Retrosynthesis, visualization and machine learning tools
Project description

- RetroChem -
📜 Package information
RetroChem, is a one-step retrosynthesis engine pip installable python package, which is based on SMILES inputs. It includes tools for model training, template preprocessing, and visualization of reaction predictions. Developped as part of a project for the practical Programming in Chemistry at EPFL (2025).
🪄 Features
- Simple to use retrosynthesis engine.
- Draw molecules or use molecule name (eg. paracetamol).
- Gives clean and chemically correct molecules.
- Works well for known molecules (eg. GHB, EDTA and etc).
- Predicts possible reactants and reaction steps.
- Gives the predictions in order of confidence
👥 Contributors
- Giulio Matteo Garotti, second year chemical engineer at EPFL
- Florian Follet, second year chemist at EPFL
- Noah Paganzzi, second year chemical engineer at EPFL
- Jacques Maurice Grandjean, second year chemical engineer at EPFL
View contributors : Contributors on GitHub
View commit activity: Commit activity on GitHub
View code frequency : Code frequency on GitHub
🔍 Retrosynthesis, what is it?
Organic retrosynthesis is a problem-solving technique used in organic chemistry to design a synthetic route for a target molecule by breaking it down into simpler precursor structures. This process, known as retrosynthetic analysis, helps chemists plan the step-by-step synthesis of complex organic compounds by working backward from the desired product.
The approach involves two main stages:
Disconnection: The target molecule is “disconnected” at strategic bonds to identify simpler molecules that could be used as starting materials. These disconnections are guided by known chemical reactions and functional group transformations. Each disconnection leads to one or more synthons, which are idealised fragments representing the functional components of the molecule. For example, a carbon–carbon bond may be disconnected to yield a nucleophile and an electrophile.
Reagent Identification: Once the synthons are defined, they are translated into real, purchasable or synthesizable compounds known as synthetic equivalents. These are then used to build the molecule in the forward direction. This step requires the selection of appropriate reagents, protecting groups, and reaction conditions to ensure a feasible and efficient synthesis.
A retrosynthetic tree is often constructed to visualise all possible synthetic pathways, with the target molecule at the top and potential starting materials branching below. Chemists use both strategic bonds and known reaction patterns (like aldol condensations, Grignard additions, or Diels-Alder reactions) to inform these choices.
- Retrosynthesis is widely applied in:
- Pharmaceutical chemistry, for the development of active pharmaceutical ingredients (APIs).
- Natural product synthesis, for recreating complex bioactive compounds from simpler building blocks.
- Material science, for designing functional molecules with specific properties.
- Green chemistry, to plan more sustainable and efficient synthetic routes.
The logic of retrosynthesis also lends itself well to computational chemistry. Algorithms and machine learning tools are increasingly used to automate retrosynthetic analysis, generating synthetic routes based on reaction databases and predictive models.
The theoretical underpinning of retrosynthesis often involves mapping the retrosynthetic steps to known reaction mechanisms and using heuristics based on reactivity and selectivity. The goal is to find the shortest, most cost-effective, and most reliable pathway to the target compound.
Let us now walk through how to apply retrosynthetic analysis to a real molecule using this framework!
🕹️ How to install it
Firstly it is advised to create a CONDA environment:
#Open bash or terminal
#Name the environment as you wish and specify python 3.10
conda create -n env.name python==3.10
#Activate your environment
conda activate env.name
#Alternative way of activating it
source activate env.name
Clone this repository and install the python package locally:
#Clone the repository
git clone https://github.com/Flo-fllt/RetroChem.git
#Naviguate to the RetroChem folder
cd RetroChem
#Install the package locally in editable mode, make sure to activate your environment before doing so
pip install retrochem
#This will install the retrosynth package on your machine. You can now run the program with:
retrochem
The streamlit web page for RetroChem will be opened on your default browser.
💡 Requirements
python>=3.10
streamlit
pandas
numpy
joblib
rdkit
scikit-learn
matplotlib
pillow
streamlit-ketcher
💻 Guide
Once the RetroChem web page is open the retrosynthesis engine is ready to run!
