Python tool to read and plot Geophysical Survey Systems Incorporated (GSSI) radar data in DZT format
Project description
readgssi
Copyleft 🄯 2017-2022
This program was written to read and process ground-penetrating radar files from instruments made by Geophysical Survey Systems Incorporated (GSSI), although I have no affiliation with nor endorsement for the afforementioned organization.
The demands of ice- and ground-penetrating radar (GPR) surveying, as in many types of scientific fieldwork, require that both quality control and time savings are critical to a successful field campaign. This software provides a way to quickly read, process, and display radar data produced by GSSI radar antennas and control units. readgssi was designed to be used in the field to quality-check entire folders of data files by converting radar profiles to portable network graphics (PNG) images, saving users valuable time versus performing the equivalent actions by hand in RADAN, especially in the case of projects with large file counts.
readgssi has the capability to read DZT and DZG files with the same pre-extension name and plot the data contained in those files. readgssi is also currently able to translate most DZT files to CSV and numpy binary (see future). The file read parameters are based on GSSI's DZT file description, similar to the ones available on pages 55-57 of the SIR-3000 manual. File structure is, unfortunately, prone to change at any time, and although I've been able to test with files from several systems, I have not encountered every iteration of file header yet. If you run into trouble, please create a github issue.
Full documentation, including a tutorial, is available at https://readgssi.readthedocs.io.
Questions, feature requests, and bugs: please open a github issue. Kindly provide the error output, describe what you are attempting to do, and attach the DZT/DZG file(s) causing you trouble.
requirements
Strongly recommended to install via anaconda:
Install via pip:
installation
Once you have anaconda running, installing is pretty easy.
You can install via the anaconda cloud:
conda create -n readgssi -c iannesbitt readgssi
# and then activate by doing:
conda activate readgssi
or via pip/PyPI:
conda config --add channels conda-forge
conda create -n readgssi python==3.7 pandas h5py pytz obspy
conda activate readgssi
pip install readgssi
That should allow you to run the commands below.
installing from source:
If you choose to install a specific commit rather than the latest working release of this software, you may download this package, unzip to your home folder, open a command line, then install in the following way:
pip install ~/readgssi
usage
To display the help text:
$ readgssi -h
usage:
readgssi -i input.DZT [OPTIONS]
required flags:
OPTION | ARGUMENT | FUNCTIONALITY
-i, --input | file: /dir/f.DZT | input DZT file
optional flags:
OPTION | ARGUMENT | FUNCTIONALITY
-o, --output | file: /dir/f.ext | output file. if not set, will be named similar to input
-f, --format | string, eg. "csv" | output format (CSV and DZT are the only working formats currently available from the command line)
-p, --plot | +integer or "auto" | plot size. will be x inches high or "auto". default: 10. see also -D to set DPI
-D, --dpi | positive integer | set the plot DPI for figure making. defaults to 150
-T, --titleoff | | turn the plot title off (useful for figure making)
-x, --xscale | string, eg. "dist" | x units. will attempt to convert the x-axis to distance, time, or trace units based on header values
-z, --zscale | string, eg. "time" | z units. attempt to convert the x-axis to depth, time, or sample units based on header values
-e, --zoom | list of +int [LRUD] | set a zoom to automatically jump to. list order is [left,right,up,down] and units are the same as axis
-n, --noshow | | suppress matplotlib popup window and simply save a figure (useful for multi-file processing)
-c, --colormap | string, eg. "Greys" | specify the colormap (https://matplotlib.org/users/colormaps.html#grayscale-conversion)
-g, --gain | positive float | gain constant (higher=greater contrast, default: 1)
-A, --absval | | Displays the absolute value of the vertical gradient of the array when plotting. Good for displaying faint array features.
-r, --bgr | +integer or zero | horizontal background removal (useful to remove ringing). zero=full width; positive=window size (after stacking)
-R, --reverse | | reverse (flip array horizontally)
-w, --dewow | | trinomial dewow algorithm
-t, --bandpass | +int-+int (MHz) | triangular FIR bandpass filter applied vertically (positive integer range in megahertz; ex. 70-130)
-b, --colorbar | | add a colorbar to the radar figure
-a, --antfreq | positive integer | set antenna frequency. overrides header value
-s, --stack | +integer or "auto" | set trace stacking value or "auto" to autostack to ~2.5:1 x:y axis ratio
-N, --normalize | | distance normalize; reads .DZG NMEA data file if it exists; otherwise tries to read CSV with lat, lon, and time fields
-P, --pausecorr | | pause correction; fixes decoupling of DZG and DZT trace numbers during survey pauses using low velocity GPS marks
-d, --spm | positive float | specify the samples per meter (spm). overrides header value
-m, --histogram | | produce a histogram of data values
-E, --epsr | float > 1.0 | user-defined epsilon sub r (sometimes referred to as "dielectric") if set, ignores value in DZT header
-Z, --zero | +int or list of int | timezero: skip samples before direct wave. samples are removed from the top of the trace. use list for multi-channel
naming scheme for exports:
CHARACTERS | MEANING
Ch0 | Profile from channel 0 (can range from 0 - 3)
Dn | Distance normalization
Tz233 | Time zero at 233 samples
S8 | Stacked 8 times
Rv | Profile read in reverse (flipped horizontally)
Bgr75 | Background removal filter with window size of 75
Dw | Dewow filter
Bp70-130 | triangular FIR filter applied from 70 to 130 MHz
G30 | 30x contrast gain
Abs | Color scale represents absolute value of vertical gradient
Z10.20.7.5 | zoom from 10-20 axis units on the x-axis and 5-7 on the z-axis
From a unix command line:
readgssi -i DZT__001.DZT
Simply specifying an input DZT file like in the above command (-i file) will display a host of data about the file including:
- name of GSSI control unit
- antenna model
- antenna frequency
- samples per trace
- bits per sample
- traces per second
- L1 dielectric as entered during survey
- sampling depth
- speed of wave at given dielectric
- number of traces
- number of seconds
basic functionality
CSV output
readgssi -i DZT__001.DZT -o test.csv -f CSV
Translates radar data array to CSV format, if that's your cup of tea. One might use this to export to Matlab. One CSV will be written per channel. The script will rename the output to 'test_100MHz.csv' automatically. No header information is included in the CSV.
readgssi -i DZT__001.DZT -s 8 -r 0 -o test.csv -f CSV
Applies 8x stacking, and background removal filters before exporting to CSV (see explanations below).
DZT output (experimental)
readgssi -i DZT__001.DZT -R -Z 0 -o test.DZT -f DZT
This exports the array as a second DZT file after reversing the direction of the survey line.
The following operations will stick when writing files:
- stacking
- filtering
- distance normalization
- reversing direction
- zero (top of array will be shaved off; to avoid this, set
-Z 0. this will be fixed in the future.) - manually setting header values, ror example samples per meter (
-d 10), epsilon_r (-E 3.1), and antenna frequency (-a 100)
The following operations will not stick:
- absolute value of gradient
- gain
- zoom
- plot features like axis units (obviously)
Please note the following about DZT output:
- While DZTs will be written, there will be no accompanying DZX written. This functionality may be implemented in the future but for now is a bit too complex of an operation for readgssi. This could have consequences if you're planning to re-import to RADAN after this operation.
- Single channel files only are supported at this time. Multi-channel files will be written but the values on the second channel will not be correct. If you wish to use readgssi to re-write multiple channels, you will have to separate the channels using RADAN or another software first.
numpy object output
readgssi -i DZT__001.DZT -o test.npy -f numpy
This command will save the array in numpy binary format. Header information will not be saved, however.
