Package for predicting 5EU in nanopore reads and predicting RNA halflives
Project description
RNAkinet
RNAkinet is a project dedicated to detecting 5eu-modified reads directly from the raw nanopore sequencing signal. Furthermore, it offers tools to calculate transcript halflives.
Usage
Installation
pip install rnakinet
Predict 5EU in POD5 files
rnakinet-inference --path <pod5_file_or_directory> --model-name rnakinet_r10_5EU --output <predictions_name.csv>
This creates a csv file with columns read_id - the read id, 5eu_mod_score - the raw prediction score from 0 to 1, 5eu_modified_prediction - Boolean column, True if the read is predicted to be modified by 5EU, False otherwise
Nvidia GPU is recommended to run this command. If you want to run inference on a CPU-only machine, use the --use-cpu option. This will substantially increase runtime.
FAST5 input is not currently supported. Support will be reimplemented in a future release; for now, inference can still be run by converting Fast5 reads to POD5 first.
Pass one or more POD5 files or directories via --path. Use --model-name to select the packaged pretrained model for your flow-cell chemistry, for example rnakinet_r9_5EU or rnakinet_r10_5EU.
Users who have trained their own RNAkinet models can use --model-path (must be used in conjunction with --arch) to run inference on custom models in place of --model-name.
Example
rnakinet-inference --path data/experiment/pod5_dir --model-name rnakinet_r10_5EU --output preds.csv
Calculate transcript halflives
rnakinet-predict-halflives --transcriptome-bam <path_to_transcriptome_alignment.bam> --predictions <predictions_name.csv> --tl <experiment_tl> --output <halflives_name.csv>
The --tl parameter is the duration for which the cells were exposed to 5EU in hours
The --predictions parameter is the output file of the 5EU prediction step described above
This creates a csv file with columns transcript - the transcript identifier from your BAM file, reads - the amount of reads available for the given transcript, percentage_modified - the percentage of reads of the given transcript that were predicted to contain 5EU, pred_t5 - the predicted halflife of the given transcript
Example
rnakinet-predict-halflives --transcriptome-bam alignments/experiment/transcriptome_alignment.bam --predictions preds.csv --tl 2.0 --output halflives.csv
Note that the calculated halflives pred_t5 are the most reliable for transcripts with high read count.
The following plots show correlation of halflives computed from RNAkinet predictions with experimentaly measured halflives [1] as we increase read count requirement. We recommend users to acknowledge this and put more confidence in halflife predictions for transcripts with high read count, and less confidence for transcripts with low read count.
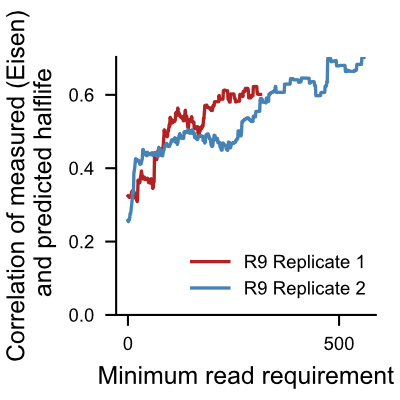
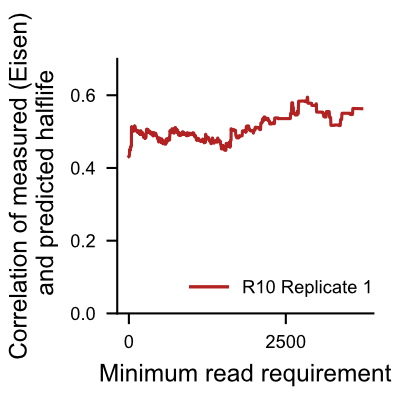
[1] Eisen,T.J., Eichhorn,S.W., Subtelny,A.O., Lin,K.S., McGeary,S.E., Gupta,S. and Bartel,D.P. (2020) The Dynamics of Cytoplasmic mRNA Metabolism. Mol. Cell, 77, 786-799.e10.
Model Training with Snakemake
Setup Conda Env
To run the snakefile, install environment.yaml using a conda environment
conda env create -f environment.yaml
Run snakemake from within the created conda environment rather than loading a module, etc.
Weights and Biases API
Obtiain an API key for Weights and Biases, and create a file .env (in the same directory as your Snakefile) that contains the line WANDB_API_KEY=YOUR_API_KEY. This will be used for tracking and visualizing model performance.
GPU and snakemake profile
Please ensure that the format for compute resource allocation is compatible with your snakemake profile and GPU/cluster requirements. Specifically, GPU names/quantities can be modified from GPUS_FOR_RULES in config.yml
Cite
Vlastimil Martinek, Jessica Martin, Cedric Belair, Matthew J Payea, Sulochan Malla, Panagiotis Alexiou, Manolis Maragkakis, Deep learning and direct sequencing of labeled RNA captures transcriptome dynamics, NAR Genomics and Bioinformatics, Volume 6, Issue 3, September 2024, lqae116, https://doi.org/10.1093/nargab/lqae116
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file rnakinet-2.0.1.tar.gz.
File metadata
- Download URL: rnakinet-2.0.1.tar.gz
- Upload date:
- Size: 2.1 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c506f1514daa94b8c6bd1890934a43b5578beb43e50294ec6bc5911049bebd61
|
|
| MD5 |
86140f70e6e6577e4a143c31307a7b5d
|
|
| BLAKE2b-256 |
e4a6806b15687764530ab791e2ffe70b101ee039c20a9c4d1ee12277ffafb689
|
File details
Details for the file rnakinet-2.0.1-py3-none-any.whl.
File metadata
- Download URL: rnakinet-2.0.1-py3-none-any.whl
- Upload date:
- Size: 2.1 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1df44c9c90697cf92aa21b2cfabb372ed124fcb4388e3b4b5bf12ce6542e7e71
|
|
| MD5 |
0d3a8f2f73e867f0c099f47ac76f0eed
|
|
| BLAKE2b-256 |
e14ee0850b8bf79a395c6a13ca2398dbacba56914d21ac204c819e0463fc39a6
|