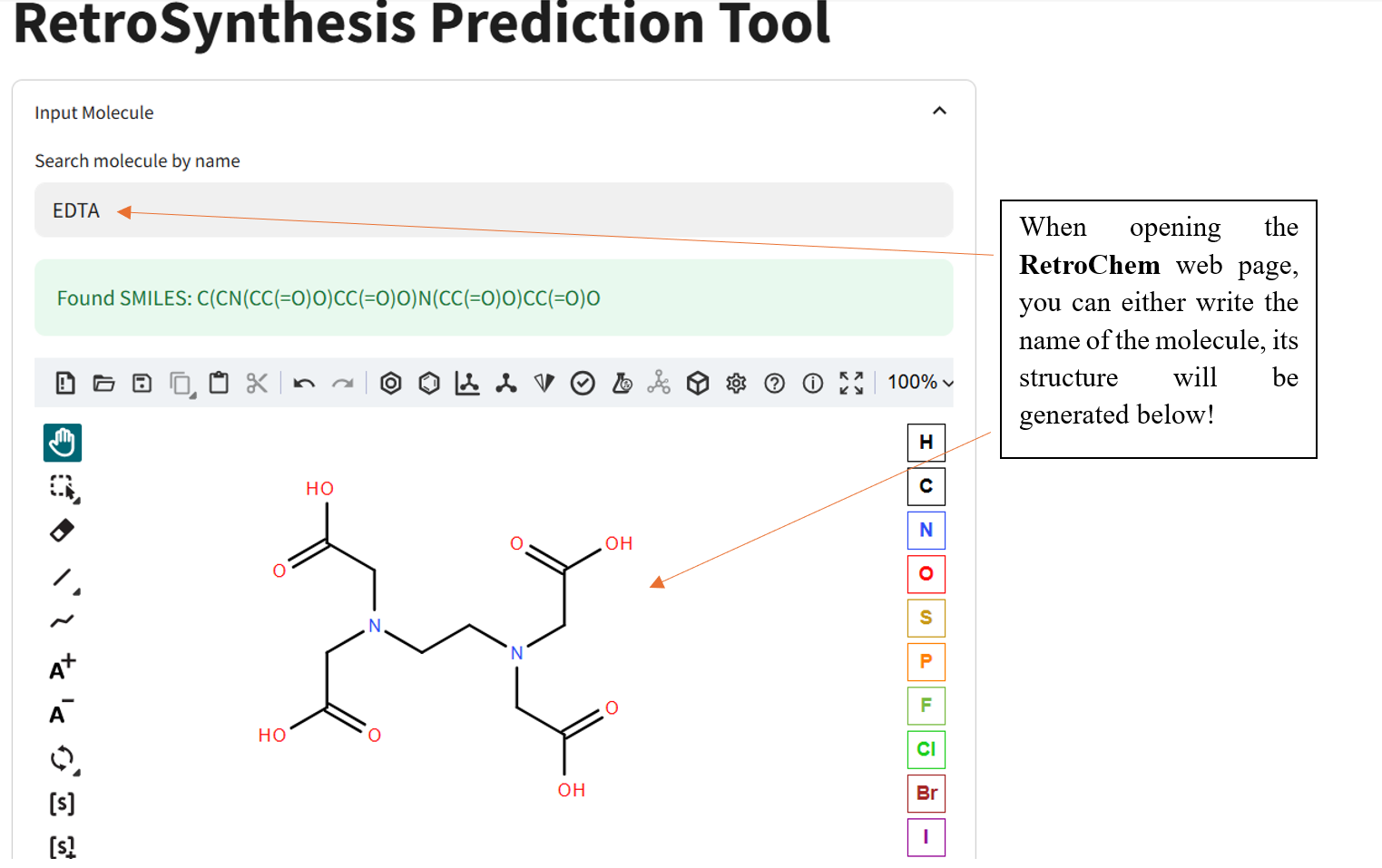
If the program has a difficult time finding reactants, try another molecule and especially known molecules.
