Causal Time Series Modeling of Supraglacial Lake Evolution in Greenland under Distribution Shift
Project description
RIC-TSC: Causal Time Series Modeling of Supraglacial Lake Evolution in Greenland under Distribution Shift
This repository provides the implementation for "Causal Time Series Modeling of Supraglacial Lake Evolution in Greenland under Distribution Shift" (Paper), which is accepted for publication in ICMLA 2025. We introduce a regionally-informed causal framework (RIC-TSC) that discovers lagged environmental drivers of supraglacial lake (SGL) evolution across Greenland and uses these causal signals for robust sequence modeling under spatial distribution shift.
Introduction
Supraglacial lakes (SGLs) exhibit complex spatiotemporal behaviors such as rapid drainage, slow drainage, refreezing, and burial. Accurate classification of lake evolution is critical to understanding meltwater runoff and ice sheet stability.
This repository presents a causally-informed modeling framework that identifies invariant environmental drivers across Greenland using Joint PCMCI+ (J-PCMCI+), and also captures region-specific causal mechanisms in individual basins. These causal predictors are then used in downstream sequence modeling to improve robustness and generalization under distribution shifts. We assess performance in global, in-distribution (ID), and out-of-distribution (OOD) settings.
Methodology
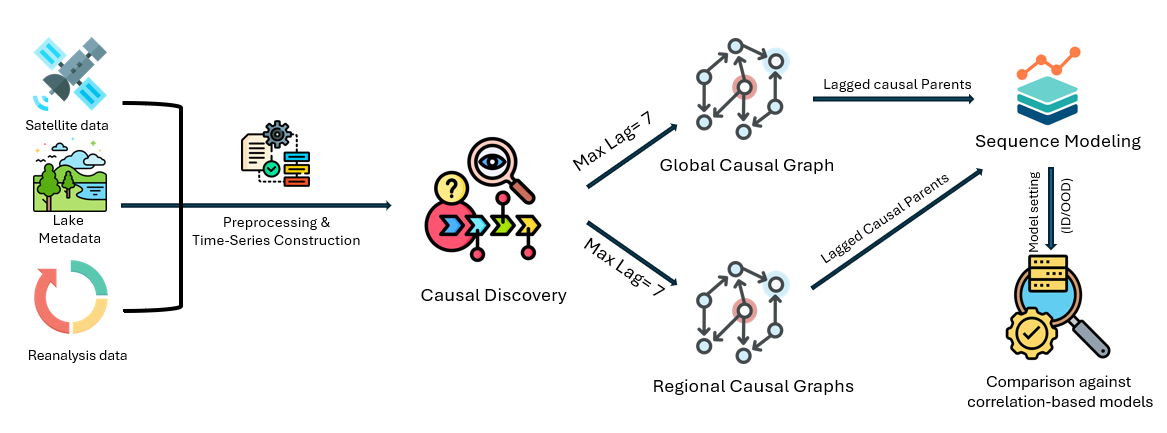
We construct daily multivariate time series from satellite and reanalysis sources:
- Sentinel-1 SAR (HV backscatter anomaly)
- Sentinel-2 and Landsat-8 optical imagery (NDWI-based water fraction, solar zenith)
- CARRA-West reanalysis (temperature, humidity, pressure, SST, etc.)
J-PCMCI+ is applied globally and per region to identify lagged causal parents of HV_anom (horizontally transmitted, vertically received backscatter anomaly), a proxy for lake water presence. These causal features are then used for lake evolution classification.
Installation
Install the package in editable mode for development:
git clone [https://github.com/ehfahad/RIC-TSC.git](https://github.com/ehfahad/RIC-TSC.git)
cd RIC-TSC
pip install -e .
