A package for operating Quantum Chemistry programs using qcdata standardized data structures. Compatible with TeraChem, psi4, QChem, NWChem, ORCA, Molpro, geomeTRIC and many more.
Project description
qccompute
A package for running Quantum Chemistry programs using qcdata standardized data structures. Compatible with TeraChem, psi4, Crest, xTB, QChem, NWChem, ORCA, Molpro, geomeTRIC and many more.
qccompute works in harmony with a suite of other quantum chemistry tools for fast, structured, and interoperable quantum chemistry.
The QC Suite of Programs
- qcconst - Physical constants, conversion factors, and a periodic table with clear source information for every value.
- qcdata - Beautiful and user-friendly data structures for quantum chemistry, featuring seamless Jupyter Notebook visualizations.
- qcinf - Cheminformatics algorithms and structure utilities such as
rmsdand alignment, using standardized qcdata data structures. - qccodec - A library for efficient parsing of quantum chemistry data into structured
qcdataobjects. - qccompute - A package for running quantum chemistry programs using
qcdatastandardized data structures. Compatible withTeraChem,psi4,QChem,NWChem,ORCA,Molpro,geomeTRIC, and many more, featuring seamless Jupyter Notebook visualizations. - BigChem - A distributed application for running quantum chemistry calculations at scale across clusters of computers or the cloud. Bring multi-node scaling to your favorite quantum chemistry program, featuring seamless Jupyter Notebook visualizations of your data.
ChemCloud- A web application and associated Python client for exposing a BigChem cluster securely over the internet, featuring seamless Jupyter Notebook visualizations.
Installation
python -m pip install qccompute
Quickstart
qccompute uses the qcdata data structures to drive quantum chemistry programs in a standardized way. This allows for a simple and consistent interface to a wide variety of quantum chemistry programs. See the qcdata library for documentation on the input and output data structures.
The compute function is the main entry point for the library and is used to run a calculation.
from qcdata import Structure, ProgramInput
from qccompute import compute
from qccompute.exceptions import ExternalProgramError
# Create the Structure
h2o = Structure.open("h2o.xyz")
# Define the program input
prog_input = ProgramInput(
structure=h2o,
calctype="energy",
model={"method": "hf", "basis": "sto-3g"},
keywords={"purify": "no", "restricted": False},
)
# Run the calculation; will return a ProgramOutput or raise an exception
try:
prog_output = compute("terachem", prog_input, collect_files=True)
except ExternalProgramError as e:
# External QQ program failed in some way
prog_output = e.prog_output
prog_output.input_data # Input data used by the QC program
prog_output.success # Will be False
prog_output.data # Any half-computed results before the failure
prog_output.logs # Logs from the calculation
prog_output.plogs # Shortcut to print out the logs in human readable format
prog_output.traceback # Stack trace from the calculation
prog_output.ptraceback # Shortcut to print out the traceback in human readable format
raise e
else:
# Calculation succeeded
prog_output.input_data # Input data used by the QC program
prog_output.success # Will be True
prog_output.data # All structured data and files from the calculation
prog_output.data.files # Any files returned by the calculation
prog_output.logs # Logs from the calculation
prog_output.plogs # Shortcut to print out the logs in human readable format
prog_output.provenance # Provenance information about the calculation
prog_output.extras # Any extra information not in the schema
One may also call compute(..., raise_exc=False) to return a ProgramOutput object rather than raising an exception when a calculation fails. This may allow easier handling of failures in some cases.
from qcdata import Structure, ProgramInput
from qccompute import compute
from qccompute.exceptions import ExternalProgramError
# Create the Structure
h2o = Structure.open("h2o.xyz")
# Define the program input
prog_input = ProgramInput(
structure=h2o,
calctype="energy",
model={"method": "hf", "basis": "sto-3g"},
keywords={"purify": "no", "restricted": False},
)
# Run the calculation; will return a ProgramOutput object
prog_output = compute("terachem", prog_input, collect_files=True, raise_exc=False)
if not prog_output.success:
# Same as except block above
else:
# Same as else block above
Alternatively, the compute_args function can be used to run a calculation with the input data structures passed in as arguments rather than as a single ProgramInput object.