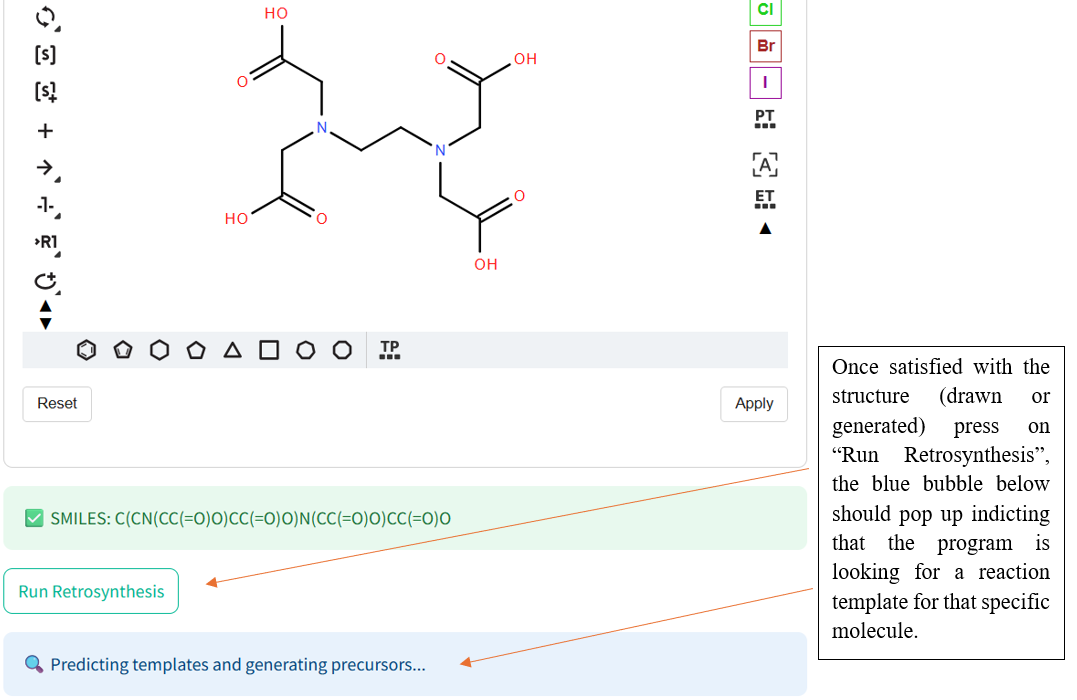
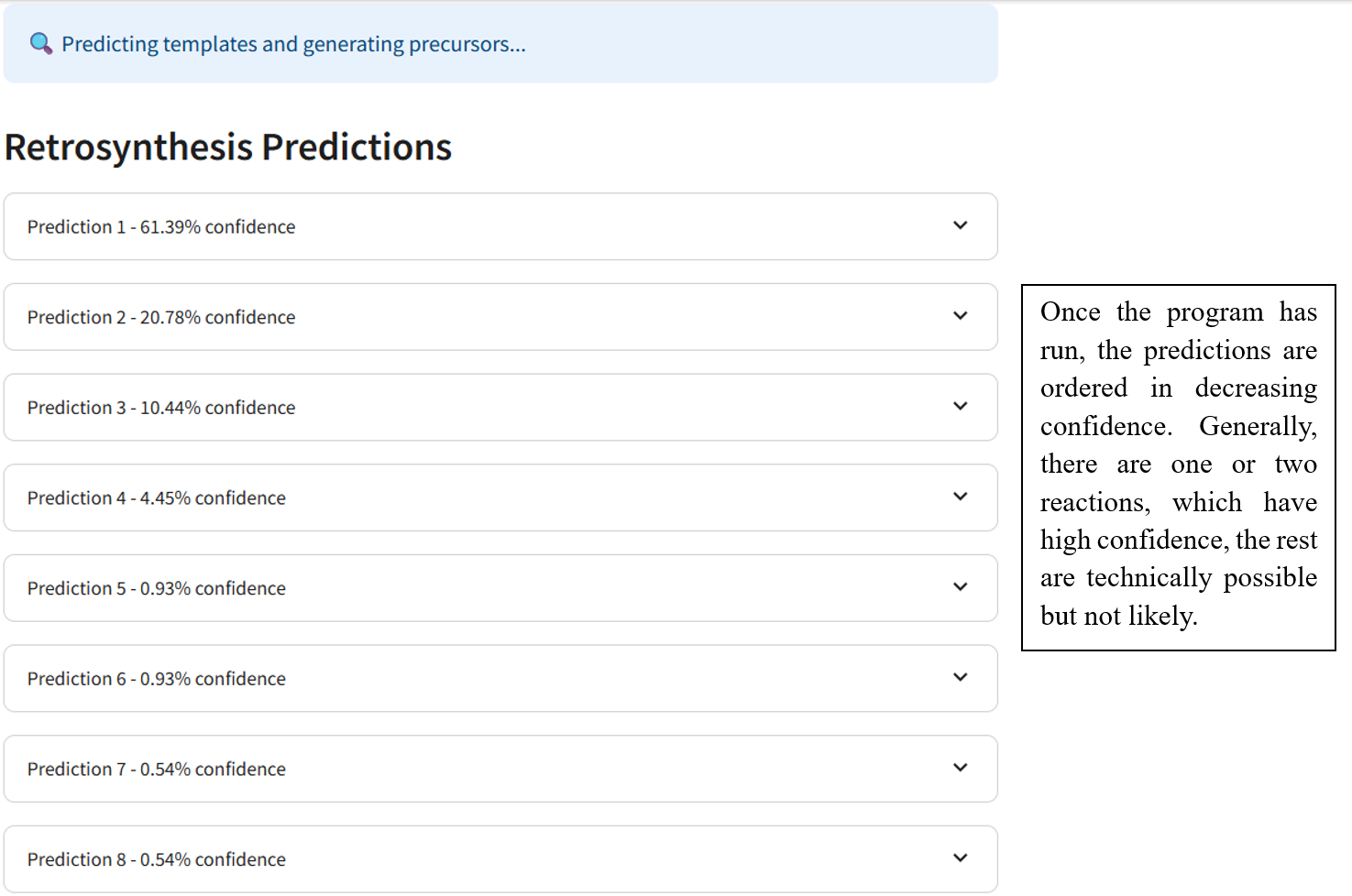