GPRPy-compatible output
readgssi -i DZT__001.DZT -o test -f gprpy
This command saves the array in numpy binary format, and preserves the header as a JSON file as well. GPRPy will soon support importing this type of file.
plotting
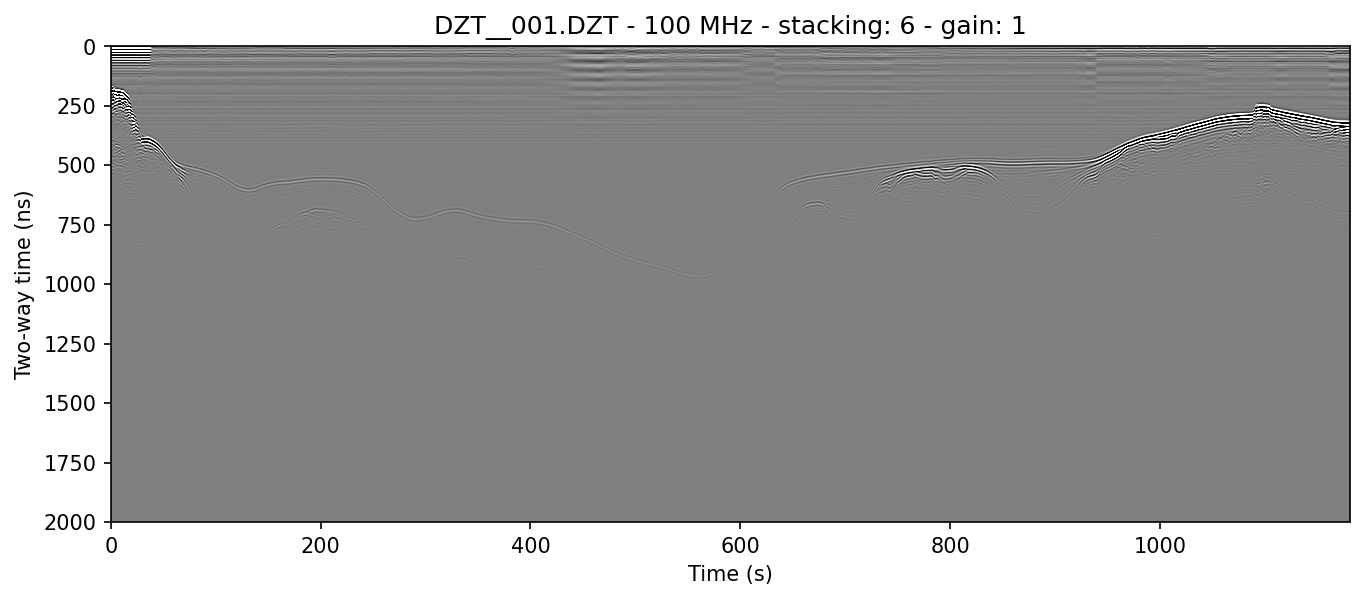
example 1A: without gain
readgssi -i DZT__001.DZT -o 1a.png -p 5 -s auto
The above command will cause readgssi to save and show a plot named "DZT__001c0Tz233S6G1.png" with a y-size of 5 inches at 150 dpi (-p 5) and the autostacking algorithm will stack the x-axis to some multiple of times shorter than the original data array for optimal viewing on a monitor, approximately 2.5*y (-s auto). The plot will be rendered in the gray color scheme.
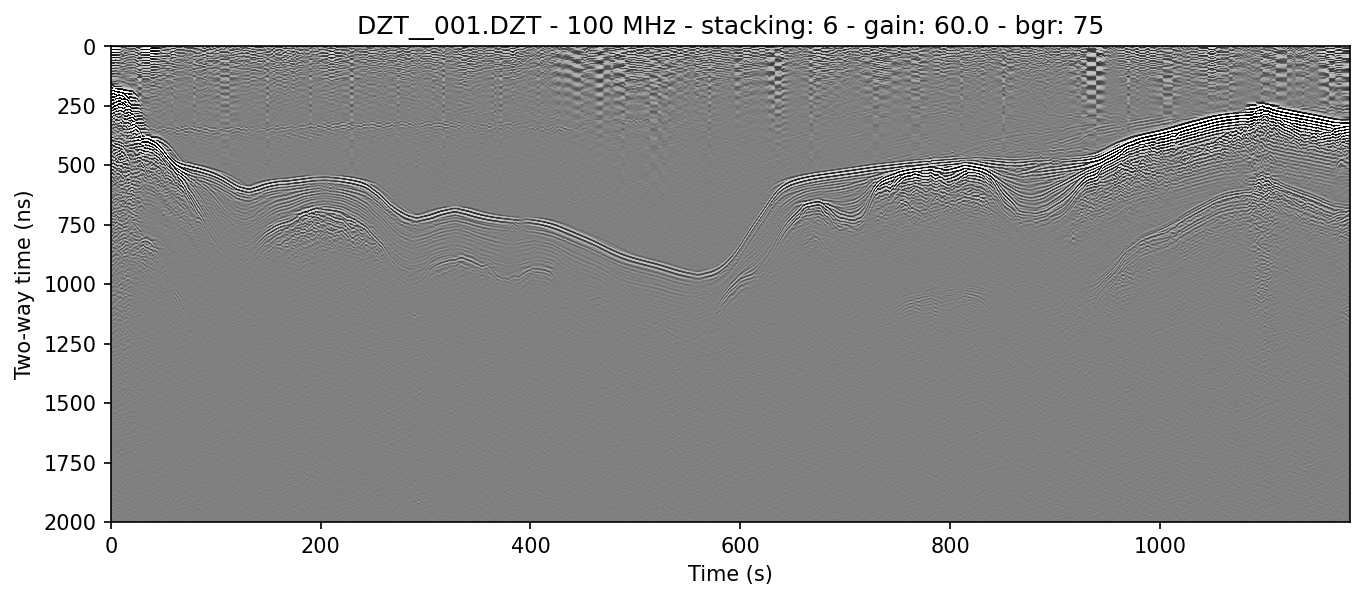
example 1B: with gain
readgssi -i DZT__001.DZT -o 1b.png -p 5 -s auto -g 60 -r 75
This will cause readgssi to create a plot from the same file, but matplotlib will save the plot as "1b.png" (-o 1b.png). The script will plot the y-axis size (-p 5) and automatically stack the x-axis to (-s auto). The script will plot the data with a gain value of 50 (-g 50), which will increase the plot contrast by a factor of 50. Next readgssi will run the background removal (-r 75) filter. (To apply this horizontal background removal filter across the entire line horizontally, use -r 0. More about this in examples 2b and 2c.)
