BIDs App to retrieve the haemodynamic response function from resting state fMRI data
Project description
Resting state HRF estimation and deconvolution.
Please refer to https://github.com/compneuro-da/rsHRF for MATLAB version
The basic idea
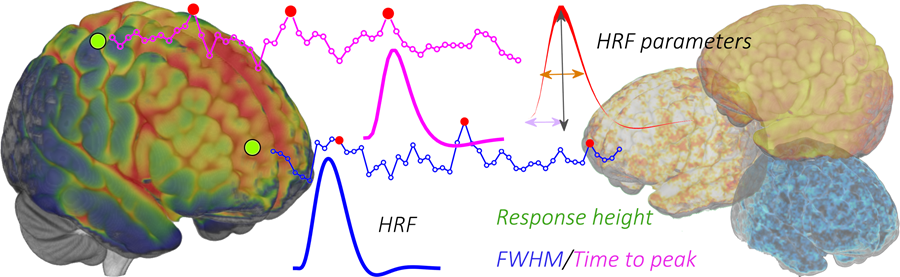
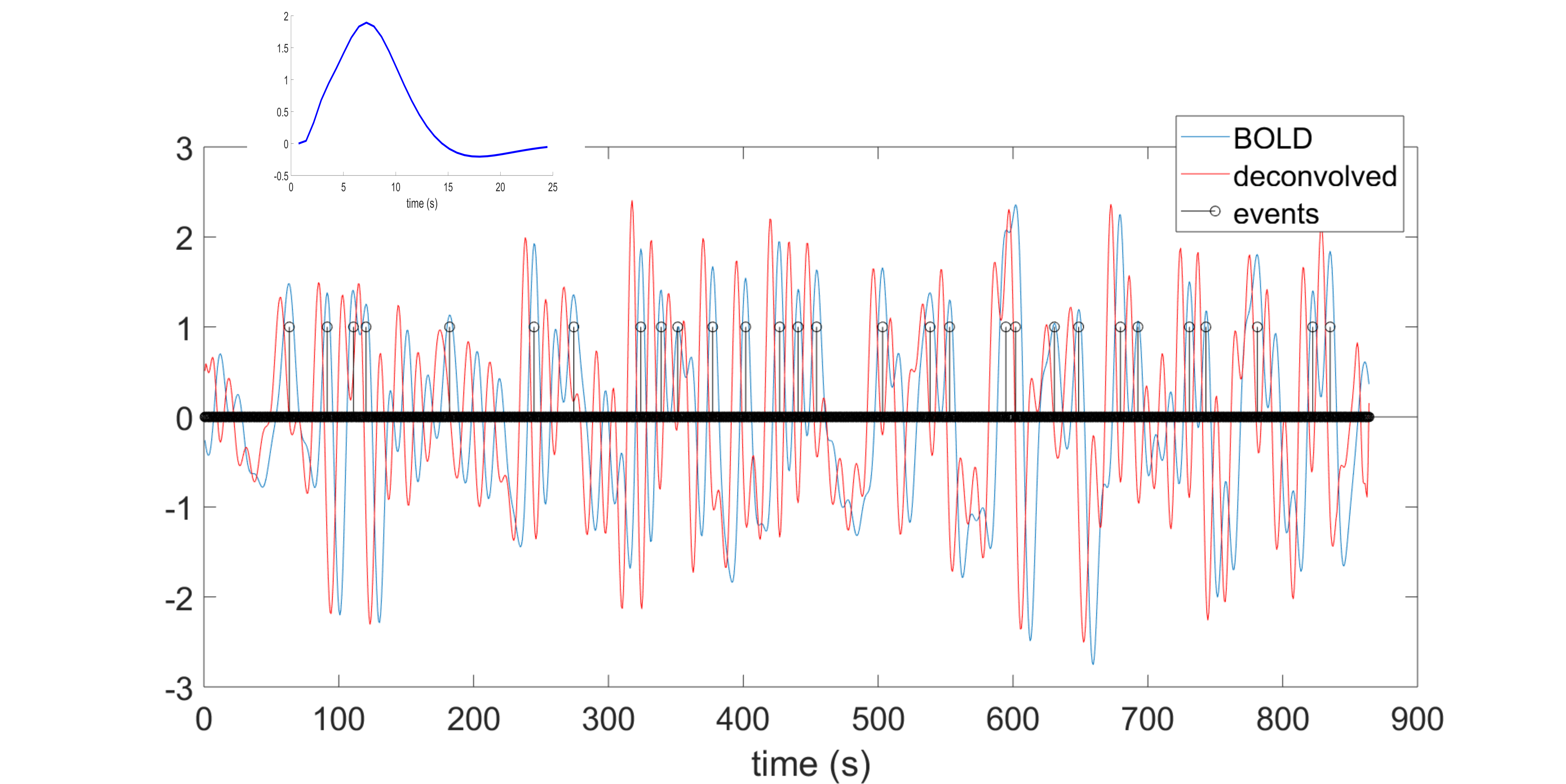
This toolbox is aimed to retrieve the onsets of pseudo-events triggering an hemodynamic response from resting state fMRI BOLD voxel-wise signal. It is based on point process theory, and fits a model to retrieve the optimal lag between the events and the HRF onset, as well as the HRF shape, using a choice of basis functions (the canonical shape with two derivatives, (smoothed) Finite Impulse Response, mixture of gammas).
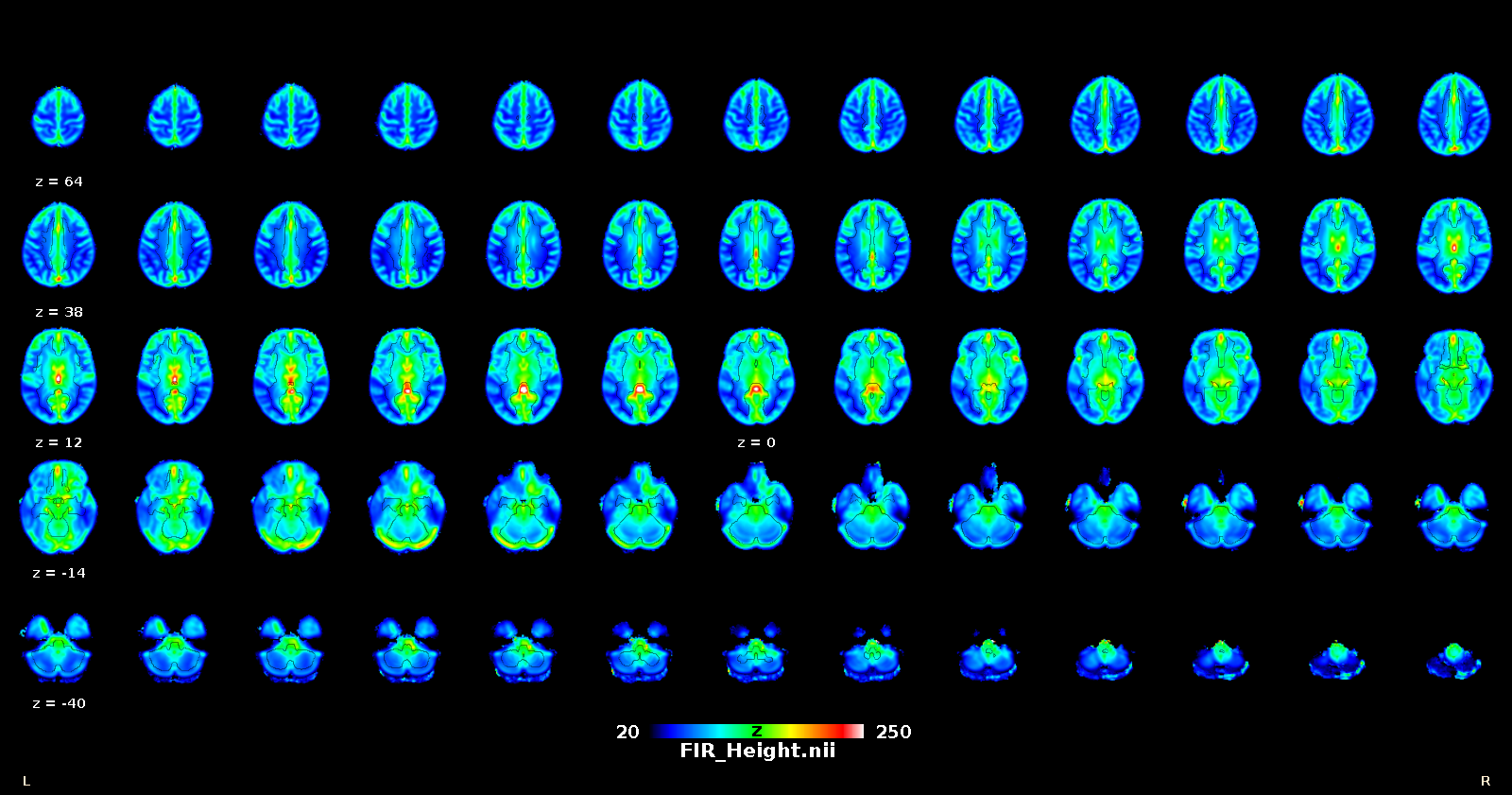
Once that the HRF has been retrieved for each voxel, it can be deconvolved from the time series (for example to improve lag-based connectivity estimates), or one can map the shape parameters everywhere in the brain (including white matter), and use the shape as a pathophysiological indicator.
How to use the toolbox
The input is voxelwise BOLD signal, already preprocessed according to your favorite recipe. Important thing are:
- bandpass filter in the 0.01-0.08 Hz interval (or something like that)
- z-score the voxel BOLD time series
To be on the safe side, these steps are performed again in the code.
The input can be images (3D or 4D), or directly matrices of [observation x voxels].
It is possible to use a temporal mask to exclude some time points (for example after scrubbing).
The demos allow you to run the analyses on several formats of input data.
Python Package and BIDS-app
A BIDS-App has been made for easy and reproducible analysis. Its documentation can be accessed at:
https://bids-apps.neuroimaging.io/rsHRF/
Collaborators
-
Guorong Wu
-
Nigel Colenbier
-
Sofie Van Den Bossche
-
Daniele Marinazzo
-
Madhur Tandon (Python - BIDS)
-
Asier Erramuzpe (Python - BIDS)
-
Amogh Johri (Python - BIDS)
References
-
Wu, G. R., Colenbier, N., Van Den Bossche, S., Clauw, K., Johri, A., Tandon, M., & Marinazzo, D. (2021). rsHRF: A toolbox for resting-state HRF estimation and deconvolution. Neuroimage, 244, 118591. open access journal page
-
Guo-Rong Wu, Wei Liao, Sebastiano Stramaglia, Ju-Rong Ding, Huafu Chen, Daniele Marinazzo*. "A blind deconvolution approach to recover effective connectivity brain networks from resting state fMRI data." Medical Image Analysis, 2013, 17:365-374. Open access institutional repo
-
Guo-Rong Wu, Daniele Marinazzo. "Sensitivity of the resting state hemodynamic response function estimation to autonomic nervous system fluctuations." Philosophical Transactions of the Royal Society A, 2016, 374: 20150190. Open access institutional repo
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file rshrf-1.6.2.tar.gz.
File metadata
- Download URL: rshrf-1.6.2.tar.gz
- Upload date:
- Size: 52.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
01a6dee7623859f20a9f71272f0d3672ac47c34eab98fe0564590404c8d8691b
|
|
| MD5 |
925afe20d2dd804673ab288e70ab1830
|
|
| BLAKE2b-256 |
641e1a46017e70a434171be49524c9a614c843ef3fd0d1775baab3f87adfb51f
|
File details
Details for the file rshrf-1.6.2-py3-none-any.whl.
File metadata
- Download URL: rshrf-1.6.2-py3-none-any.whl
- Upload date:
- Size: 64.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.9.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
adb987e3e903e8548776d13bf0423a45f47f9b93331ed9c851a878f8c790070d
|
|
| MD5 |
1a0f163c56b4fbf500a1a604b09990ca
|
|
| BLAKE2b-256 |
9549157a098996e5a275dedf52a5b28737069a7cfa8831aae4bee2afef0f951e
|