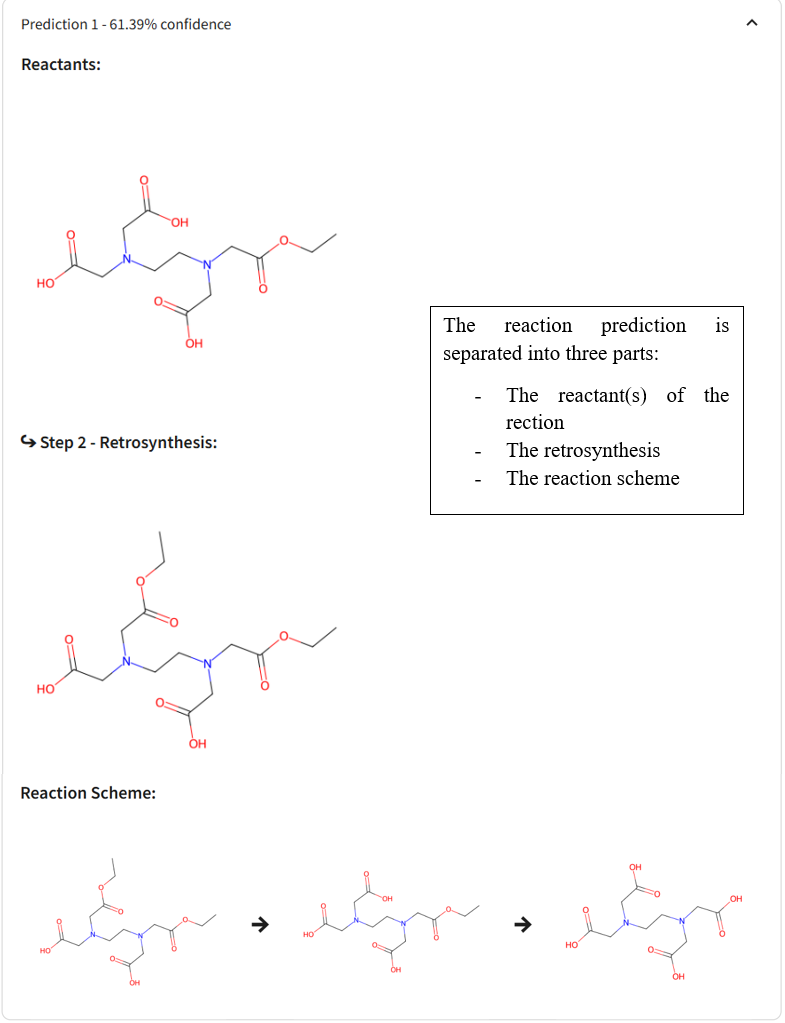
🛟 Need help?
