A package for mRNA sequence translation stochastic simulations
Project description
Python 2.7 or 3.5+ Version of the Single Molecule Translation Simulator (MatLab) by Dr. Luis Aguilera
Computational Design and Interpretation of Single-RNA Translation Experiments
Translated by Will Raymond - 2018/2019
rSNAPsim - RNA Sequence to NAscent Protein Simulation
Project Goal
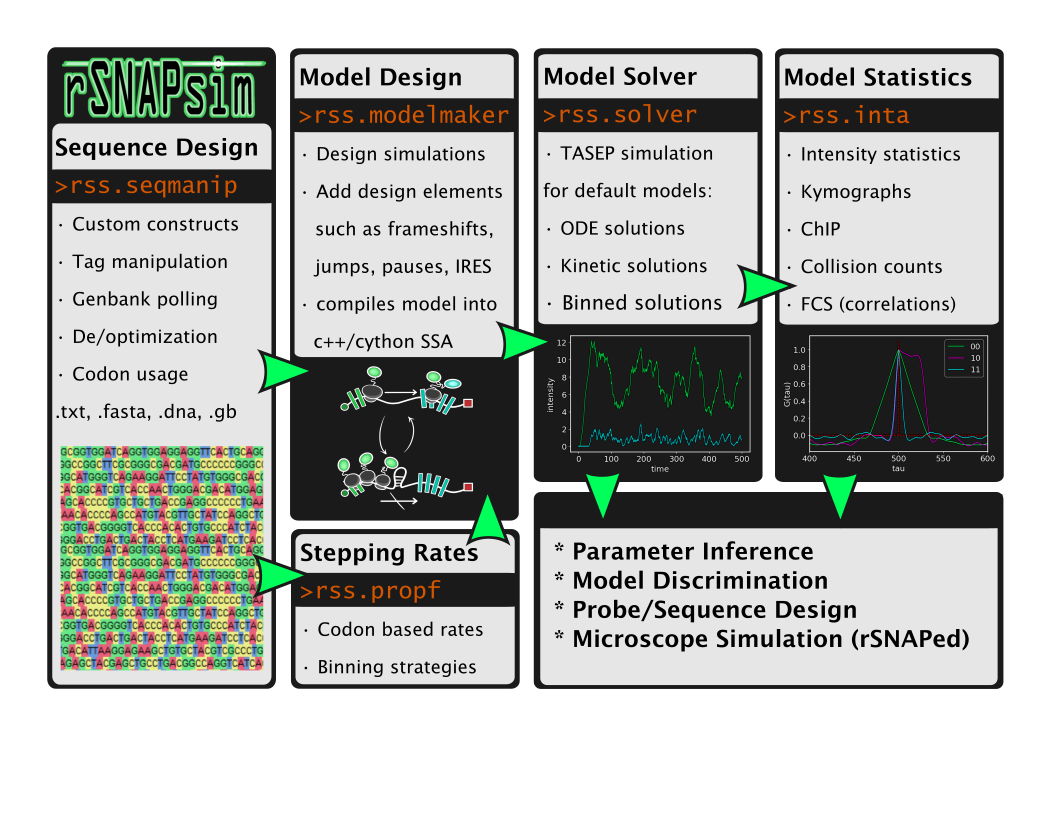
Provide a Python module that takes nucleotide sequence as an input and does the following:
- Choose a file or pull a file from GeneBank
- Analyzes the sequence and identifies proteins
- Detects or adds fluorescent tags
- Simulates translation trajectories and converts to intensity vectors of A.U. under various conditions
- Constructs with Rare codons only or Common codons, FRAP or Harringtonite assays
- Provides analyses of the trajectories
- Allows the user to save or export the data
- Commandline / GUI implementations
Documentation
Tutorials, Module Documentation, Installiation and more [LINK TO MUNSKY GROUP WEBSITE]
Dependencies:
Instillation
Within a conda enviroment:
conda install eigen
pip install rsnapsim-ssa-cpp
pip install rsnapsim
Within a Google Colab:
!apt install libeigen3-dev
!ln -sf /usr/include/eigen3/Eigen /usr/include/Eigen
!pip install rsnapsim-ssa-cpp
!pip install rsnapsim
!pip install --upgrade rsnapsim
Compilation of the C++
The c++ model should attempt to compile when you pip install the ssa-cpp module, however in the event that it cannot here are some common errors:
-
cannot include eigen3/Eigen/Dense
- This means eigen was not installed correctly from the conda installiation, you may have to manually download eigen and pass the argument to the setup.py command.
python setup.py build_ext --inplace -I[PATH TO EIGEN FOLDER]
- This means eigen was not installed correctly from the conda installiation, you may have to manually download eigen and pass the argument to the setup.py command.
-
gcc not found
Example Colab Notebooks
- Simulating Translation
- Simulating Constructs with Different codon usages
- Intensity Analyses
- Harringtonine / FRAP simulations
- Model Maker/ Designer
- MW/Diffusion Calculations
Future work
- Example notebooks of all functions
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file rsnapsim-0.0.49.tar.gz.
File metadata
- Download URL: rsnapsim-0.0.49.tar.gz
- Upload date:
- Size: 90.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.2 importlib_metadata/4.6.4 pkginfo/1.7.1 requests/2.26.0 requests-toolbelt/0.9.1 tqdm/4.62.1 CPython/3.8.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e25e7731c62200f3762747b87e5f723a25604373fc66c439db51de09e3f9f4dd
|
|
| MD5 |
cd10b50784a361dd32f6f5c064ea311e
|
|
| BLAKE2b-256 |
9ddd8ef670a84175bea98871e428d0ffe20ee964413de63edb1626fb0b47687f
|
File details
Details for the file rsnapsim-0.0.49-py3-none-any.whl.
File metadata
- Download URL: rsnapsim-0.0.49-py3-none-any.whl
- Upload date:
- Size: 124.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.2 importlib_metadata/4.6.4 pkginfo/1.7.1 requests/2.26.0 requests-toolbelt/0.9.1 tqdm/4.62.1 CPython/3.8.10
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fcb10fd1bfe175b40e2deace06973faf2442ed4d6bcab13e1d546ff2c9c1ad96
|
|
| MD5 |
710ca55c5760c679872412f8d8d55b8a
|
|
| BLAKE2b-256 |
78f175639fe0906462d4f9d14d5096374cee09056d1255b8d8bde1e411716281
|