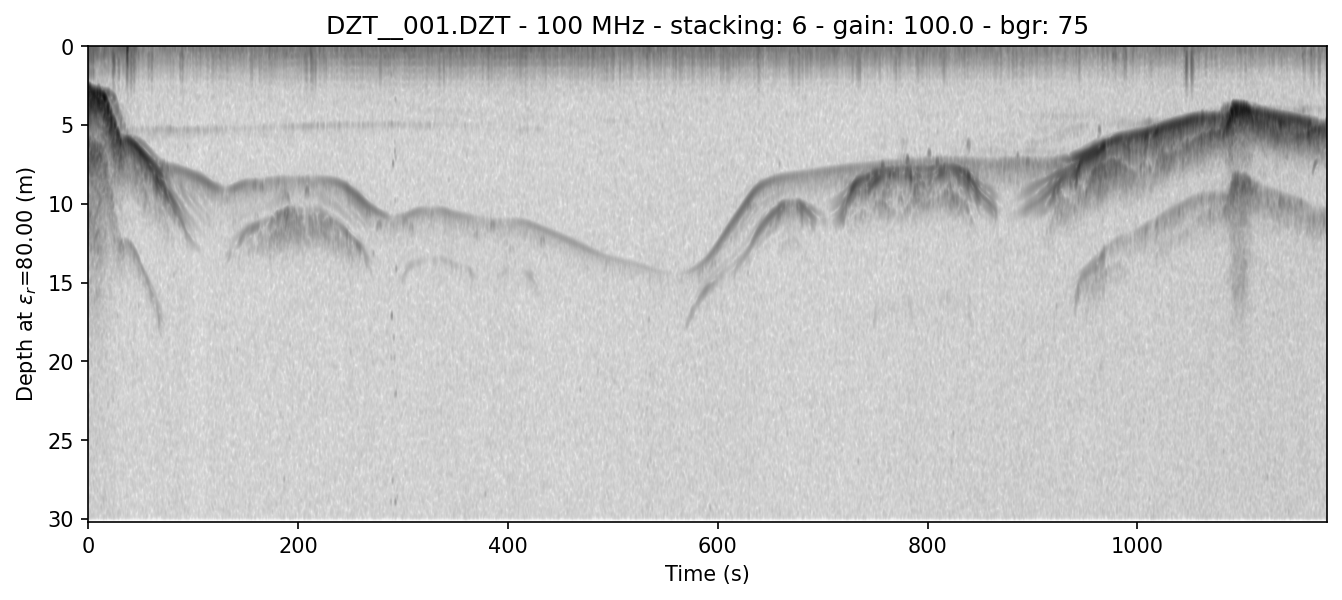
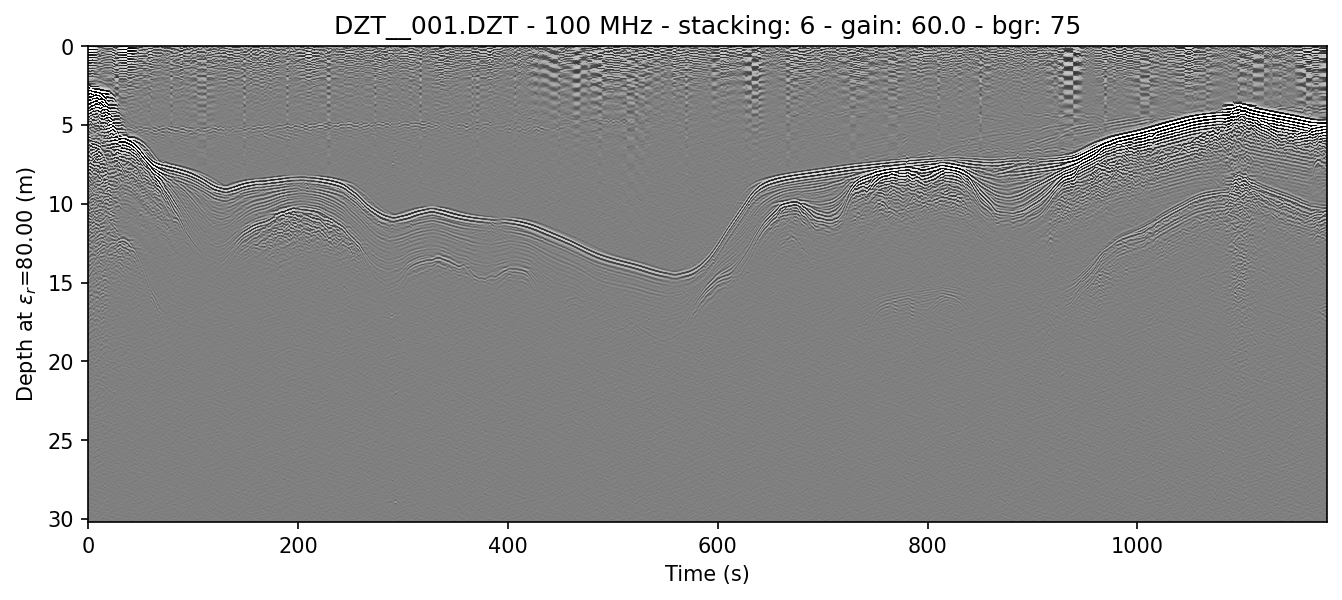
example 1C: display the vertical axis in meters and manually set the Epsilon_r
readgssi -i DZT__001.DZT -o 1c.png -p 5 -s auto -g 60 -r 75 -z m -E 80
This will set the Epsilon_r—that is, the relative permittivity of the first layer medium—to 80 (-E 80) and the Z axis to meters (-z m). The resulting plot will have a depth scale in meters.
Epsilon_r should be set in the header, but if it is set incorrectly, this is the way to adjust it on the fly.
example 1D: make a high-quality figure
readgssi -i DZT__001.DZT -o 1d.png -p 5 -s auto -g 60 -r 75 -z m -E 80 -T -D 300
Okay, now our figure looks great and we want to put it in a publication! Time to turn off the title (-T) and pump up the DPI to printer quality (-D 300). For AGU poster-quality images, you may even need -D 600.
One tricky thing to keep in mind is that the smaller the plot size (-p) the larger the relative size of the text in the figure.
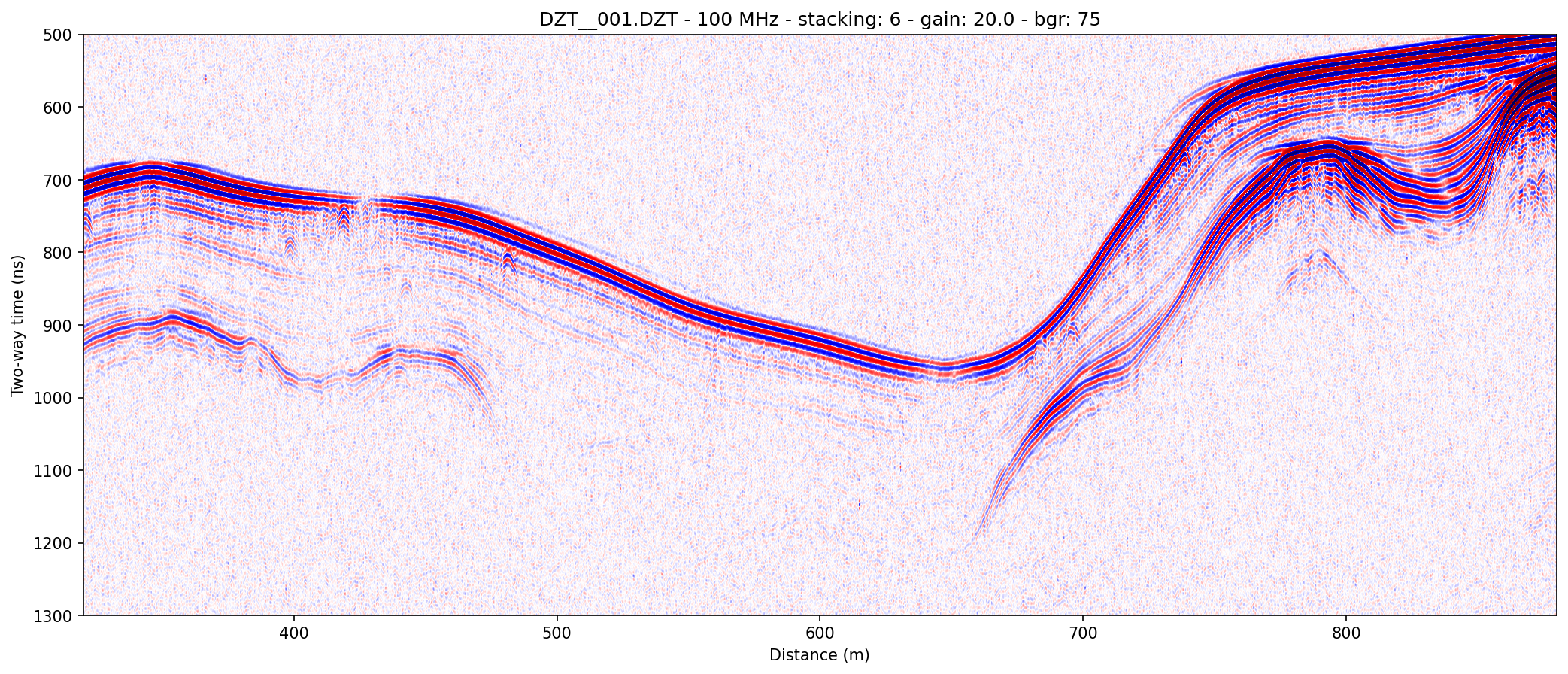
example 1E: changing colormaps and gain settings
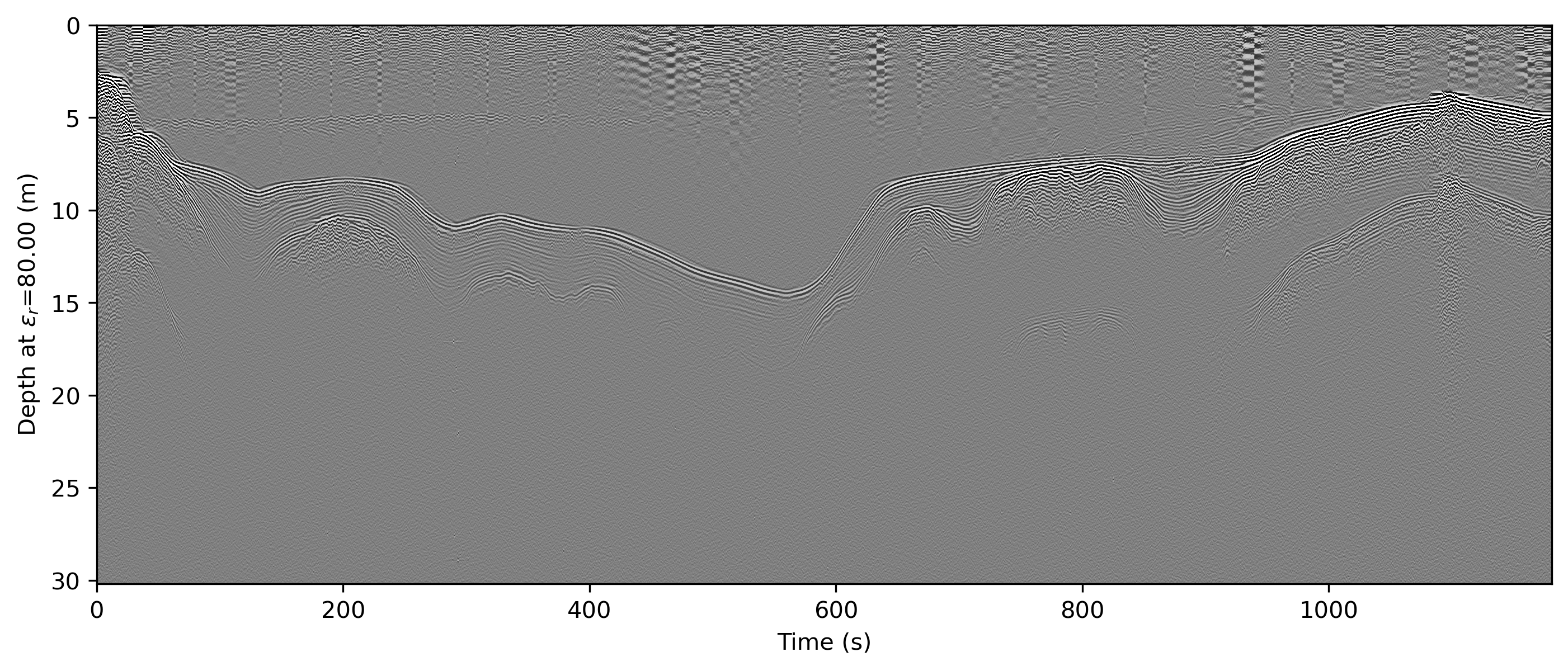
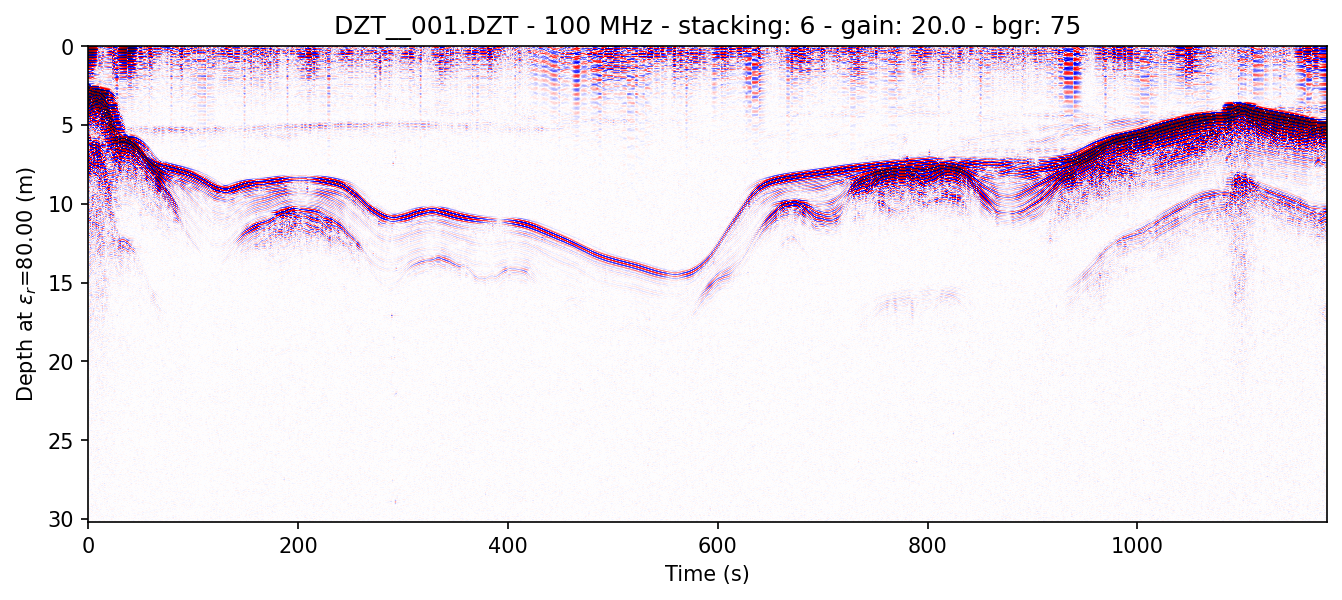
readgssi -i DZT__001.DZT -o 1e.png -p 5 -s auto -g 20 -r 75 -z m -E 80 -c seismic
