A forward-model simulation for Quantitative Susceptibility Mapping
Project description
QSM Forward Model
This package provides a Python API and CLI for simulating input data for Quantitative Susceptibility Mapping (QSM), including BIDS-compliant magnitude and phase MRI. This is also known as the QSM forward problem or forward model. A good quality forward model is important for testing and evaluating QSM algorithms under controlled conditions and in developing deep-learning models for QSM.
Based on Marques, J. P., et al. (2021). QSM reconstruction challenge 2.0: A realistic in silico head phantom for MRI data simulation and evaluation of susceptibility mapping procedures. Magnetic Resonance in Medicine, 86(1), 526-542. https://doi.org/10.1002/mrm.28716
Includes code for:
- Field model (forward multiplication with dipole kernel based on chi)
- Signal model (magnitude and phase simulation based on field/M0/R1/R2star)
- Phase offset model
- Noise model
- Shim field model
- k-space cropping
Install
pip install qsm-forward
Example using simulated sources
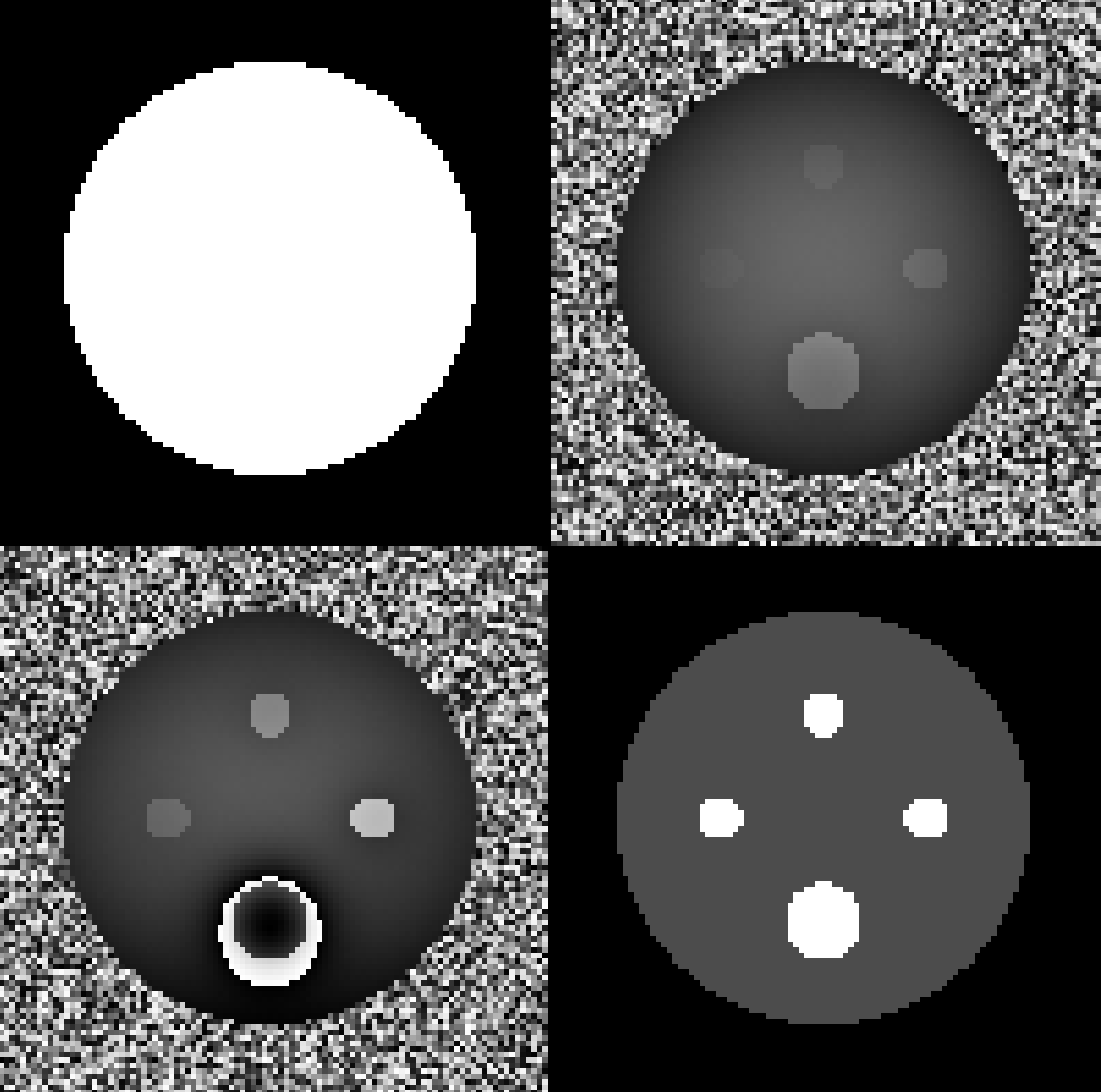
In this example, we simulated susceptibility sources (spheres and rectangles) to generate a BIDS directory:
import qsm_forward
if __name__ == "__main__":
recon_params = qsm_forward.ReconParams()
recon_params.subject = "simulated-sources"
recon_params.peak_snr = 100
recon_params.random_seed = 42
tissue_params = qsm_forward.TissueParams(
chi=qsm_forward.generate_susceptibility_phantom(
resolution=[100, 100, 100],
background=0,
large_cylinder_val=0.005,
small_cylinder_radii=[4, 4, 4, 7],
small_cylinder_vals=[0.05, 0.1, 0.2, 0.5]
)
)
qsm_forward.generate_bids(tissue_params, recon_params, "bids")
bids/
└── sub-simulated-sources
└── ses-1
├── anat
