Querying functional and structural niches on spatial transcriptomics data
Project description
Querying functional and structural niches on spatial transcriptomics data
Overview
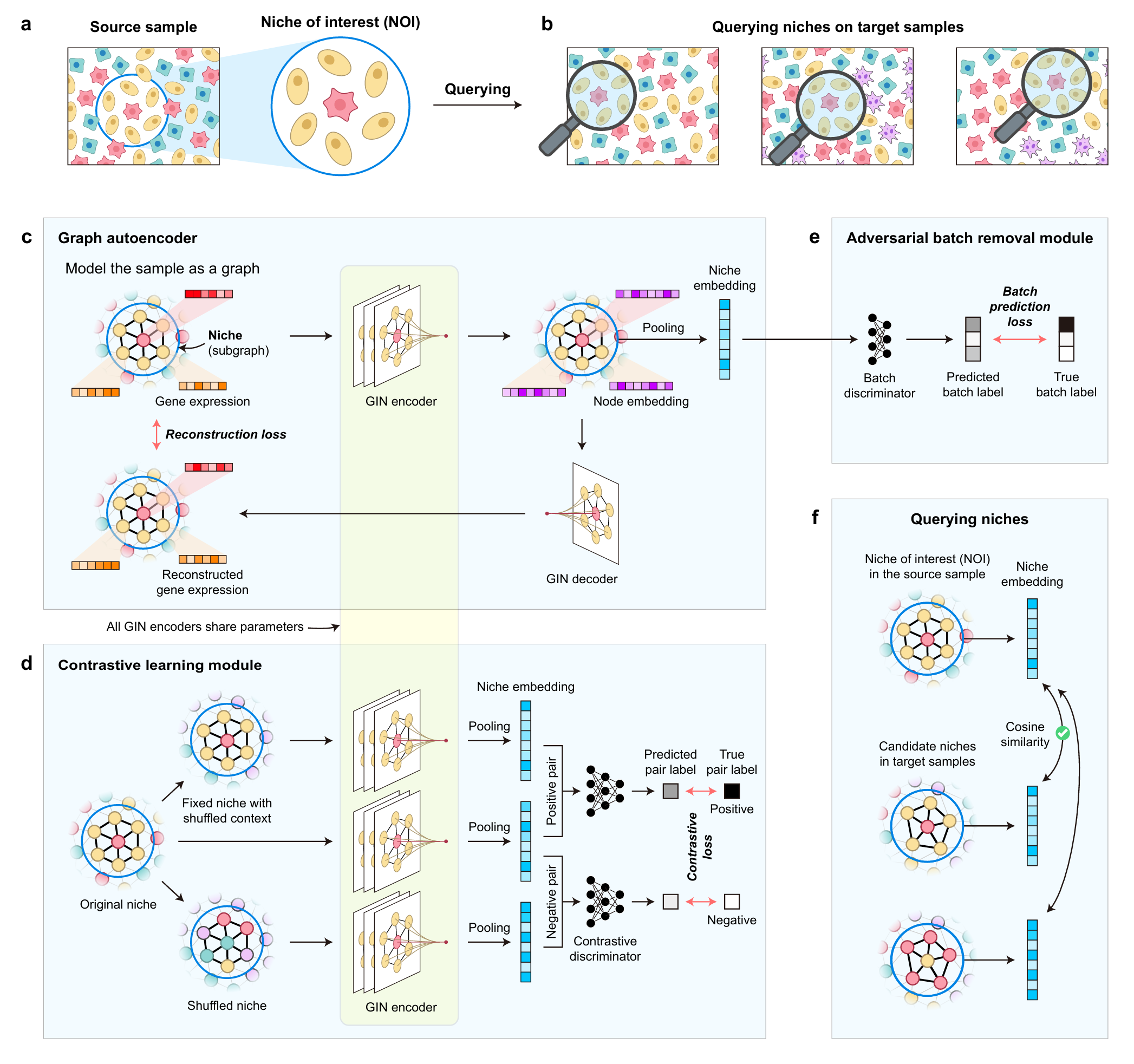
Cells in multicellular organisms coordinate to form functional and structural niches. With spatial transcriptomics enabling gene expression profiling in spatial contexts, it has been revealed that spatial niches serve as cohesive and recurrent units in physiological and pathological processes. These observations suggest universal tissue organization principles encoded by conserved niche patterns, and call for a query-based niche analytical paradigm beyond current computational tools. In this work, we defined the Niche Query Task, which is to identify similar niches across ST samples given a niche of interest (NOI). We further developed QueST, a specialized method for solving this task. QueST models each niche as a subgraph, uses contrastive learning to learn discriminative niche embeddings, and incorporates adversarial training to mitigate batch effects. In simulations and benchmark datasets, QueST outperformed existing methods repurposed for niche querying, accurately capturing niche structures in heterogeneous environments and demonstrating strong generalizability across diverse sequencing platforms. Applied to tertiary lymphoid structures in renal and lung cancers, QueST revealed functionally distinct niches associated with patient prognosis and uncovered conserved and divergent spatial architectures across cancer types. These results demonstrate that QueST enables systematic, quantitative profiling of spatial niches across samples, providing a powerful tool to dissect spatial tissue architecture in health and disease.
Getting started
Installation
We recommend using a conda environment
conda create -n quest python==3.9.19
Install necessary dependencies first before the installation of QueST
conda activate quest
pip install -r requirements.txt
Finally, QueST is available on PyPI and can be installed via
pip install quest-niche
Usage
See detailed usage on Read the Docs website.
Data Availability
- For complete reference datasets and pretrained model weights, please refer to https://doi.org/10.6084/m9.figshare.29432396.v1
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file quest_niche-0.1.5.tar.gz.
File metadata
- Download URL: quest_niche-0.1.5.tar.gz
- Upload date:
- Size: 19.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
763bb477c88ed7bd8e030738de6477618c7fcbcf3c1d77b21794a73ac295d2eb
|
|
| MD5 |
ca73e63f6d402730eeebf3a3f7cb85c8
|
|
| BLAKE2b-256 |
baa608120b78b6a302964414b340ad2f35e6e1a5c9e12c85979a8dad88d91394
|
File details
Details for the file quest_niche-0.1.5-py3-none-any.whl.
File metadata
- Download URL: quest_niche-0.1.5-py3-none-any.whl
- Upload date:
- Size: 20.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cbfb75a1eb9e425ea2643724539482a9ca558fe48cda596cdd2142808f9bfcdc
|
|
| MD5 |
991bf69dcceccf940be5dd4820b406a7
|
|
| BLAKE2b-256 |
ea5593a89757640e0f74bc156870e085efd15fe680858d891abfdb63246632f8
|