Here, a horizontal background removal is applied, but gain is turned down (-g 20). The script uses matplotlib's "seismic" colormap (-c seismic) which is specifically designed for this type of waterfall array plotting. Even without gain, you will often be able to easily see very slight signal perturbations. Given its use of red, however, it is not terribly colorblind-friendly for either of the two most common types of human colorblindness, which is why it is not used as the default colormap.
example 1F: absolute value of vertical gradient
readgssi -i DZT__001.DZT -o 1f.png -p 5 -s auto -g 100 -r 75 -z m -E 80 -c gray_r -A -t 80-120
While we're on the topic of colormaps, it's worth pointing out that you can tell readgssi to calculate the vertical derivative (the "gradient") of the profile and display its absolute value using the -A flag. This gradient display is a good way to highlight areas of polarity change regardless of positive or negative values. It is particularly useful for highlighting glacial bed material through ice, for example. Here (in a lake profile) we set both -A to plot the absolute value of vertical gradient and also a FIR filter (-t 80-120) explained below. We also set the colormap to the reverse of the usual one (-c gray_r) so that black indicates high values.
filtering
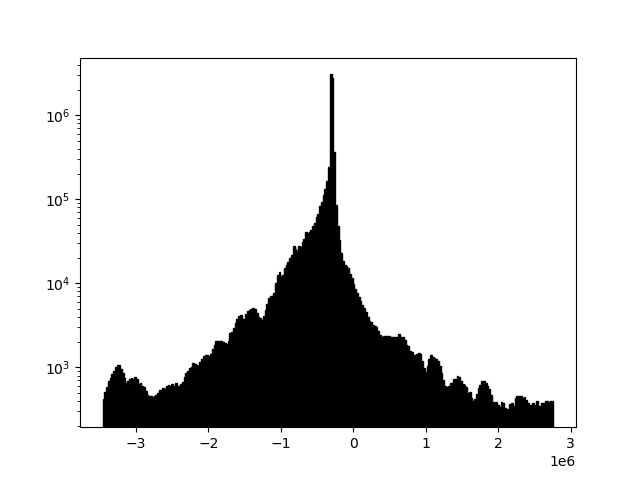
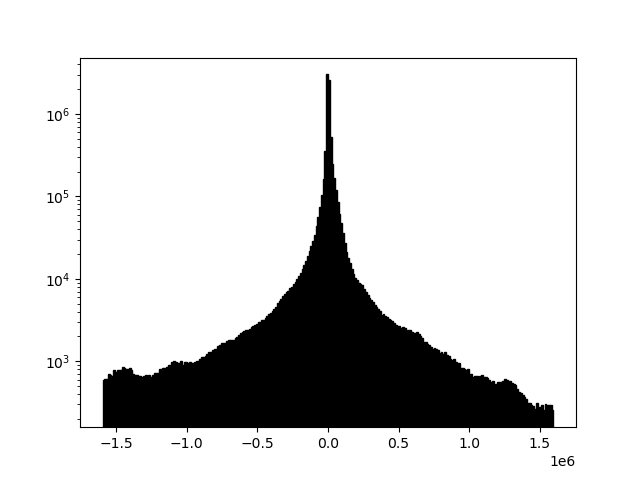
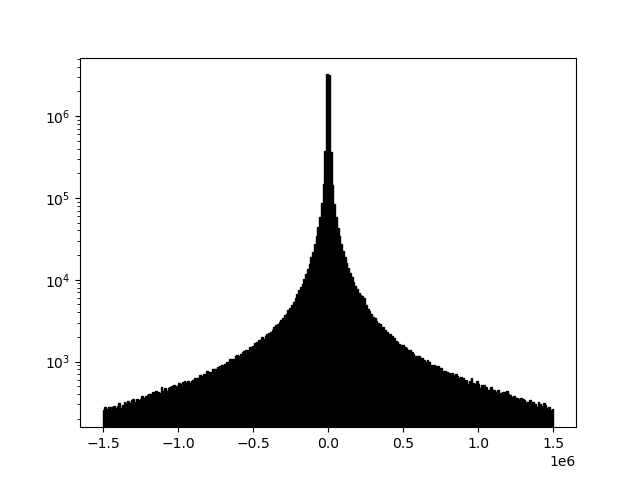
Each of the plots in this section is displayed with a histogram to illustrate changing array values.
example 2A: no background removal
readgssi -i DZT__002.DZT -o 2a.png -p 5 -s auto -g 30 -m
Sometimes, files will look "washed out" due to a skew relative to the mean of the data. This is easily correctable. Here readgssi will create a plot of size 5 and stack 5x (-p 5 -s 5). Matplotlib will use the default "gray" colormap and save a PNG of the figure, but the script will suppress the matplotlib window (using the -n flag, useful for processing an entire directory full of DZTs at once). Finally, the -m flag will draw a histogram for each data channel. Note how the histogram changes when filters are applied.
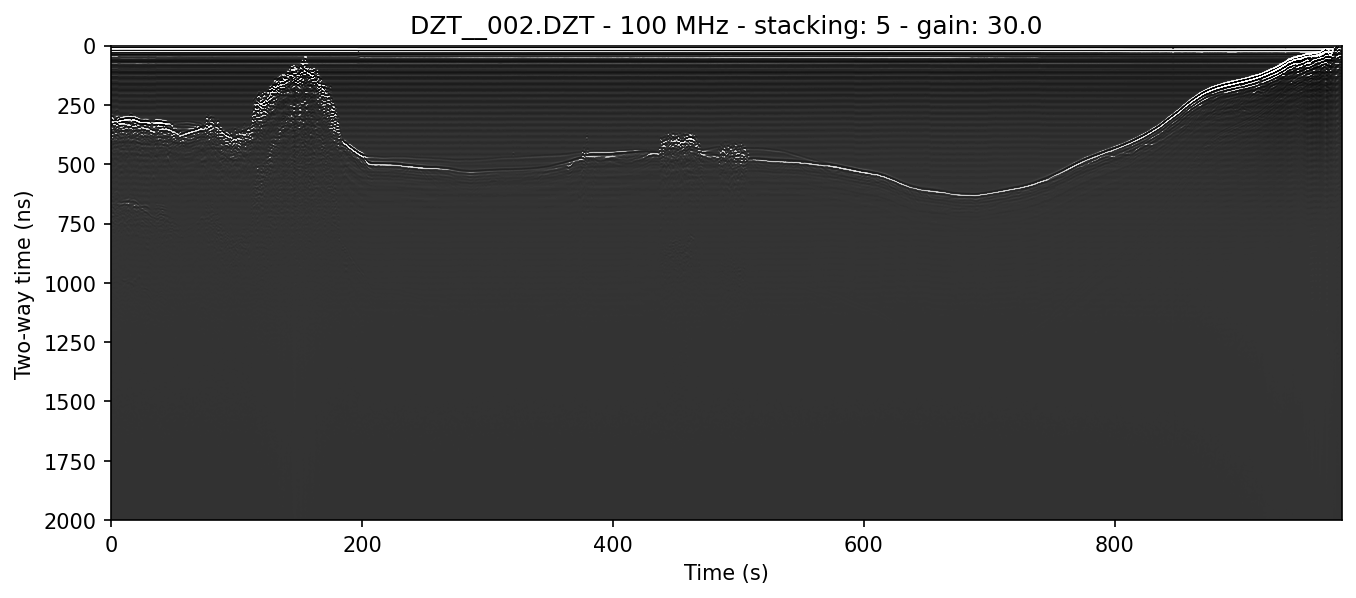
example 2B: horizontal mean BGR algorithm applied
The flag to get rid of the skew (or any horizontally uniform noise) is -r, also known as background removal or BGR for short. -r has two modes, one set by -r 0 and one set when the option after the -r flag is greater than zero. When this BGR option is zero, the program simply subtracts the average of each profile row from the array. When it's greater than 0, readgssi will implement a moving window mean, the size of which is set in post-stack traces (see example 2C).
The command below does the same thing as example 2A, except -r 0 applies full width horizontal mean background removal to the profile. Note the difference in ringing artifacts and skew between examples 2a and 2b.
readgssi -i DZT__002.DZT -o 2b.png -p 5 -s auto -g 30 -m -r 0
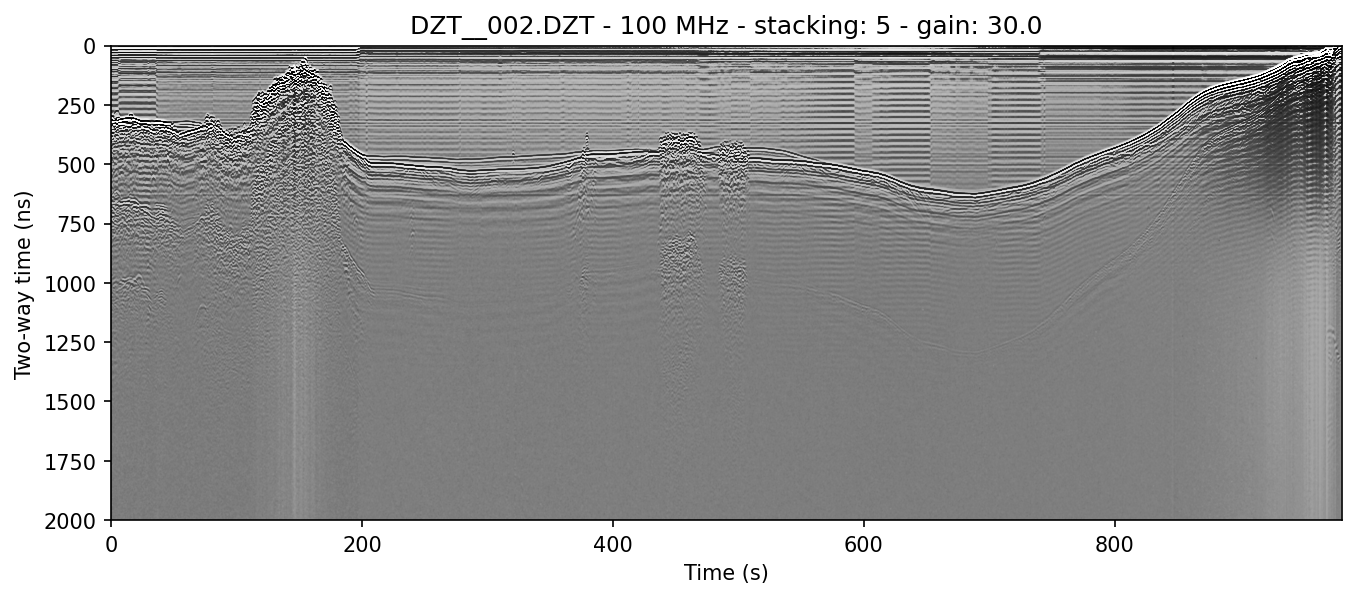
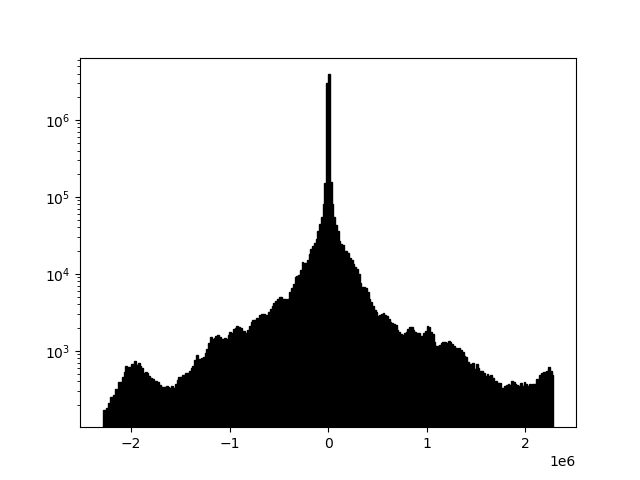
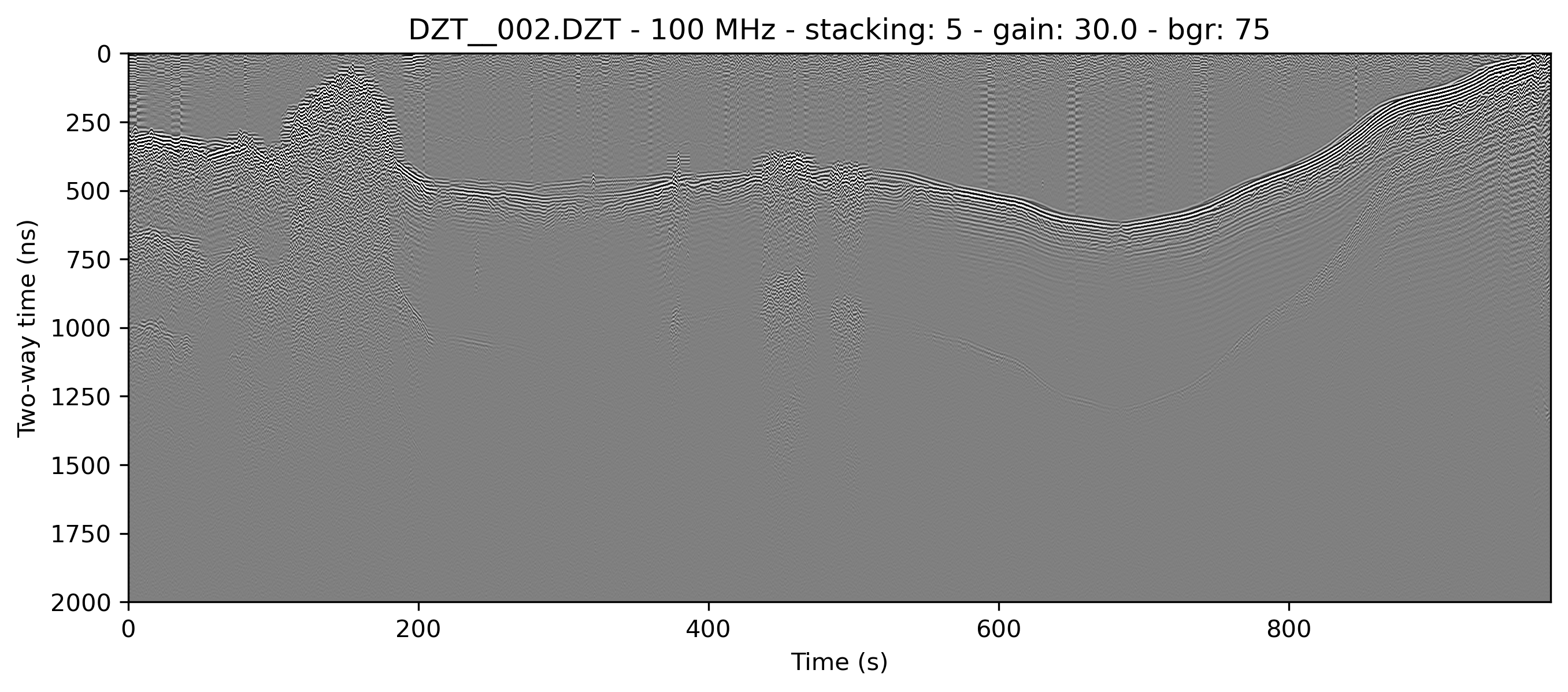
example 2C: moving window horizontal mean
readgssi -i DZT__002.DZT -o 2c.png -p 5 -s auto -g 30 -m -r 75
Same as above but with a 75-trace wide moving window mean (-r 75). This width represents post-stack traces. This is, for all intents and purposes, the same as RADAN's "BOXCAR" method of horizontal noise removal, but much, much faster because it's implemented with scipy. Areas beyond the left and right edges are treated as zeros. Notice that the noise in the water column is nearly entirely wiped out, but real data is extended with lateral wisps the size of half of the window, which is a side-effect of this method. Note that the histogram (-m) has a fairly even distribution around the mean, which generally indicates that the image should be fairly readable.
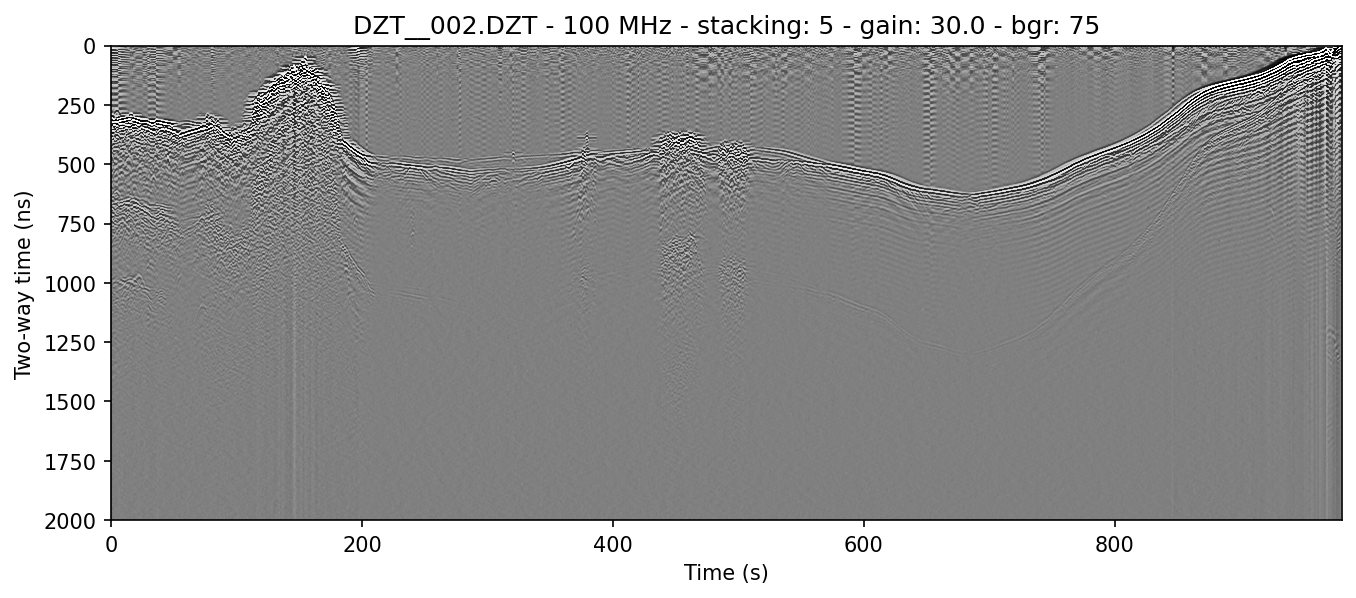
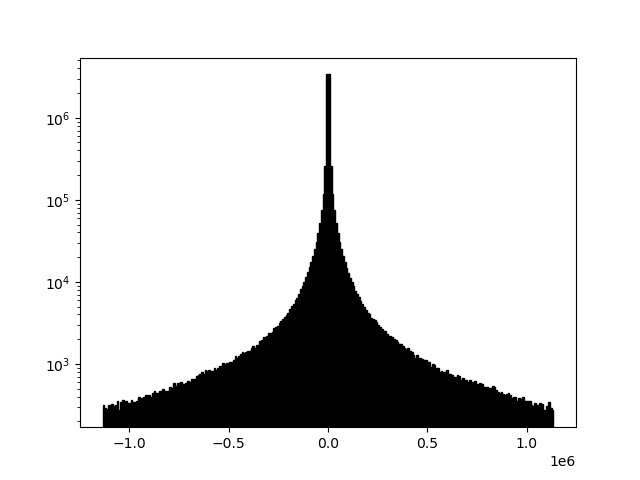
example 2D: vertical triangular FIR filter
readgssi -i DZT__002.DZT -o 2d.png -p 5 -s auto -g 30 -m -t 80-120
This function is functionally the same as RADAN's vertical triangular FIR filter, but again implemented with scipy. It is used to filter out unwanted frequencies on a per-shot basis, which makes it very effective as a profile cleaning tool. The most effective results are achieved by filtering a tight window of frequencies around the center frequency of the antenna you are using. In this case, we are using a 100 MHz antenna and filtering from 80-120 MHz.