Directory Structure
RIC-TSC/
├── src/rictsc/ # Core package logic
│ ├── utils/ # Refactored helper functions
│ ├── preprocessing.py # Preprocessing module
│ ├── causality.py # Causal feature module
│ └── classification.py # RICTSCClassifier API
├── causality/ # J-PCMCI+ causal discovery notebooks
├── data/ # Raw, processed, and causal datasets
├── figures/ # Methodology diagrams and experiment visualizations
├── results/ # Output metrics, confusion matrices, GMM plots
├── tests/ # Package sanity tests
├── pyproject.toml # Package metadata and dependencies
├── run_global_classification.py # Global pooled classification script
└── run_regionwise_classification.py # Region-wise ID and OOD classification script
Quickstart
1. Command Line Interface
Run the pipeline directly from the terminal using the installed entry points:
# Step 1: Preprocess time series for all lakes
rictsc-preprocess
# Step 2: Extract region-specific causal datasets
rictsc-causal
2. Python API
Integrate the RIC-TSC classifier into your own scripts:
from rictsc import RICTSCClassifier
import pandas as pd
# Initialize the classifier
model = RICTSCClassifier(seed=42)
# Load data and fit model on causal features
df = pd.read_csv("data/region_causal_datasets/CW_causal_timeseries.csv")
model.fit(df, feature_cols=["HV_anom_lag1", "S2_water", "r2"], label_col="label")
# Predict on new sequences
predictions = model.predict(test_df, feature_cols=["HV_anom_lag1", "S2_water", "r2"])
Experiments
We evaluate RIC-TSC under three experimental settings:
- Global: Train/test on pooled lake data from all six regions using an 80/20 split stratified by region.
- In-Distribution (ID): For each region, an 80/20 train/test split is applied to that region’s lakes.
- Out-of-Distribution (OOD): Train on a single region and test on the remaining five, assessing generalization beyond the training domain.
Each setting compares two models:
- Causal Model: Trained only on the lagged causal parents discovered by J-PCMCI+ for each region.
- Baseline Model: Trained using all available features, with no causal feature selection or temporal lag filtering.
Performance is reported using overall accuracy, macro-averaged F1, precision, and recall.
Reproducing Results
To reproduce the experiments from the paper:
- Clone the repository:
git clone https://github.com/ehfahad/RIC-TSC.git - Run the experiment scripts:
python run_global_classification.py
python run_regionwise_classification.py
Output Structure
results/
├── global_classification/
│ └── global_classification_results.csv # Metrics for global experiment comparing causal vs. baseline models
│
├── region_specific_classification/
│ ├── id_results.csv # Region-wise ID results comparing causal vs. baseline models
│ └── ood_results.csv # OOD results where models are trained on one region and tested on the other five
Citation
Please cite the work as:
@inproceedings{hossain2025rictsc,
title={Causal Time Series Modeling of Supraglacial Lake Evolution in Greenland under Distribution Shift},
author={Emam Hossain and Muhammad Hasan Ferdous and Devon Dunmire and Aneesh Subramanian and Md Osman Gani},
booktitle={2025 International Conference on Machine Learning and Applications (ICMLA)},
year={2025},
organization={IEEE}
}
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file rictsc-0.1.3.tar.gz.
File metadata
- Download URL: rictsc-0.1.3.tar.gz
- Upload date:
- Size: 15.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b5d3eacd7633b60a7dd4777f6e66730760dfc7994c17efa50a6d2e224d7d4794
|
|
| MD5 |
86ea4132c3b0938f325a97e61b83c4a1
|
|
| BLAKE2b-256 |
c0451baecf4b8b7fa7557d9bd22c21f73e42c1525ab6d6c2ddf95eacb7525dca
|
File details
Details for the file rictsc-0.1.3-py3-none-any.whl.
File metadata
- Download URL: rictsc-0.1.3-py3-none-any.whl
- Upload date:
- Size: 15.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8b3bb5241126b1ae0f5f1e1689ba380fbdb5a2e1c8dce635b07d7e267b623b1f
|
|
| MD5 |
9a505a1394749163ebaeb21d1c92f4c6
|
|
| BLAKE2b-256 |
76539a66522b4c116d436a198946d552ea7df9d3de4b92475ccc51d40f9f6140
|











