Fast and memory-efficient ANN with a subset-search functionality
Project description

Reconfigurable Inverted Index (Rii): IVFPQ-based fast and memory efficient approximate nearest neighbor search method with a subset-search functionality.
Reference:
- Y. Matsui, R. Hinami, and S. Satoh, "Reconfigurable Inverted Index", ACM Multimedia 2018 (oral). [paper] [project]
Summary of features
 |
 |
|---|---|
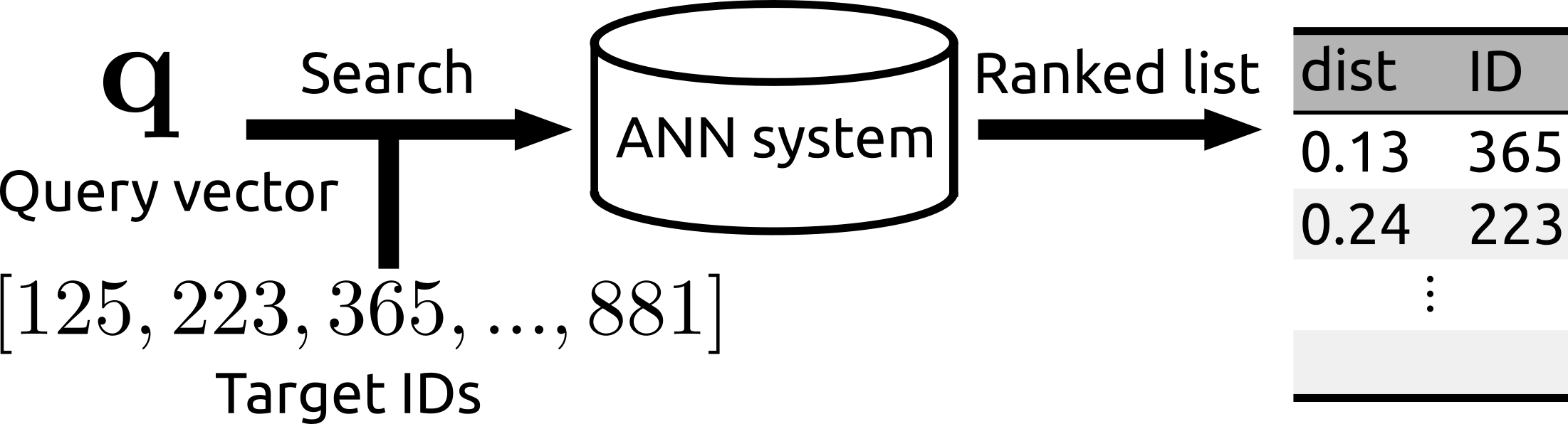
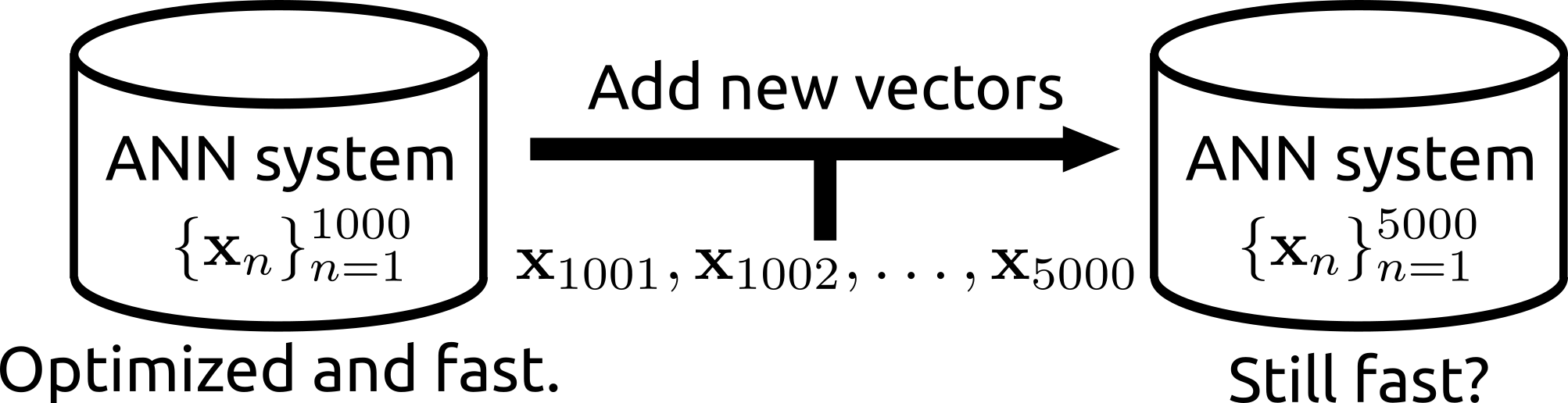
| The search can be operated for a subset of a database. | Rii remains fast even after many new items are added. |
- Fast and memory efficient ANN. Rii enables you to run billion-scale search in less than 10 ms.
- You can run the search over a subset of the whole database
- Rii Remains fast even after many vectors are newly added (i.e., the data structure can be reconfigured)
Installing
You can install the package via pip. This library works with Python 3.6+ on linux/mac/wsl/Windows10
pip install rii
For windows (maintained by @ashleyabraham)
Installing in Windows 10 via pip install
Requires MS Visual Studio Build tools C++ 14.0 or 14.1 toolset or above to compile and install via pip install
Pre-compiled binary for Windows 10
Pre-compiled binaries doesn't require MS Visual Studio Build tools
#Python 3.8
pip install https://github.com/ashleyabraham/rii/releases/download/v0.2.8/rii-0.2.8-cp38-cp38-win_amd64.whl
#Python 3.7
pip install https://github.com/ashleyabraham/rii/releases/download/v0.2.8/rii-0.2.8-cp37-cp37m-win_amd64.whl
OpenMP
OpenMP requires libomp140_x86_64.dll to compile in windows, which is part of MS Visual Studio Build tools and it is not redistributable.