If for some reason the program does not work or an issue occurs during the search of retrosynthesis templates there are a few steps you can take to ensure everything is in order.
First make sure that you are in the correct environment where you installed the package:
#Show what current environnement you are in
which python
If not:
#Activate the environnement where you downloaded the retrochem package
conda activate env.name
Then navigate to the top-level RetroChem repository folder:
cd retrochem
# Confirm your current location
pwd
# It should end with: /your/path/to/retrochem
❗️ Warning: If your path ends in /retrochem/retrochem, you are one level too deep, go back using:
cd ..
Thirdly check that you have the latest version of RetroChem
#Shows you which version of retrochem you are using
pip show retrochem
#Updates the package
pip install --upgrade retrochem
#If the issue persists try uninstalling the package, then installing it again with the latest version specified
pip uninstall retrochem
pip install retrochem==x.x.x #change the x.x.x to the current version (check PyPi page below)
You can compare the version you have downloaded to that of the newest available version, which can be found on the PiPy page:
If for some reason the issue is not resolved, it may derive from a pip issue, in this case checking for a pip update may solve it:
#For a virtual environment
pip install --upgrade pip
#For Linux/MacOS
python3 -m pip install --upgrade pip
For questions on how to set-up/download the necesseties (ANACONDA and etc) we welcome you to take a look at this:
🤖 How to retrain our model
To retrain the retrosynthesis prediction model, use the notebook located in:
retrain_model/Train_model.ipynb
By default, this notebook is configured to work with the original USPTO-50K dataset.
- Retrain with the default dataset
- Simply run the two cell blocks in the notebook:
- The first one combines the three original datasets (train, valid, and test) into one file.
- The second cell block loads the combined file, retrains the model, and saves:
retrain_model/new_mlp_classifier_model.pkl
retrain_model/new_scaler.pkl
retrain_model/new_label_encoder.pkl
- Retrain with your own data (not recommended)
- Skip the first cell block
- Update the file path in the second cell block:
# Replace this path with your own
combined_file_path = os.path.join(data_dir, "your_data.csv")
- Ensure your CSV includes the following required columns:
RxnSmilesClean, PseudoHash, RetroTemplate, TemplateHash
Disclaimer: Retraining with custom data is not recommended unless you are familiar with the dataset structure and preprocessing requirements. The model was trained with curated USPTO-50K data, and using inconsistent or improperly formatted data may lead to poor performance or errors.
📚 Want more information?
Here is the link to the report of the project, logging detailed information regarding the model we used and how it was trained along with the general context of this project.
🕺Your turn!
With all this information you should be more than ready to give RetroChem a try and don't forget to let us know what you think!
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file retrochem-0.2.0.tar.gz.
File metadata
- Download URL: retrochem-0.2.0.tar.gz
- Upload date:
- Size: 47.6 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a6fcc777cd45c8531833b063178311d1f5df28ad95d538cfc1915835b626f761
|
|
| MD5 |
45e02e7a44972e65262352c093119252
|
|
| BLAKE2b-256 |
71d84a176b6c22ae2226f4542cdf256d2630df2e17ef2a167d12df6532307053
|
File details
Details for the file retrochem-0.2.0-py3-none-any.whl.
File metadata
- Download URL: retrochem-0.2.0-py3-none-any.whl
- Upload date:
- Size: 44.8 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e13132010ff09c68a307b018a47950140085a10433dc86f564ada0bc68189a97
|
|
| MD5 |
33617c33c61ba3b4e7b76fced11a9bac
|
|
| BLAKE2b-256 |
295490bd97f895ec3c1f564dd1b3680a57fede5f90cf982a3071d6271c05b048
|






























