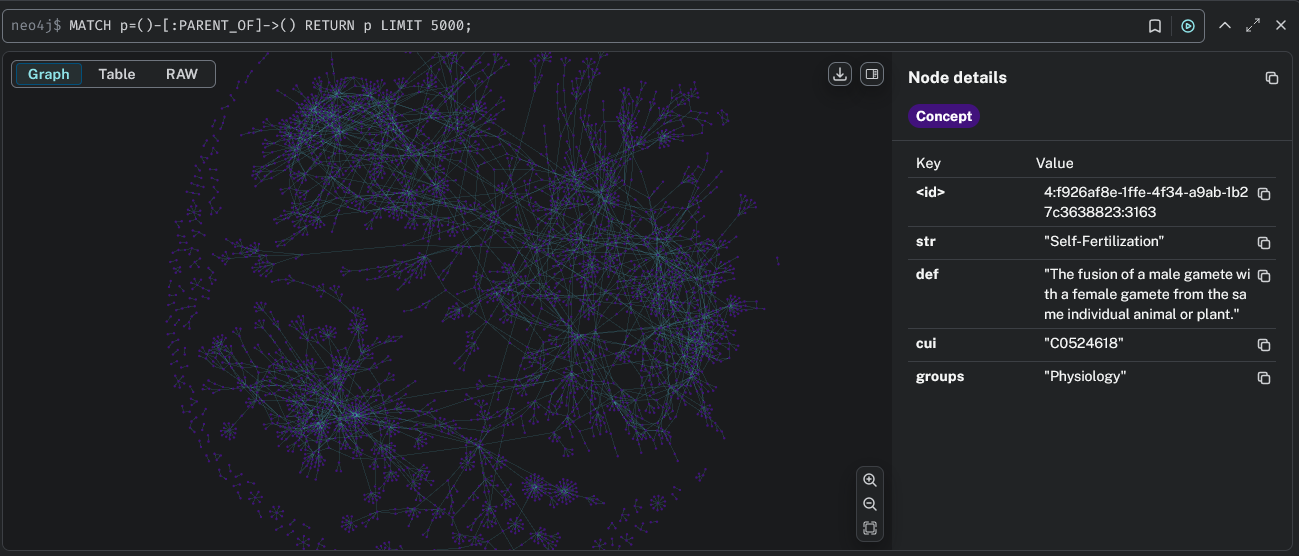
Required Files
Semantic Group Mapping
You will need the SemGroups.txt file, available at:
https://www.nlm.nih.gov/research/umls/knowledge_sources/semantic_network/SemGroups.txt
Information Content File
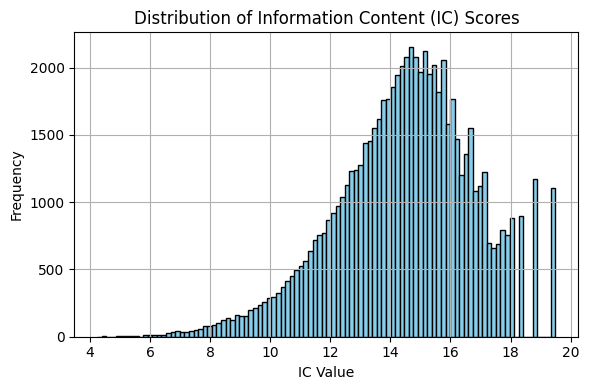
To use IC-based similarity measures (Resnik, Lin, Jiang-Conrath), you must provide an information content (IC) file mapping CUIs to IC values.
A precomputed IC file (cui_ic_pubmed.parquet) is included with the package based on ~72,000 CUIs using term frequency estimates from a large PubMed abstracts corpus: yanjx21/PubMed on Hugging Face.
Note: Information Content-based measures are dependent on the corpus used to determine term frequencies. As such, you may want to explore different corpora when exploring these metrics. The file provided here is for prototyping/experimentation purposes.
Basic Usage
Start by importing the module like so:
from pyumls-similarity import UMLSSemanticEngine
Then, you can use the routines below to get you started. Start by initiating the connection to the MySQL UMLS database. Load the SemGroups.txt file.
You can also optionally load the transformer model you want to use for your embeddings and the cui_ic.parquet file if you are interested in information content-based metrics
# Step 1: Set up MySQL connection info
mysql_info = {
"host": "localhost",
"user": "root",
"password": "your_password",
"database": "umls2024"
}
# Step 2: Instantiate the engine and load resources
umls_utils = UMLSSemanticEngine(mysql_info)
umls_utils.load_transformer_model(model_name="NeuML/pubmedbert-base-embeddings")
tui_to_group = umls_utils.load_semantic_group_mapping("SemGroups.txt")
umls_utils.load_cui_ic_from_parquet("ic/cui_ic_pubmed.parquet")
Next, you'll want to construct the networks for whatever ontology you are interested in. In the example below, I'm creating the graphs for the MeSH ontology using PAR relationships (in G_MSH) and including RB and RN relationships (in G_MSH_expanded).