from qcdata import Structure
from qccompute import compute_args
# Create the Structure
h2o = Structure.open("h2o.xyz")
# Run the calculation
prog_output = compute_args(
"terachem",
h2o,
calctype="energy",
model={"method": "hf", "basis": "sto-3g"},
keywords={"purify": "no", "restricted": False},
files={...},
collect_files=True
)
The behavior of compute() and compute_args() can be tuned by passing in keyword arguments like collect_files shown above. Arguments can modify which scratch directory location to use, whether to delete or keep the scratch files after a calculation completes, what files to collect from a calculation, whether to stream the program logs in real time as the program executes, and whether to propagate a wavefunction through a series of calculations. Arguments also include hooks for passing in update functions that can be called as a program executes in real time. See the compute method docstring for more details.
See the /examples directory for more examples.
✨ Visualization ✨
Visualize all your results with a single line of code!
First install the visualization module:
python -m pip install qcdata[view]
or if your shell requires '' around arguments with brackets:
python -m pip install 'qcdata[view]'
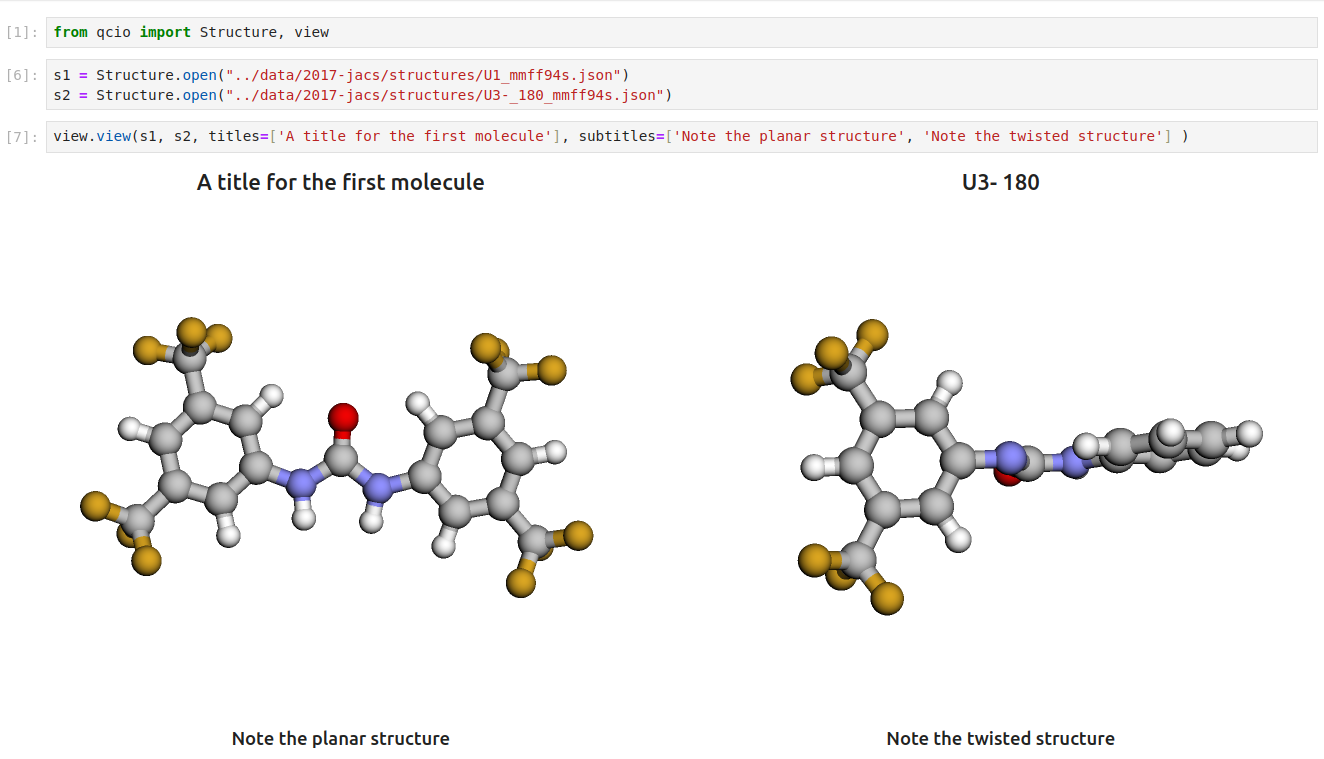
Then in a Jupyter notebook import the qcdata view module and call view.view(...) passing it one or any number of qcdata objects you want to visualize, including Structure objects or any ProgramOutput object. You may also pass arrays of titles and/or subtitles to add additional information to the molecular structure display. If no titles are passed, qcdata will look for Structure identifiers such as a name or SMILES to label the Structure.
Seamless visualizations for ProgramOutput objects make results analysis easy!
Single point calculations display their results in a table.
If you want to use the HTML generated by the viewer to build your own dashboards, use the functions inside qcdata.view.py that begin with generate_ to create HTML you can insert into any dashboard.
Support
If you have any issues with qccompute or would like to request a feature, please open an issue.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file qccompute-0.13.0.tar.gz.
File metadata
- Download URL: qccompute-0.13.0.tar.gz
- Upload date:
- Size: 495.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2cdd04a9c704792cfec4b934da1f4b20b8916d2d56f7b7fdda32624ee5942ee8
|
|
| MD5 |
3ad1dc1bd6c765d1f068c1ab473ad1b3
|
|
| BLAKE2b-256 |
932a1f92b5ba943cee2388ed2066249dabb8875d898e0f13932da655b83e09a5
|
Provenance
The following attestation bundles were made for qccompute-0.13.0.tar.gz:
Publisher:
publish-to-pypi.yaml on coltonbh/qccompute
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
qccompute-0.13.0.tar.gz -
Subject digest:
2cdd04a9c704792cfec4b934da1f4b20b8916d2d56f7b7fdda32624ee5942ee8 - Sigstore transparency entry: 1186876950
- Sigstore integration time:
-
Permalink:
coltonbh/qccompute@e06ecf0dfd8f9868b5a7f1b5a8181f075a939956 -
Branch / Tag:
refs/tags/0.13.0 - Owner: https://github.com/coltonbh
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-to-pypi.yaml@e06ecf0dfd8f9868b5a7f1b5a8181f075a939956 -
Trigger Event:
push
-
Statement type:
File details
Details for the file qccompute-0.13.0-py3-none-any.whl.
File metadata
- Download URL: qccompute-0.13.0-py3-none-any.whl
- Upload date:
- Size: 32.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9d5018a7fc08c93aa2b193e94d8eac6a558e0e9ee0b1fbd91bb6545f33175977
|
|
| MD5 |
017541b01a50829bb603722a9472e574
|
|
| BLAKE2b-256 |
0bf1722ca0f17f61c3dda112c69ce977a549750c2f50d456c4f263bf8424a2cf
|
Provenance
The following attestation bundles were made for qccompute-0.13.0-py3-none-any.whl:
Publisher:
publish-to-pypi.yaml on coltonbh/qccompute
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
qccompute-0.13.0-py3-none-any.whl -
Subject digest:
9d5018a7fc08c93aa2b193e94d8eac6a558e0e9ee0b1fbd91bb6545f33175977 - Sigstore transparency entry: 1186877028
- Sigstore integration time:
-
Permalink:
coltonbh/qccompute@e06ecf0dfd8f9868b5a7f1b5a8181f075a939956 -
Branch / Tag:
refs/tags/0.13.0 - Owner: https://github.com/coltonbh
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-to-pypi.yaml@e06ecf0dfd8f9868b5a7f1b5a8181f075a939956 -
Trigger Event:
push
-
Statement type: