Single-Cell Multi-modality Integration via cell type filtered Anchors using Contrastive learning
Project description
scMIAC: Single-Cell Multi-modality Integration via cell type filtered Anchors using Contrastive learning
Overview
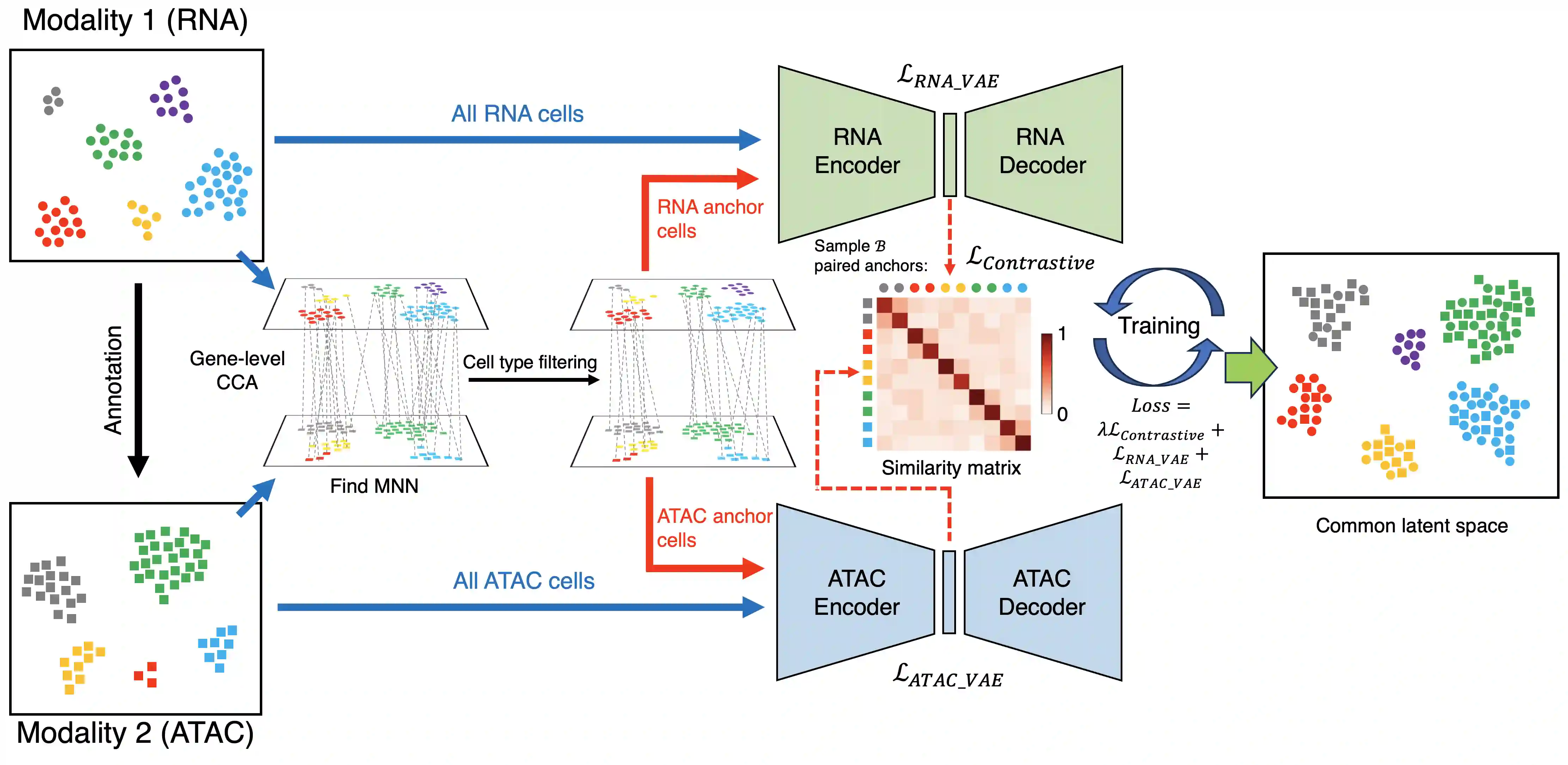
scMIAC is a comprehensive framework for single-cell multi-modality data integration, designed to tackle the most challenging problem in single-cell integration: diagonal integration (integrating unpaired cells from different feature spaces across modalities).
The methodological innovations of scMIAC include:
- scMIAC utilizes cell type information to select high-quality anchor cells for contrastive learning, improving integration of challenging cells such as imbalanced, rare, or isolated cell types.
- scMIAC innovatively introduces contrastive learning to diagonal integration task, where previous methods could only be applied to horizontal or vertical integration scenarios.
- As a diagonal integration approach, scMIAC preserves each modality's original biological characteristics through modality-specific VAEs, which serves as a regularizer preventing over-emphasis on modality alignment.
Installation
-
Create and activate a Conda environment:
conda create -n scmiac python=3.11 conda activate scmiac
-
Clone the repository and install:
git clone https://github.com/TianLab-Bioinfo/scMIAC.git cd scMIAC pip install .
Usage
scMIAC provides two usage modes:
1. CLI Mode - Quick start for non-interactive, command-line workflow:
scmiac train \
--rna-h5ad data/10x/input/adata_rna_10x.h5ad \
--atac-h5ad data/10x/input/adata_atac_10x.h5ad \
--output-dir data/10x/output/scmiac_results/ \
--rna-latent-key X_pca \
--atac-latent-key lsi49 \
--rna-celltype-key cell_type \
--atac-celltype-key pred
scmiac train -h # For viewing all available parameters
MUST Required parameters:
--rna-h5ad: Path to RNA AnnData file--atac-h5ad: Path to ATAC AnnData file--output-dir: Output directory
Output files:
anchors.csv: Anchor pairsrna_vae.pth: RNA VAE model weightsatac_vae.pth: ATAC VAE model weightsrna_embeddings.csv: RNA cell embeddingsatac_embeddings.csv: ATAC cell embeddingsscmiac_latent_umap.png: UMAP visualization
2. API Mode - For flexible research and experimentation:
Refer to the Full API Tutorial & Examples for detailed usage and examples.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file scmiac-0.1.0.tar.gz.
File metadata
- Download URL: scmiac-0.1.0.tar.gz
- Upload date:
- Size: 49.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8806ffb4adb68e34a61eec82d3a353fe5867ec941a92274d8f83961367627a9a
|
|
| MD5 |
cb61c8ddc886bea0e5f1c1e6dea693f5
|
|
| BLAKE2b-256 |
150dcb87ad318de8f89d48ceadece06bf15953b9dcd50703bcd99eaec5c48e4a
|
File details
Details for the file scmiac-0.1.0-py3-none-any.whl.
File metadata
- Download URL: scmiac-0.1.0-py3-none-any.whl
- Upload date:
- Size: 52.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
48ebc40258e859740345a538d2f36ad29b14a86f2c097ab2e6528df395b8f485
|
|
| MD5 |
a81f1a3fae43118cb2fd6ab4647f05c0
|
|
| BLAKE2b-256 |
dc99fceb5a07bd106140a56327786cccdc53b554e1431368500959831409a63b
|