# Step 3: Build graphs from UMLS MRREL
G_MSH = umls_utils.build_mesh_graph_from_mrrel()
UG_MSH = G_MSH.to_undirected()
G_MSH_expanded = umls_utils.build_mesh_graph_expanded()
UG_MSH_expanded = G_MSH_expanded.to_undirected()
Then, there are different inputs that can be used for the semantic calculations. You can either provide a list of tuples containing the concept pairs that you want to compare or you can provide a piece of text. Below is an example of a workflow using the tuple list.
# Step 4: Define concept pairs
concept_pairs = [
('Heart', 'Lung'),
('Kidney', 'Liver'),
('Asthma', 'Influenza'),
('Diabetes Mellitus', 'Hypertension'),
('Stroke', 'Alzheimer Disease'),
('Breast', 'Prostate'),
('Blood', 'Platelets'),
('Vaccination', 'Antibody Formation'),
('MRI', 'Computed Tomography'),
('Anxiety', 'Depression'),
('Amoxicillin', 'Vancomycin'),
('Surgery', 'Wound Healing'),
('Fluoxetine', 'PTSD'),
('Breast Cancer', 'Prostate Cancer'),
]
# Step 5: Run semantic similarity analysis
df_results = umls_utils.generate_concept_path_analysis(
concept_pairs=concept_pairs,
UG=UG_MSH,
UG_expanded=UG_MSH_expanded,
tui_to_group=tui_to_group,
sab="MSH",
max_depth=18,
similarity_metrics=["path", "wup", "lch", "transformer", "resnik", "lin"],
verbose=False
)
To process a piece of text you can do so similarly as shown below:
# Step 4: Load your NER model from HuggingFace
umls_utils.load_ner_model(model_name="HUMADEX/english_medical_ner")
# Step 5: Provide your piece of text
sample_text = """
The patient presented with symptoms of both asthma and influenza, experiencing shortness of breath and fever.
Prior history includes diabetes mellitus and hypertension, managed through insulin therapy and dietary changes.
Imaging techniques such as MRI and computed tomography were utilized to assess internal damage. Elevated blood pressure was noted alongside low platelet count.
A family history of breast cancer and prostate cancer was also reported. The patient received a vaccination and demonstrated antibody formation within two weeks.
Mood assessments revealed moderate anxiety and depression. A surgical plan was developed to accelerate wound healing post-intervention."
"""
# Step 6: Run semantic similarity analysis
df_results_ner = umls_utils.run_pipeline_on_text(
sample_text,
UG_MSH_expanded,
tui_to_group,
sab="MSH",
verbose=True,
similarity_metrics=["path", "wup", "lch"]
)
Regardless of the method, you'll end up with a dataframe that provides detailed information about each concept pair like their CUI, their STR value in the ontology, their DEF in the ontology, the exact path between them (if it exists), their lowest common subsumer, semantic groups, and several others as shown below.
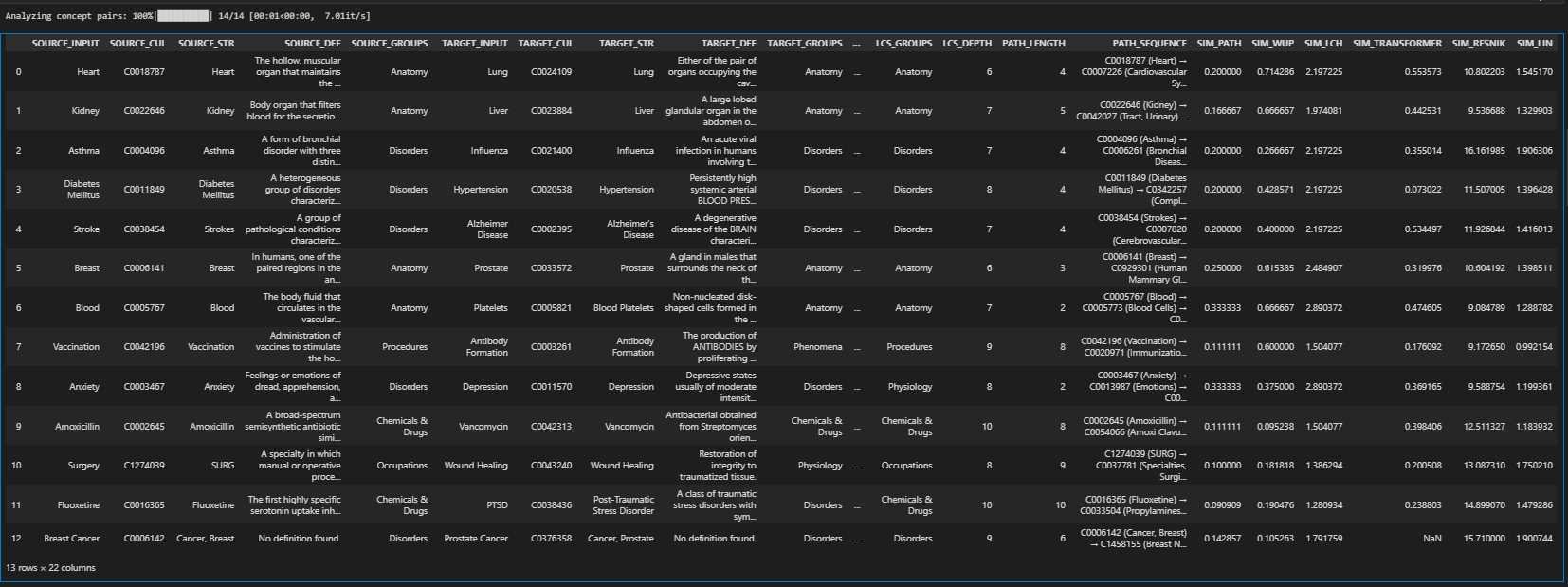
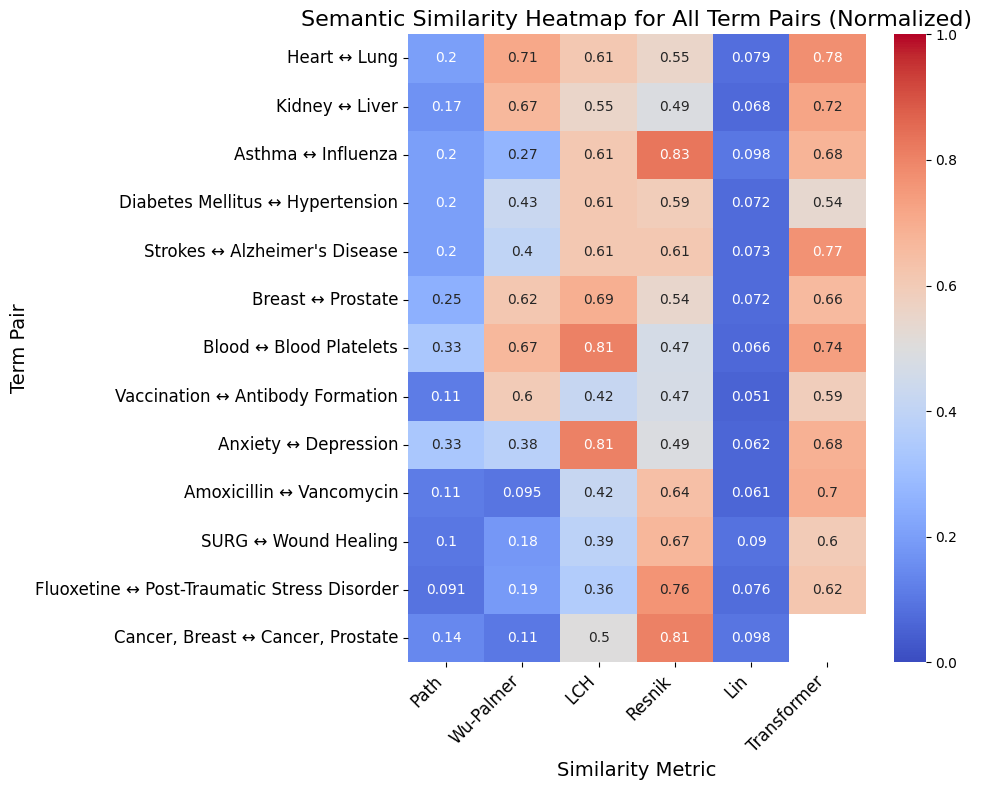
You can also visualize the similarity metrics via a heatmap using:
umls_utils.plot_all_similarity_metrics_heatmap(df_results, max_depth=18, normalize=True)
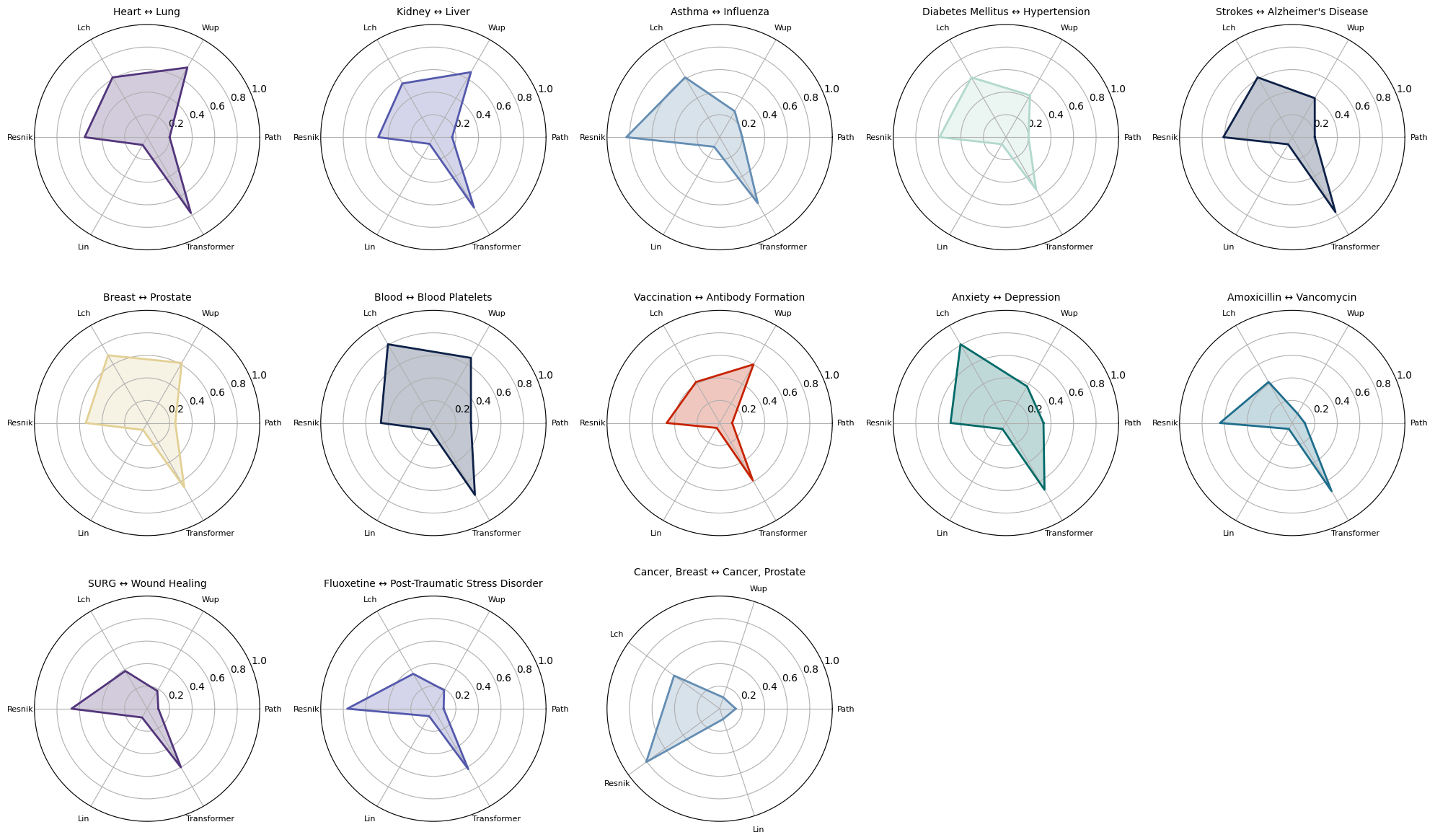
Or you can also visualize radar plots for the concept pairs as shown here:
umls_utils.plot_all_similarity_metrics_radar(df_results, max_depth=18, normalize=True, plots_per_row=5)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file pyumls_similarity-0.1.3.tar.gz.
File metadata
- Download URL: pyumls_similarity-0.1.3.tar.gz
- Upload date:
- Size: 29.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
527ee6a7de304dfeae932e1dec6bec069e2ec0d9198ce253b6d79243f6dc816b
|
|
| MD5 |
3b9546d1284d348c33d83328c1845944
|
|
| BLAKE2b-256 |
fce385e64ced4f3a6dfd4dff6b588aa6462446b4a3e8a4d552936c1aa8576df0
|
File details
Details for the file pyumls_similarity-0.1.3-py3-none-any.whl.
File metadata
- Download URL: pyumls_similarity-0.1.3-py3-none-any.whl
- Upload date:
- Size: 26.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d416842a54658d40658018d04fa6c95df0d85140ae08dba96f5bd5166ef39261
|
|
| MD5 |
92f38db7b58163edcd6a7873cf6bd717
|
|
| BLAKE2b-256 |
bac0cbb6774b77c2a018c64e818ecd4aa1af319cfebb8db5b5be5399b78f670a
|