In order to use OpenMP 3.0 /openmp:llvm flag is used which causes warnings of multiple libs loading, use at your descretion when used with other parallel processing library loadings. To supress use
SET KMP_DUPLICATE_LIB_OK=TRUE
SIMD
The /arch:AVX2 flag is used in MSVC to set appropriate SIMD preprocessors and compiler intrinsics
Documentation
Usage
Basic ANN
import rii
import nanopq
import numpy as np
N, Nt, D = 10000, 1000, 128
X = np.random.random((N, D)).astype(np.float32) # 10,000 128-dim vectors to be searched
Xt = np.random.random((Nt, D)).astype(np.float32) # 1,000 128-dim vectors for training
q = np.random.random((D,)).astype(np.float32) # a 128-dim vector
# Prepare a PQ/OPQ codec with M=32 sub spaces
codec = nanopq.PQ(M=32).fit(vecs=Xt) # Trained using Xt
# Instantiate a Rii class with the codec
e = rii.Rii(fine_quantizer=codec)
# Add vectors
e.add_configure(vecs=X)
# Search
ids, dists = e.query(q=q, topk=3)
print(ids, dists) # e.g., [7484 8173 1556] [15.06257439 15.38533878 16.16935158]
Note that you can construct a PQ codec and instantiate the Rii class at the same time if you want.
e = rii.Rii(fine_quantizer=nanopq.PQ(M=32).fit(vecs=Xt))
e.add_configure(vecs=X)
Furthermore, you can even write them in one line by chaining a function.
e = rii.Rii(fine_quantizer=nanopq.PQ(M=32).fit(vecs=Xt)).add_configure(vecs=X)
Subset search
# The search can be conducted over a subset of the database
target_ids = np.array([85, 132, 236, 551, 694, 728, 992, 1234]) # Specified by IDs
# For windows, you must specify dtype=np.int64 as follows.
# target_ids = np.array([85, 132, 236, 551, 694, 728, 992, 1234], dtype=np.int64)
ids, dists = e.query(q=q, topk=3, target_ids=target_ids)
print(ids, dists) # e.g., [728 85 132] [14.80522156 15.92787838 16.28690338]
Data addition and reconfiguration
# Add new vectors
X2 = np.random.random((1000, D)).astype(np.float32)
e.add(vecs=X2) # Now N is 11000
e.query(q=q) # Ok. (0.12 msec / query)
# However, if you add quite a lot of vectors, the search might become slower
# because the data structure has been optimized for the initial item size (N=10000)
X3 = np.random.random((1000000, D)).astype(np.float32)
e.add(vecs=X3) # A lot. Now N is 1011000
e.query(q=q) # Slower (0.96 msec/query)
# In such case, run the reconfigure function. That updates the data structure
e.reconfigure()
e.query(q=q) # Ok. (0.21 msec / query)
I/O by pickling
import pickle
with open('rii.pkl', 'wb') as f:
pickle.dump(e, f)
with open('rii.pkl', 'rb') as f:
e_dumped = pickle.load(f) # e_dumped is identical to e
Util functions
# Print the current parameters
e.print_params()
# Delete all PQ-codes and posting lists. fine_quantizer is kept.
e.clear()
# You can switch the verbose flag
e.verbose = False
# You can merge two Rii instances if they have the same fine_quantizer
e1 = rii.Rii(fine_quantizer=codec)
e2 = rii.Rii(fine_quantizer=codec)
e1.add_configure(vecs=X1)
e2.add_configure(vecs=X2)
e1.merge(e2) # Now e1 contains both X1 and X2
Examples
Author
Credits
- The logo is designed by @richardbmx (#4)
- The windows implementation is by @ashleyabraham (#42)
- Fixing the build (#50) and supporting for ARM CPUs (#58) by @timzag
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file rii-0.2.12.tar.gz.
File metadata
- Download URL: rii-0.2.12.tar.gz
- Upload date:
- Size: 84.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
85ccd0fb5a8a193990128843596bee2306792ff8b520d1f77c01de1f6bece50c
|
|
| MD5 |
de26e95aa81fe595244f3d239eae4cce
|
|
| BLAKE2b-256 |
8a6fc17fc238ae73e6c535e13461e9da489daf41b0036116a3951fa66858e0c8
|
Provenance
The following attestation bundles were made for rii-0.2.12.tar.gz:
Publisher:
publish.yml on matsui528/rii
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
rii-0.2.12.tar.gz -
Subject digest:
85ccd0fb5a8a193990128843596bee2306792ff8b520d1f77c01de1f6bece50c - Sigstore transparency entry: 208445288
- Sigstore integration time:
-
Permalink:
matsui528/rii@c0ed0a2b934cbca716f7d40c82aef70bc0cb4100 -
Branch / Tag:
refs/tags/v0.2.12 - Owner: https://github.com/matsui528
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@c0ed0a2b934cbca716f7d40c82aef70bc0cb4100 -
Trigger Event:
release
-
Statement type:















