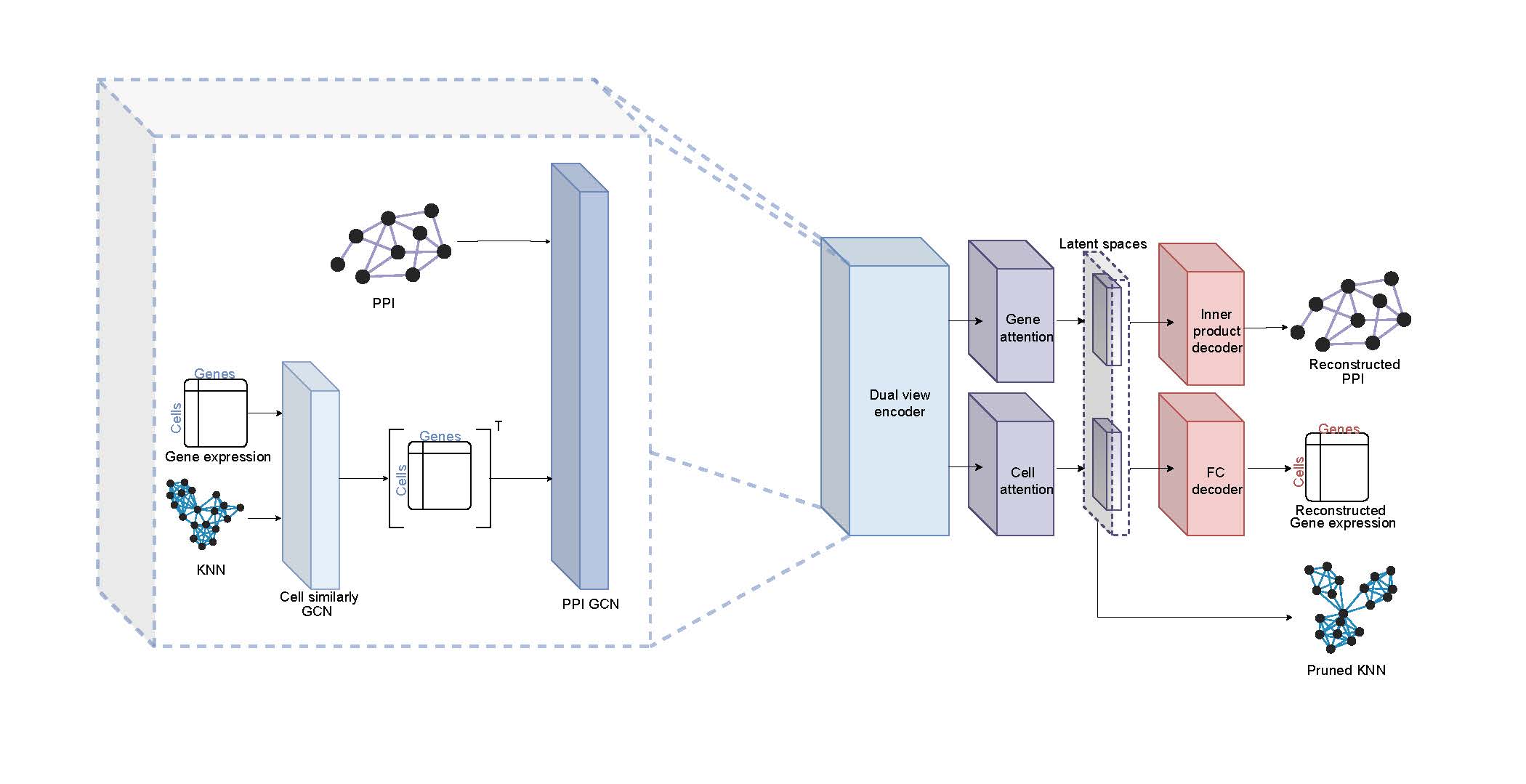
Our method employs a unique dual-graph architecture based on graph neural networks (GNNs), enabling the joint representation of gene expression and PPI network data
Project description
scNET: Learning Context-Specific Gene and Cell Embeddings by Integrating Single-Cell Gene Expression Data with Protein-Protein Interaction Information
Overview
Recent advances in single-cell RNA sequencing (scRNA-seq) techniques have provided unprecedented insights into tissue heterogeneity. However, gene expression data alone often fails to capture changes in cellular pathways and complexes, which are more discernible at the protein level. Additionally, analyzing scRNA-seq data presents challenges due to high noise levels and zero inflation. In this study, we propose a novel approach to address these limitations by integrating scRNA-seq datasets with a protein-protein interaction (PPI) network. Our method employs a unique bi-graph architecture based on graph neural networks (GNNs), enabling the joint representation of gene expression and PPI network data. This approach models gene-to-gene relationships under specific biological contexts and refines cell-cell relations using an attention mechanism, resulting in new gene and cell embeddings.
