Utitilies for constructing and manipulating models for non-local structural dependencies in genomic sequences
Project description
Quasinet

Description
Infer non-local structural dependencies in genomic sequences. Genomic sequences are esentially compressed encodings of phenotypic information. This package provides a novel set of tools to extract long-range structural dependencies in genotypic data that define the phenotypic outcomes. The key capabilities implemented here are as follows:
- Compute the Quasinet (Q-net) given a database of nucleic acid sequences. The Q-net is a family of conditional inference trees that capture the predictability of each nucleotide position given the rest of the genome. The constructed Q-net for COVID-19 and Influenza A H1N1 HA 2008-9 is shown below.
| COVID-19 | INFLUENZA |
|---|---|
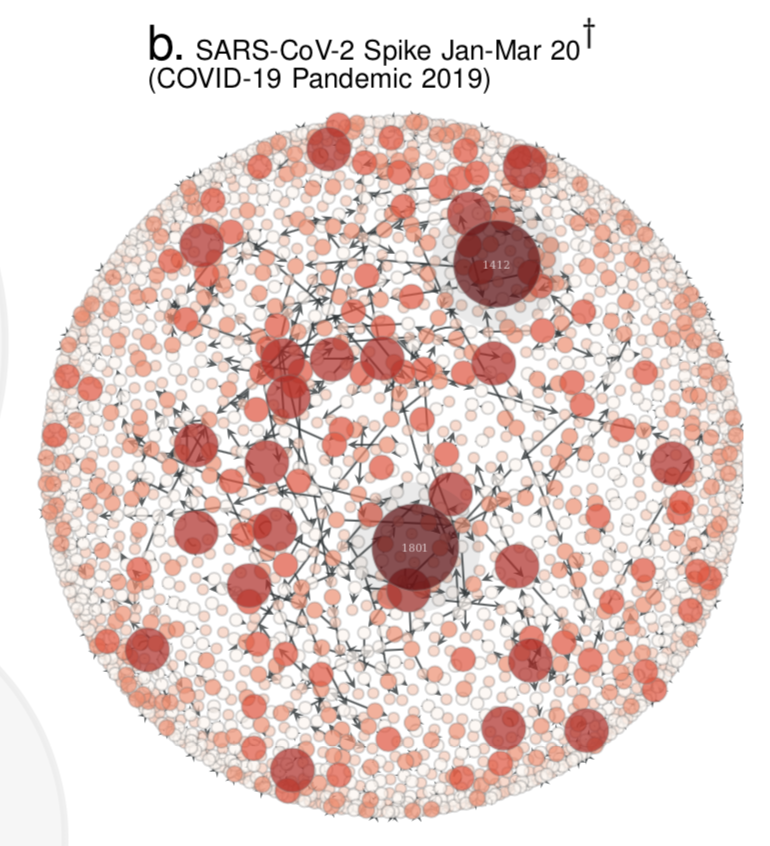 |
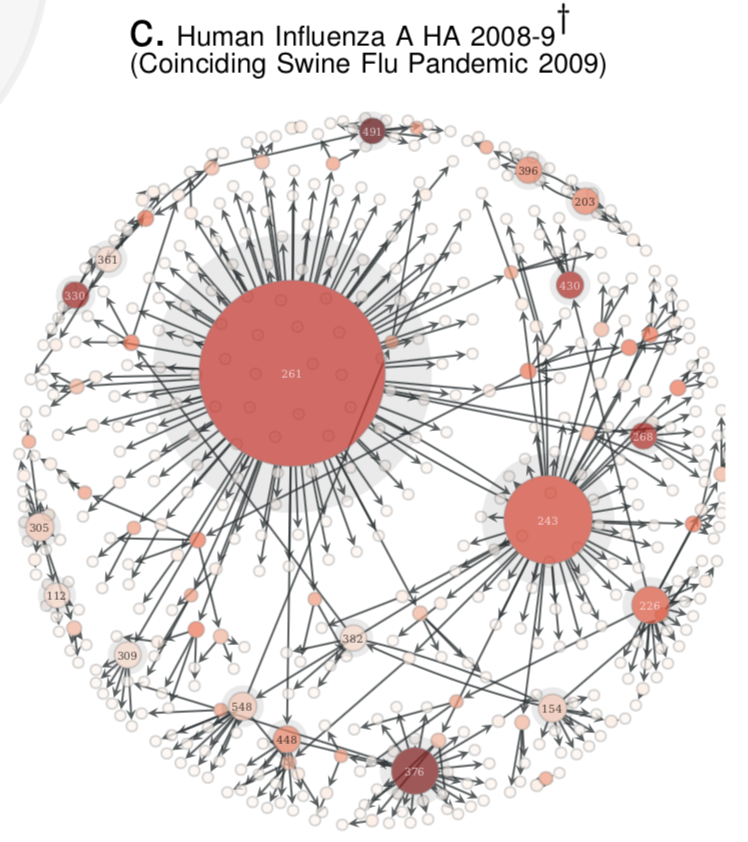 |
-
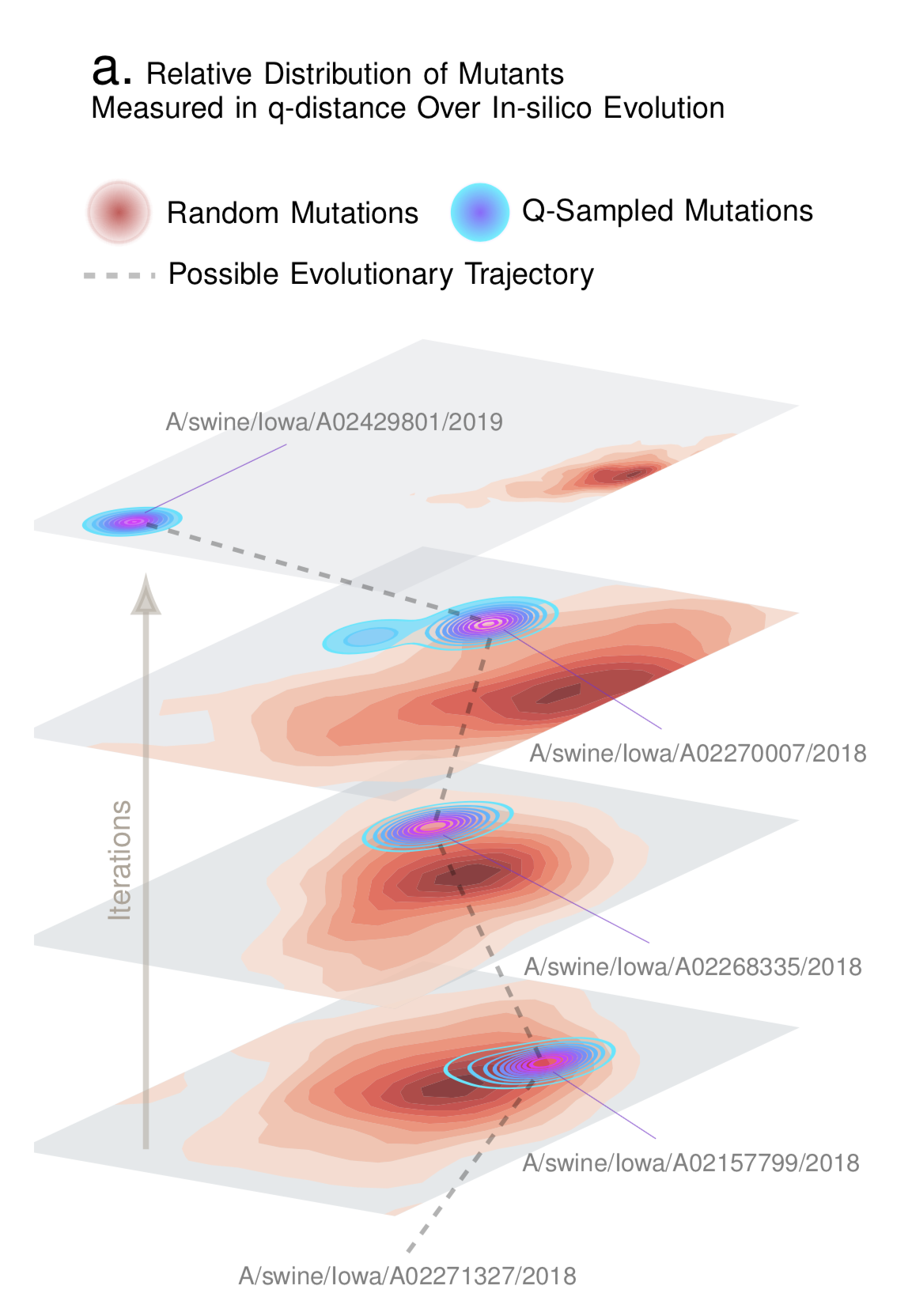
Compute a structure-aware evolution-adaptive notion of distance between genomes, which is demonstrably more biologically relevant compared to the standard edit distance.
-
Draw samples in-silico that have a high probability of being biologically correct. For example, given a database of Influenza sequences, we can generate a new genomic sequence that has a high probability of being a valid influenza sequence.
Installation
To install with pip:
pip install quasinet
NOTE: If trying to reproduce the paper below, please use pip install quasinet==0.0.58
Dependencies
- scikit-learn
- scipy
- numpy
- numba
- pandas
- joblib
- biopython
Usage
from quasinet import qnet
# initialize qnet
myqnet = qnet.Qnet()
# train the qnet
myqnet.fit(X)
# compute qdistance
qdist = qnet.qdistance(seq1, seq2, myqnet, myqnet)
Examples
Examples are located here.
Documentation
For more documentation, see here.
Papers
For reference, please check out our paper:
Authors
You can reach the ZED lab at: zed.uchicago.edu
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file quasinet-0.0.63.tar.gz.
File metadata
- Download URL: quasinet-0.0.63.tar.gz
- Upload date:
- Size: 14.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/50.3.0 requests-toolbelt/0.9.1 tqdm/4.47.0 CPython/3.7.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
53efe0bb96b137c1b3f1e374569499bea96467de8ca6cbeec97693c7cabe99da
|
|
| MD5 |
20eb54e46471ecb7c3ca12a7fa8df566
|
|
| BLAKE2b-256 |
caeb8b55e8b371e63d10a1fe7a4715a8482d234fcc2fd6fb13252bf0c9dcecf2
|
File details
Details for the file quasinet-0.0.63-py3-none-any.whl.
File metadata
- Download URL: quasinet-0.0.63-py3-none-any.whl
- Upload date:
- Size: 15.1 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.2.0 pkginfo/1.5.0.1 requests/2.24.0 setuptools/50.3.0 requests-toolbelt/0.9.1 tqdm/4.47.0 CPython/3.7.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c360301e560e442f9ca69054b439271712dea39090d4d2d840fa920481e90662
|
|
| MD5 |
db899941b2f0d2c284fd52d28cc4aead
|
|
| BLAKE2b-256 |
3f99afae6a25caeedd1c889f495c7ed91d6346a2dc02ef9f93df7313011067a1
|