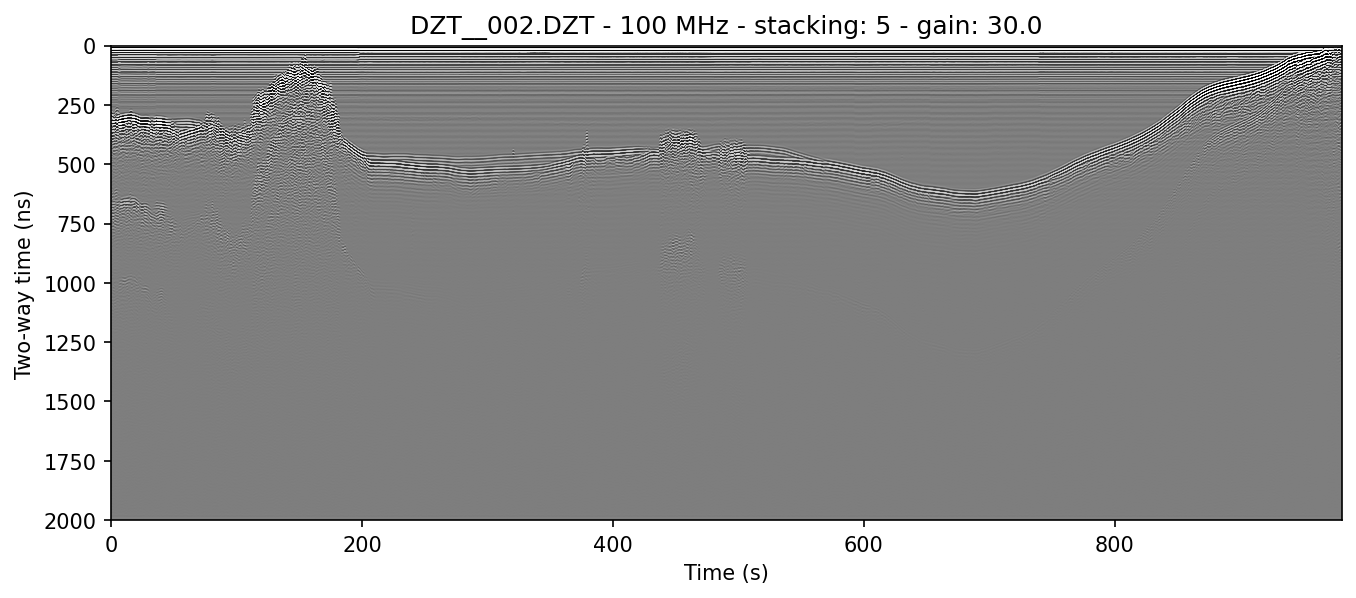
example 2E: combining filters for a cleaner profile
readgssi -i DZT__002.DZT -o 2e.png -p 5 -s auto -g 30 -m -r 75 -t 80-120
In this example, we use two concurrent filters (the horizontal windowed BGR and the vertical triangular FIR) to try and clean the image up as much as possible without overcooking it.
array manipulation
example 3A: (without) distance normalization
The default behavior of readgssi is to plot the X-axis in survey time units (seconds). This can be changed using the -x flag. To display in distance units, you must either have GPS information in DZG format, or specify the number of radar traces per meter using the -d flag. -d 24 -x meters will change the traces per meter value in the header to 24.0 and display the profile with distance in meters along the X-axis.
Files with GPS information are handled in a slightly different way. First, readgssi will read a DZG file to create an array of distance information associated with marks in the DZT. (NOTE: If your project was recorded without DZG files but you have per-line GPS mark information in GPX format, please look at gpx2dzg for a method of creating a DZG file for each survey line) After reading the DZG, the program will expand or contract the GPR array based on the speed over ground between GPS points. It will then modify the traces per meter value from the header and display the profile with distance on the X-axis.
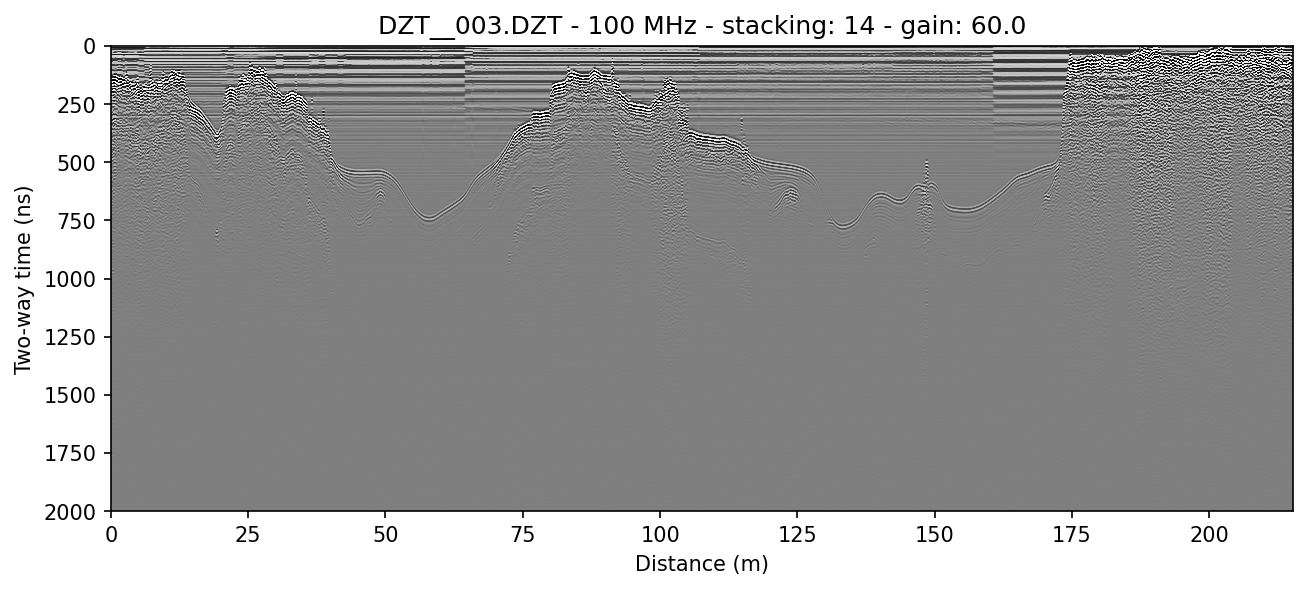
Here a file is processed and displayed without distance normalization:
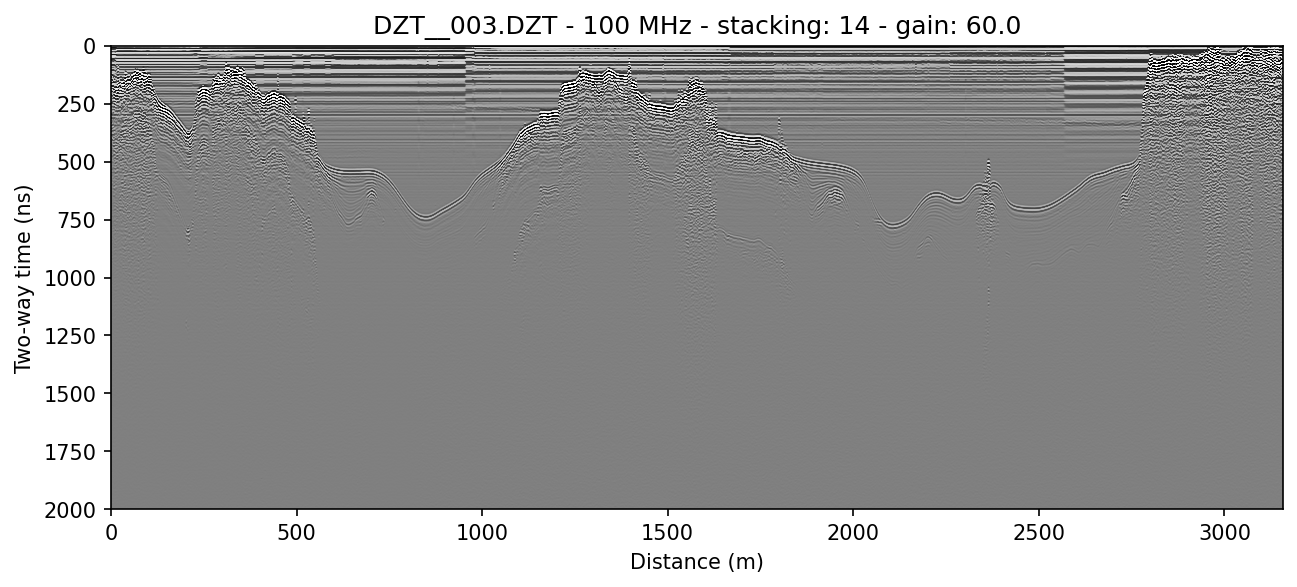
readgssi -i DZT__003.DZT -o 3a.png -p 5 -s auto -r 0 -g 60
example 3B: distance normalization using a DZG file
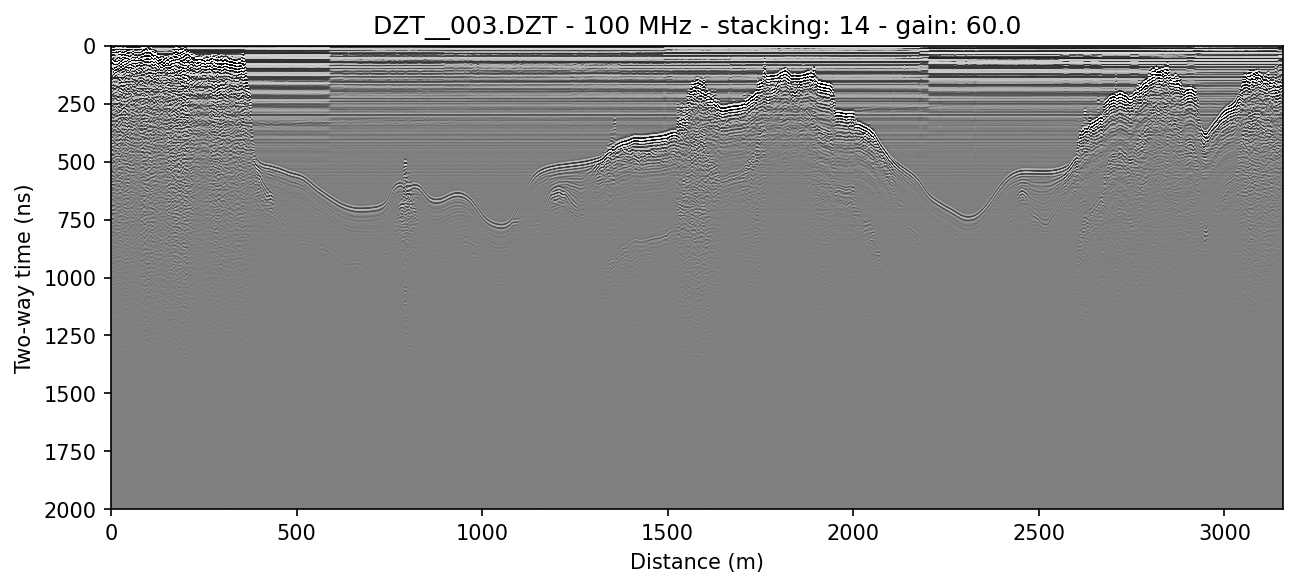
To use DZG GPS information to distance normalize the profile and display in meters traveled, use the -N and -x meters flags. readgssi will normalize the file in chunks to reduce memory usage. Here is the same file with distance normalization applied:
readgssi -i DZT__003.DZT -o 3b.png -p 5 -s auto -r 0 -g 60 -N -x meters
example 3c: reverse line direction
A simple but sometimes necessary operation is flipping a survey line back to front in order to display it congruently with others in a set. This can be done easily using -R.
advanced usage
UNIX users have a distinct advantage of being able to easily process entire folders full of DZTs with a simple command. Users who wish to do this should read up on how to construct for loops in Bash or simply follow and modify these examples below.
processing all files in a folder
This command makes use of the ls function in Bash, which lists all files that match a specific pattern. In this case, we want the pattern to be "any DZT file," which ends up being simply ls *.DZT (the * symbol is a wildcard, meaning it matches any set of characters, so in this case it would match both FILE____005.DZT and test.DZT but not Test01.dzt because the .DZT is case sensitive.).
for f in `ls *.DZT`; do readgssi -p 8 -n -r 0 -g 40 -Z 233 -z ns -N -x m -s auto -i $f; done
The structure of this command is easy to understand if you know a little bit about for loops. This command loops over every file with the extension .DZT (ls *.DZT where * indicates a wildcard) and assigns the filename to the f variable on each loop. Then, after the semicolon, bash runs readgssi for every pass of the loop. In this case, the parameters are:
-p 8 # plot with size 8
-n # suppress the matplotlib window; useful if you do not want the operation interrupted by plot windows
-r 0 # full-width background removal
-g 40 # gain of 40
-Z 233 # time zero at 233 samples
-z ns # display the depth axis in nanoseconds
-N # distance-normalize the profile
-x m # display the x-axis in meters
-s auto # apply automatic stacking
-i $f # recall the `f` variable containing this loop's filename and feed it to the input flag of readgssi
Finally, end the loop by closing the command with a linebreak ;, and the done marker.
processing specific sets of files
You can make the command even more specific by further modifying the set of files returned by the ls command. For example:
for f in `ls FILE__{010..025}.DZT`; do readgssi -p 8 -n -r 0 -g 40 -Z 233 -z ns -N -x m -s auto -i $f; done
This command will process only the 16 files in the numeric sequence between and including 010 and 025 in the set (FILE__010.DZT, FILE__011.DZT, ..., FILE__025.DZT). bash handles the zero padding for you as well. Pretty cool.
contributors
- Ian Nesbitt (@iannesbitt, author)
- François-Xavier Simon (@fxsimon)
- Thomas Paulin (@thomaspaulin)
- teshaw (@teshaw)
citation suggestion:
Ian M. Nesbitt, François-Xavier Simon, Thomas Paulin, 2018. readgssi - an open-source tool to read and plot GSSI ground-penetrating radar data. doi:10.5281/zenodo.1439119
known bugs:
- scipy 1.2.x causes errors when filtering. use scipy 1.3.0 to avoid.
future
- automatic script testing for smoother dev
- create a class for surveyline objects, similar to
obspy.core.trace.Trace - GPS transcription from CSV with fields like
mark name, lon, lat, elev, time - ability to use GPS altitude to adjust z position across profile
- ability to ground truth depths as horizontal lines
- GUI-based geologic/dielectric layer picking
- layer velocity calculation (using minimum of clustered hyperbola tail angle measurements, or manual input)
- velocity-based depth adjustments
- ability to incorporate ground truth measurements
- velocity gradient/angle of incidence-based array migration
- translation to common geophysical formats (HDF5, SEGY, etc.)
- integration with
vistafor 3D visualization of location-aware arrays
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file readgssi-0.0.22.tar.gz.
File metadata
- Download URL: readgssi-0.0.22.tar.gz
- Upload date:
- Size: 69.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.7.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
74d7b68cbecd008e3bd0b03136d148393e420b44ac8d4c2dcc0ab1071e925f32
|
|
| MD5 |
a5530520ed853c81927e7ee9ff068eae
|
|
| BLAKE2b-256 |
40f7ac395e81fa9264614990011e1495cc7dfd45c156056b9f11fe860d4b2d15
|
File details
Details for the file readgssi-0.0.22-py3-none-any.whl.
File metadata
- Download URL: readgssi-0.0.22-py3-none-any.whl
- Upload date:
- Size: 67.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.7.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0513759a2db734b6560d7937ad34d6d85440184392f4c31126f8b02ed98e82ee
|
|
| MD5 |
3958bb4b7f733f2aa3961c88dc35a438
|
|
| BLAKE2b-256 |
f5da0ee5d945f4f22ae82824945922bbcf41bf2c5336b37e1c7f881cdda8c852
|