│ ├── sub-simulated-sources_ses-1_run-1_echo-1_part-mag_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-1_part-mag_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-1_part-phase_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-1_part-phase_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-2_part-mag_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-2_part-mag_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-2_part-phase_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-2_part-phase_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-3_part-mag_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-3_part-mag_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-3_part-phase_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-3_part-phase_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-4_part-mag_MEGRE.json
│ ├── sub-simulated-sources_ses-1_run-1_echo-4_part-mag_MEGRE.nii
│ ├── sub-simulated-sources_ses-1_run-1_echo-4_part-phase_MEGRE.json
│ └── sub-simulated-sources_ses-1_run-1_echo-4_part-phase_MEGRE.nii
└── extra_data
├── sub-simulated-sources_ses-1_run-1_chi.nii
├── sub-simulated-sources_ses-1_run-1_mask.nii
└── sub-simulated-sources_ses-1_run-1_segmentation.nii
Some repesentative images including the mask, first and last-echo phase image, and ground truth susceptibility (chi):
Example using head phantom data
In this example, we generate a BIDS-compliant dataset based on the realistic in-silico head phantom. If you have access to the head phantom, you need to retain the data directory which provides relevant tissue parameters:
import qsm_forward
import numpy as np
if __name__ == "__main__":
tissue_params = qsm_forward.TissueParams(root_dir="~/data")
recon_params_all = [
qsm_forward.ReconParams(voxel_size=voxel_size, peak_snr=100, random_seed=42, session=session)
for (voxel_size, session) in [
(np.array([0.8, 0.8, 0.8]), "0p8"),
(np.array([1.0, 1.0, 1.0]), "1p0"),
(np.array([1.2, 1.2, 1.2]), "1p2")
]
]
for recon_params in recon_params_all:
qsm_forward.generate_bids(tissue_params=tissue_params, recon_params=recon_params, bids_dir="bids")
bids/
└── sub-1
├── ses-0p8
│ ├── anat
│ │ ├── sub-1_ses-0p8_run-1_echo-1_part-mag_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-1_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-1_part-phase_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-1_part-phase_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-2_part-mag_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-2_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-2_part-phase_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-2_part-phase_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-3_part-mag_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-3_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-3_part-phase_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-3_part-phase_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-4_part-mag_MEGRE.json
│ │ ├── sub-1_ses-0p8_run-1_echo-4_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-0p8_run-1_echo-4_part-phase_MEGRE.json
│ │ └── sub-1_ses-0p8_run-1_echo-4_part-phase_MEGRE.nii
│ └── extra_data
│ ├── sub-1_ses-0p8_run-1_chi.nii
│ ├── sub-1_ses-0p8_run-1_mask.nii
│ └── sub-1_ses-0p8_run-1_segmentation.nii
├── ses-1p0
│ ├── anat
│ │ ├── sub-1_ses-1p0_run-1_echo-1_part-mag_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-1_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-1_part-phase_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-1_part-phase_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-2_part-mag_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-2_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-2_part-phase_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-2_part-phase_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-3_part-mag_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-3_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-3_part-phase_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-3_part-phase_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-4_part-mag_MEGRE.json
│ │ ├── sub-1_ses-1p0_run-1_echo-4_part-mag_MEGRE.nii
│ │ ├── sub-1_ses-1p0_run-1_echo-4_part-phase_MEGRE.json
│ │ └── sub-1_ses-1p0_run-1_echo-4_part-phase_MEGRE.nii
│ └── extra_data
│ ├── sub-1_ses-1p0_run-1_chi.nii
│ ├── sub-1_ses-1p0_run-1_mask.nii
│ └── sub-1_ses-1p0_run-1_segmentation.nii
└── ses-1p2
├── anat
│ ├── sub-1_ses-1p2_run-1_echo-1_part-mag_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-1_part-mag_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-1_part-phase_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-1_part-phase_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-2_part-mag_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-2_part-mag_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-2_part-phase_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-2_part-phase_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-3_part-mag_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-3_part-mag_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-3_part-phase_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-3_part-phase_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-4_part-mag_MEGRE.json
│ ├── sub-1_ses-1p2_run-1_echo-4_part-mag_MEGRE.nii
│ ├── sub-1_ses-1p2_run-1_echo-4_part-phase_MEGRE.json
│ └── sub-1_ses-1p2_run-1_echo-4_part-phase_MEGRE.nii
└── extra_data
├── sub-1_ses-1p2_run-1_chi.nii
├── sub-1_ses-1p2_run-1_mask.nii
└── sub-1_ses-1p2_run-1_segmentation.nii
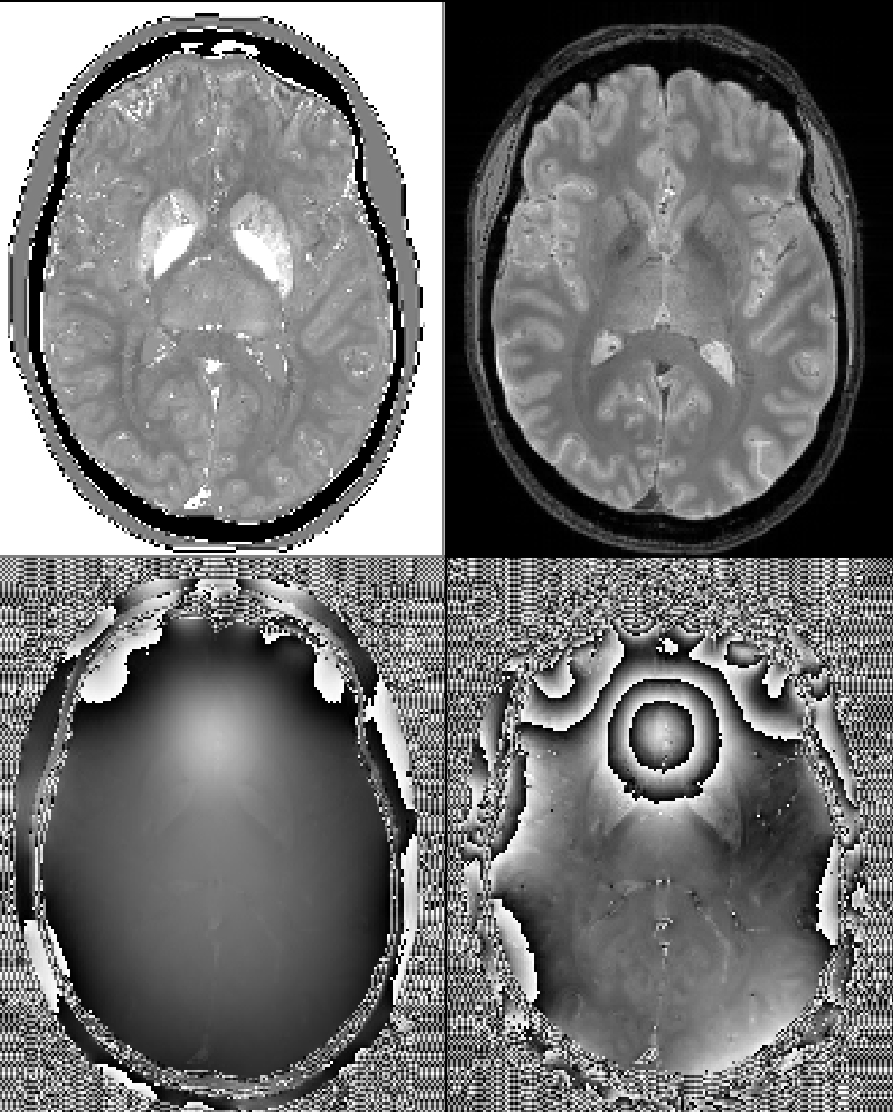
Some repesentative images including the ground truth chi map, first-echo magnitude image, and first and last-echo phase images:
Example including T1-weighted images
import qsm_forward
import numpy as np
if __name__ == "__main__":
tissue_params = qsm_forward.TissueParams(root_dir="~/data", chi="ChiModelMIX.nii.gz")
recon_params_all = [
qsm_forward.ReconParams(voxel_size=voxel_size, session=session, TEs=TEs, TR=TR, flip_angle=flip_angle, random_seed=42, suffix=suffix, save_phase=save_phase)
for (voxel_size, session, TEs, TR, flip_angle, suffix, save_phase) in [
(np.array([0.64, 0.64, 0.64]), "0p64", np.array([3.5e-3]), 7.5e-3, 40, "T1w", False),
(np.array([0.64, 0.64, 0.64]), "0p64", np.array([0.004, 0.012, 0.02, 0.028]), 0.05, 15, "T2starw", True),
]
]
for recon_params in recon_params_all:
qsm_forward.generate_bids(tissue_params=tissue_params, recon_params=recon_params, bids_dir="bids")
bids/
└── sub-1
└── ses-0p64
├── anat
│ ├── sub-1_ses-0p64_run-1_echo-1_part-mag_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-1_part-mag_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-1_part-phase_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-1_part-phase_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-2_part-mag_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-2_part-mag_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-2_part-phase_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-2_part-phase_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-3_part-mag_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-3_part-mag_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-3_part-phase_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-3_part-phase_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-4_part-mag_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-4_part-mag_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_echo-4_part-phase_MEGRE.json
│ ├── sub-1_ses-0p64_run-1_echo-4_part-phase_MEGRE.nii
│ ├── sub-1_ses-0p64_run-1_T1w.json
│ └── sub-1_ses-0p64_run-1_T1w.nii
└── extra_data
├── sub-1_ses-0p64_run-1_chi.nii
├── sub-1_ses-0p64_run-1_mask.nii
└── sub-1_ses-0p64_run-1_segmentation.nii
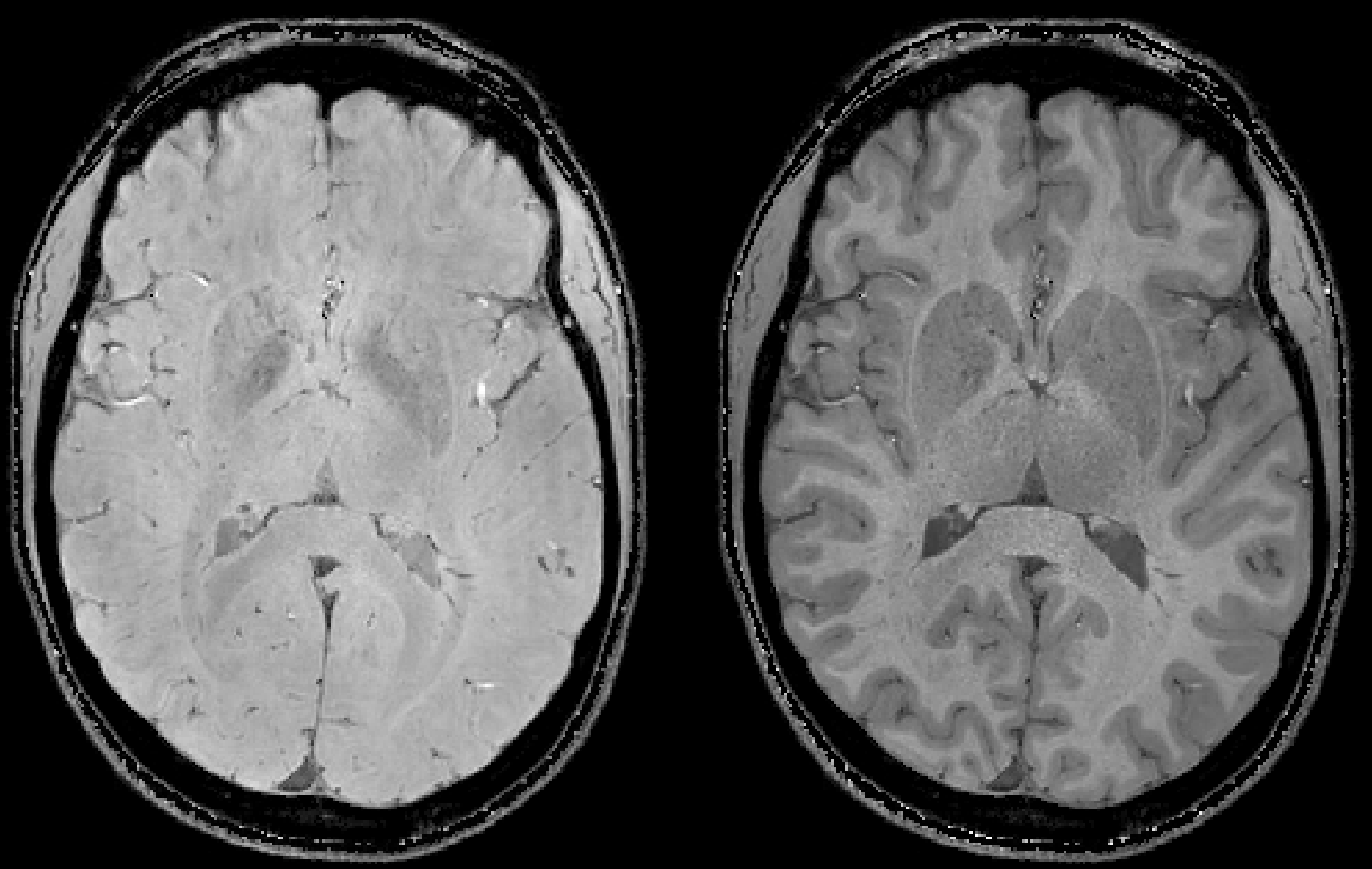
Some repesentative images including the T2starw and T1w magnitude images:
Example simulating oblique acquisition
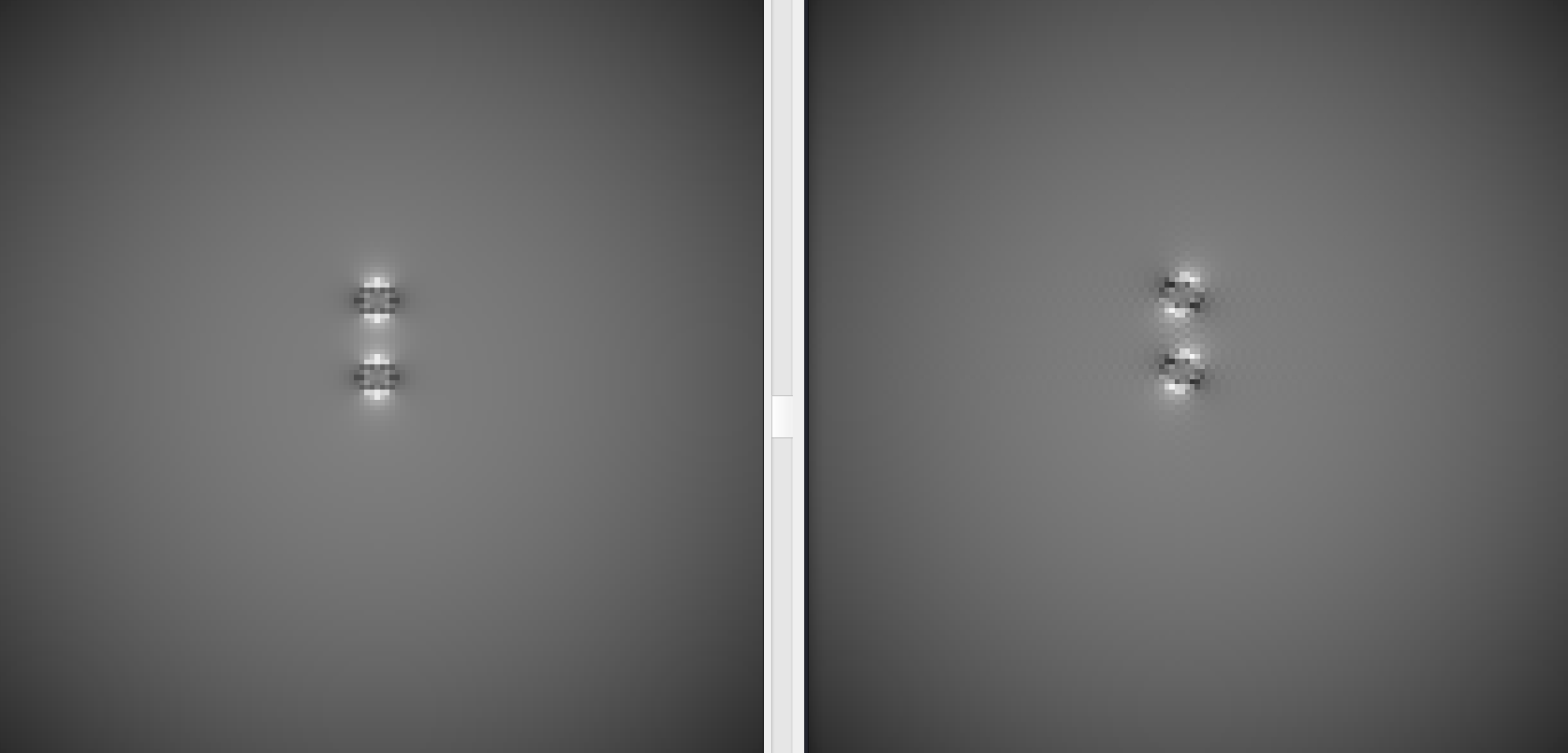
In this example, we simulated spherical susceptibility sources to generate a BIDS directory with a range of B0 directions:
On the left is the phase image with the two sources with an axial B0 direction. On the right is a phase image with the two sources with a B0 direction rotated 30 degrees about the x axis.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file qsm_forward-0.25.tar.gz.
File metadata
- Download URL: qsm_forward-0.25.tar.gz
- Upload date:
- Size: 32.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.23
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c2c67d0a32814f03ba0ddf6d3ed7d6bf8df01d67e33ab340e28ebc95f036d74c
|
|
| MD5 |
084149bad2cf3308ad6717750e0d2642
|
|
| BLAKE2b-256 |
59b1cac371597bbfdc9220b37ad83e1d3da6840356ebf48b044dc9d7827737d9
|
File details
Details for the file qsm_forward-0.25-py3-none-any.whl.
File metadata
- Download URL: qsm_forward-0.25-py3-none-any.whl
- Upload date:
- Size: 28.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.23
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9d29f1792d1ae6c2f56cc65cba3c1bffc4f6bb1f45e1690137787e2741d1971a
|
|
| MD5 |
83ed6ae51567f6f775d1329ce1fcba6e
|
|
| BLAKE2b-256 |
4ab95c20c112168c1e28c2c63d4821eb74c68ee56ccfe699ea0bfec22e2e4c8e
|