Implementation of the IDR (Irreproducible Discovery Rate) method for RNA reactivity data.
Project description

reactIDR: evaluation of the statistical reproducibility of high-throughput structural analyses towards a robust RNA structure prediction
- Published in BMC Bioinformatics
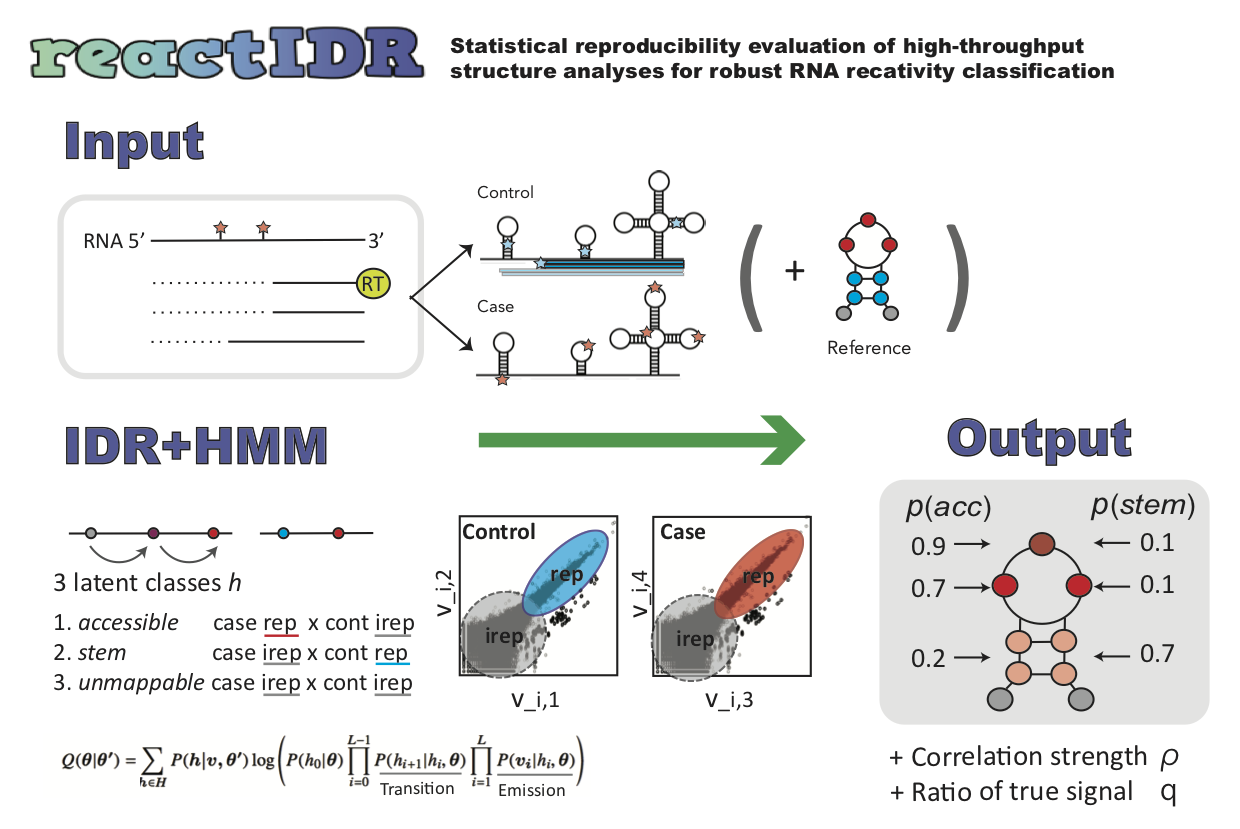
reactIDR is a Python package that evaluates statistical reproducibility across replicated high-throughput RNA structure profiling data (e.g., PARS, SHAPE-Seq, icSHAPE, DMS-Seq) to robustly infer loop and stem probabilities.
📥 Input
- Read count data (tabular format)
- PARS
- SHAPE-Seq
- icSHAPE
- DMS-Seq (assumed to enrich A/C only)
📤 Output
- Posterior probability for each site:
- Loop (signal enriched in "case")
- Stem (signal enriched in "control")
🧠 Algorithm
- IDR (Irreproducible Discovery Rate)
- Hidden Markov Model
🔧 Requirements
python >= 3.9
numpy >= 2.0.2
scipy >= 1.13.1
pandas >= 2.2.3
Optional packages for visualization:
seaborn
jupyter notebook
🚀 Installation
pip install reactIDR
▶️ Getting Started
Test datasets are provided in the example and csv_example directories. To run a demo using CSV input:
git clone https://github.com/carushi/reactIDR
cd reactIDR/csv_example
python -c "import reactIDR; reactIDR.run_reactIDR([
'-e', '0',
'--csv',
'--global',
'--case', './csv_example/case.csv',
'--output', 'test.csv',
'--param', './csv_example/default_parameters.txt'
])"
📚 More usage examples and options are available in the Wiki.
🛠️ Scripts
| Script | Description |
|---|---|
read_collapse.py |
Collapse PCR duplicates and trim barcodes (assumes gawk) |
read_truncate.py |
Extract consistent paired-end reads |
bed_to_pars_format.py |
Convert BED coverage to PARS-style format based on annotations |
| format: NAME 0;1;2;3;..... | |
tab_to_csv.py |
Append raw count data to output CSV |
📖 Reference
-
R. Kawaguchi, H. Kiryu, J. Iwakiri and J. Sese. "reactIDR: evaluation of the statistical reproducibility of high-throughput structural analyses towards a robust RNA structure prediction" BMC Bioinformatics 20 (Suppl 3) :130 (2019)" ー Selected for APBC '19 proceedings
-
- Convert bam to read count data
- Find scripts and how to use at https://github.com/carushi/RT_end_counter
-
IDR
- Li, Qunhua, et al. "Measuring reproducibility of high-throughput experiments", The annals of applied statistics, 2011.
- IDR in Python
- IDR in R
TODO
- apply to MaP analyses
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file reactidr-2.0.0.tar.gz.
File metadata
- Download URL: reactidr-2.0.0.tar.gz
- Upload date:
- Size: 142.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
99dc5972560263ff543657957bca1aef01142cffe0edfac4dcd6e7494ac9c42a
|
|
| MD5 |
8ed8bd2cd03eb8deeada808b22458d2e
|
|
| BLAKE2b-256 |
e7506a0d70fe0229803c3efb54041e8061927c3fd51998355f57f4c367aba9dd
|
File details
Details for the file reactidr-2.0.0-py3-none-any.whl.
File metadata
- Download URL: reactidr-2.0.0-py3-none-any.whl
- Upload date:
- Size: 147.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.19
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9e54b4779f34f4246f76533422a3a6acac9e55a0f0f8efcf862e6e289574baa9
|
|
| MD5 |
647122bcbcd2aebd1137ab245b03a782
|
|
| BLAKE2b-256 |
1229228024d94bcc8d886d3e0f981b9a83de3c887ae14a8641b65b4bfb18e388
|











