A Python package for scalable network analysis and high-quality visualization.
Project description
RISK
RISK (Regional Inference of Significant Kinships) is a next-generation tool for biological network annotation and visualization. It integrates community detection algorithms, rigorous overrepresentation analysis, and a modular framework for diverse network types. RISK identifies biologically coherent relationships within networks and generates publication-ready visualizations, making it a useful tool for biological and interdisciplinary network analysis.
For a full description of RISK and its applications, see:
Horecka and Röst (2025), "RISK: a next-generation tool for biological network annotation and visualization".
DOI: 10.5281/zenodo.17257418
Documentation and Tutorial
Full documentation is available at:
- Docs: https://riskportal.github.io/risk-docs
- Tutorial Jupyter Notebook Repository: https://github.com/riskportal/risk-docs
Installation
RISK is compatible with Python 3.8 or later and runs on all major operating systems. To install the latest version of RISK, run:
pip install risk-network --upgrade
Key Features of RISK
- Broad Data Compatibility: Accepts multiple network formats (Cytoscape, Cytoscape JSON, GPickle, NetworkX) and user-provided annotations formatted as term–to–gene membership tables (JSON, CSV, TSV, Excel, Python dictionaries).
- Flexible Clustering: Offers Louvain, Leiden, Markov Clustering, Greedy Modularity, Label Propagation, Spinglass, and Walktrap, with user-defined resolution parameters to detect both coarse and fine-grained modules.
- Statistical Testing: Provides permutation, hypergeometric, chi-squared, and binomial tests, balancing statistical rigor with speed.
- High-Resolution Visualization: Generates publication-ready figures with customizable node/edge properties, contour overlays, and export to SVG, PNG, or PDF.
Example Usage
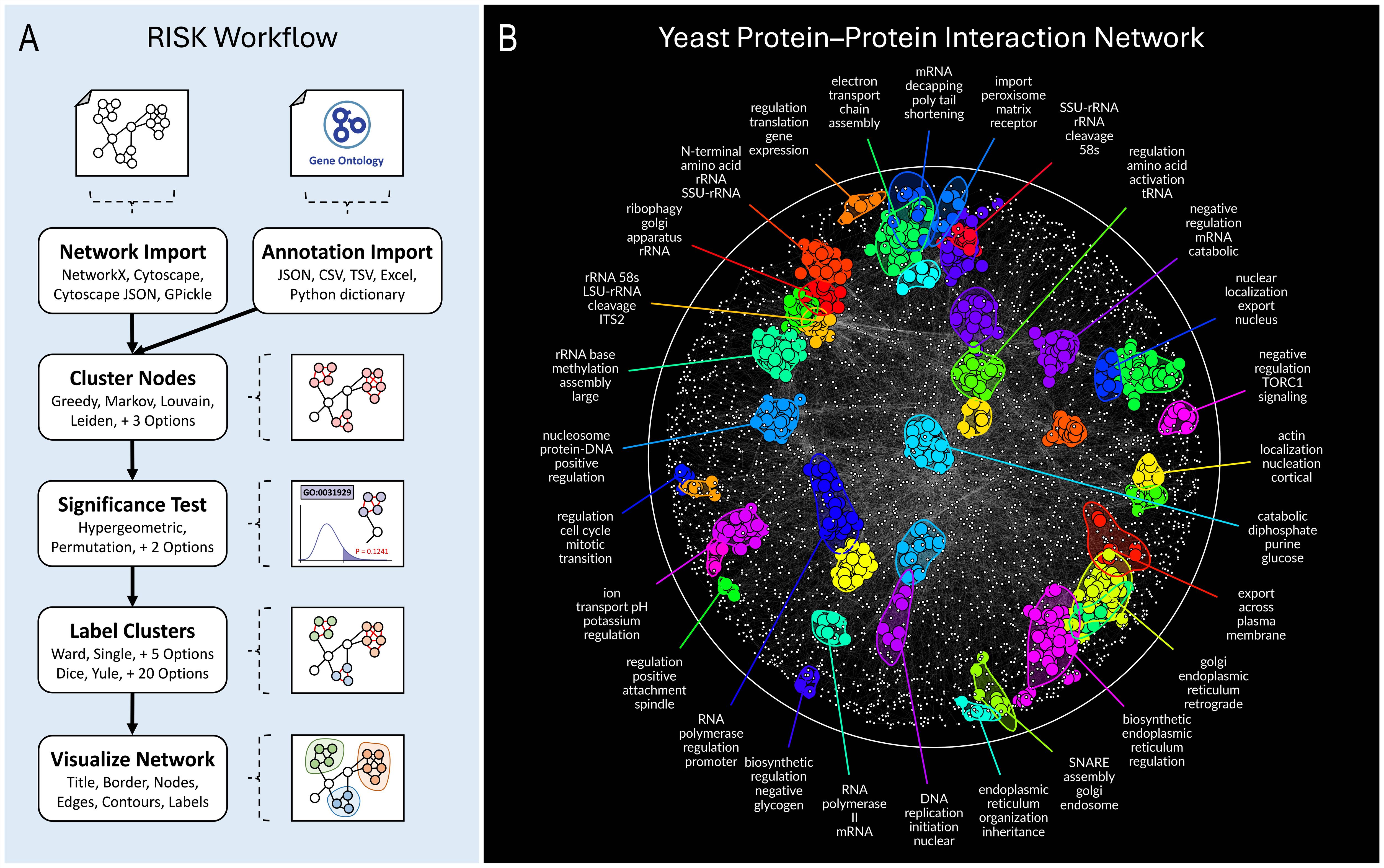
We applied RISK to a Saccharomyces cerevisiae protein–protein interaction (PPI) network (Michaelis et al., 2023; 3,839 proteins, 30,955 interactions). RISK identified compact, functional modules overrepresented in Gene Ontology Biological Process (GO BP) terms (Ashburner et al., 2000), revealing biological organization including ribosomal assembly, mitochondrial organization, and RNA polymerase activity.
Citation
If you use RISK in your research, please cite the following:
Horecka and Röst (2025), "RISK: a next-generation tool for biological network annotation and visualization".
DOI: 10.5281/zenodo.17257418
Contributing
We welcome contributions from the community:
Support
If you encounter issues or have suggestions for new features, please use the Issues Tracker on GitHub.
License
RISK is open source under the GNU General Public License v3.0.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file risk_network-0.0.16b2.tar.gz.
File metadata
- Download URL: risk_network-0.0.16b2.tar.gz
- Upload date:
- Size: 102.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4d1fc00e58b2bc450cb1af847b8562e138b09b08e63c98c2a63202a3994d7932
|
|
| MD5 |
83fcfaac4a5547f2d3bde92807503744
|
|
| BLAKE2b-256 |
de27f7561537521eaab77bd759608a3d9edf2a169da95433b187b6ca9725e07f
|
File details
Details for the file risk_network-0.0.16b2-py3-none-any.whl.
File metadata
- Download URL: risk_network-0.0.16b2-py3-none-any.whl
- Upload date:
- Size: 96.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
185fff94eed1ac60125e15c5808a4316cb4c10cc0adb4964655635b1c2e538c0
|
|
| MD5 |
c8300a9bc2a7d39b4ddec3532cb8370f
|
|
| BLAKE2b-256 |
68db1e070e6ce3364960f06dfe88d09c889a6b8e3e6fc90043cccaf75132b941
|
















