Aligned Cross-modal Integration and Characterization of Single-Cell Multiomic Data with Deep Contrastive Learning
Project description
scMDCF
scMDCF is a python package containing tools for clustering single cell multi-omics data based on cross-modality contrastive learning to learn the common latent representation and assign clustering.
Overview
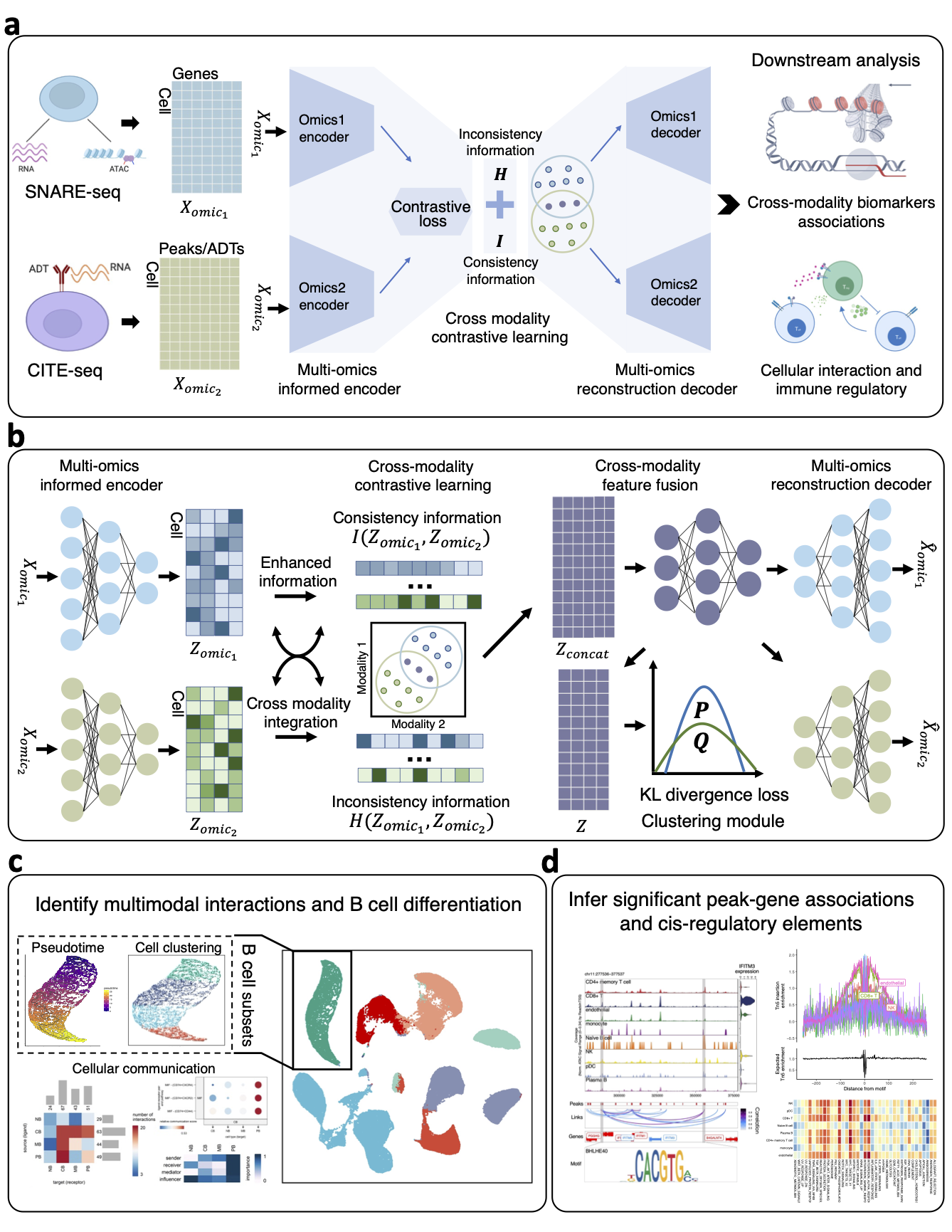
Single-cell multi-omics (scMulti-omics) technologies have revolutionized our understanding of cellular functions and interactions by enabling the simultaneous measurement of diverse cellular modalities. However, the inherent complexity, high-dimensionality, and heterogeneity of these datasets pose substantial challenges for integration and analysis across different modalities. To address these challenges, we develop a single-cell multi-omics deep learning model (scMDCF) based on contrastive learning, tailored for the efficient characterization and integration of scMulti-omics data. scMDCF features a cross-modality contrastive learning module that harmonizes data representations across different omics types, ensuring consistency while accommodating conditional entropy to preserve data heterogeneity. Furthermore, a cross-modality feature fusion module is designed to extract common low-dimensional latent representations of scMulti-omics data, effectively balancing the characteristics of these diverse omics data. Extensive empirical studies demonstrate that scMDCF outperforms existing state-of-the-art scMulti-omics models across various types of scMulti-omics data. In particular, scMDCF exhibits progressive capability in extracting cell-type specific peak-gene associations and cis-regulatory elements from SNARE-seq data, as well as in elucidating immune regulation from CITE-seq data. Furthermore, we demonstrate that in the post-BNT162b2 mRNA SARS‐CoV‐2 vaccination dataset, scMDCF successfully annotates specific vaccine-induced B cell subpopulations through integrative and multimodal analysis, uncovering dynamic interactions and regulatory mechanisms within the immune system after vaccination.
System Requirements
Hardware requirements
scMDCF package requires only a standard computer with enough RAM to support the in-memory operations.
Software requirements
OS requirements
This package is supported for Linux. The package has been tested on the following systems:
- Linux: Ubuntu 18.04
Python Dependencies
scMDCF mainly depends on the Python scientific stack.
numpy
pytorch
scanpy
pandas
scikit-learn
For specific setting, please see requirements.
Installation Guide
Install from PyPi
conda create -n scMDCF_env python=3.9.16
conda activate scMDCF_env
pip install scMDCF==1.1.3
Usage
scMDCF is a deep embedding learning method for single-cell multi-omics data clustering, which can be used to:
- CITE-seq dataset clustering. The example can be seen in the main_CITE.py
- SNARE-seq (paired RNA-seq and ATAC-seq) dataset clustering. The example can be seen in the main_SNARE.py
Data Availability
The datasets we used can be download in dataset
License
This project is covered under the MIT License.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file scMDCF-1.1.3.tar.gz.
File metadata
- Download URL: scMDCF-1.1.3.tar.gz
- Upload date:
- Size: 9.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.0.0 CPython/3.9.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9f6d10cc692ca46978065d559723fe6485a812bf4678e198bf7491ba9d050c3c
|
|
| MD5 |
2b5f87cd27ae6d07df1dbc53fa6f017d
|
|
| BLAKE2b-256 |
1ba3da9bf0b7673308024d39b65f8e37e104d1851b73bf62e2ad092f9fa168c9
|
File details
Details for the file scMDCF-1.1.3-py2.py3-none-any.whl.
File metadata
- Download URL: scMDCF-1.1.3-py2.py3-none-any.whl
- Upload date:
- Size: 8.4 kB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.0.0 CPython/3.9.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
56ba81d77b06331f42ccc929779ca2279b2bc565b1a8cf7e2f4b1cc7cb4eb3bc
|
|
| MD5 |
d32a498b1bd64e468d63909542241a83
|
|
| BLAKE2b-256 |
ec7c9951d879d7546cee7191b9d1cd1b886d39f0df255c4be0d249e27828aa1c
|