Download via PIP
pip install scnet
Download via git
To clone the repository, use the following command:
git clone https://github.com/madilabcode/scNET
We recommend using the provided Conda environment located at ./Data/scNET-env.yaml. cd scNET conda env create -f ./Data/scNET-env.yaml
import scNET
import scNET
API
To train scNET on scRNA-seq data, first load an AnnData object using Scanpy, then initialize training with the following command:
scNET.run_scNET(obj, pre_processing_flag=False, human_flag=False, number_of_batches=3, split_cells= True, max_epoch=250, model_name = project_name)
with the following args:
-
obj (AnnData, optional): AnnData obj.
-
pre_processing_flag (bool, optional): If True, perform pre-processing steps.
-
human_flag (bool, optional): Controls gene name casing in the network.
-
number_of_batches (int, optional): Number of mini-batches for the training.
-
split_cells (bool, optional): If True, split by cells instead of edges during training.
-
n_neighbors (int, optional): Number of neighbors for building the adjacency graph.
-
max_epoch (int, optional): Max number of epochs for model training.
-
model_name (str, optional): Identifier for saving the model outputs.
-
save_model_flag (bool, optional): If True, save the trained model.
Retrieve embeddings and model outputs with:
embedded_genes, embedded_cells, node_features , out_features = scNET.load_embeddings(project_name)
where:
-
embedded_genes (np.ndarray): Learned gene embeddings.
-
embedded_cells (np.ndarray): Learned cell embeddings.
-
node_features (pd.DataFrame): Original gene expression matrix.
-
out_features (np.ndarray): Reconstructed gene expression matrix
Create a new AnnData object using model outputs:
recon_obj = scNET.create_reconstructed_obj(node_features, out_features, obj)
Construct a co-embedded network using the gene embeddings:
scNET.build_co_embeded_network(embedded_genes, node_features)
Delete all files associated with a project (models, embeddings, and KNN graphs)
scNET.delete_project(project_name)
Tutorials
For a basic usage example of our framework, please refer to the following notebook: scNET Example Notebook
For a uasge example with batch integration using bbknn graph, plese refer to the following notebook: scNET Multi Batch Example Notebook
For a simple usage example on gene inference using scNET gene embedding,please refer to the following notebook: scNET Icos embedding
For a simple example of predicting functional annotations using gene embeddings, please refer to the following notebook: scNET functional annotations
For a example of how to use scNET to identify CD8+ T Cells subpopulation please refer to the following notebook: scNET subpouplation clustring
Contact
For questions or feedback, please contact ronsheinin@mail.tau.ac.il or open a GitHub issue.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file scnet-0.2.4.tar.gz.
File metadata
- Download URL: scnet-0.2.4.tar.gz
- Upload date:
- Size: 7.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.11.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
21abcbadd81683eb53d0f2504ff52107fde4c5a449fd23efda9fe77936fce72d
|
|
| MD5 |
9b74d753f68d27140e6902abd23327e0
|
|
| BLAKE2b-256 |
fe579b6057d03c4b7b07c2fee11dd8e4591e818c7d718293a439aee840b297b7
|
File details
Details for the file scnet-0.2.4-py3-none-any.whl.
File metadata
- Download URL: scnet-0.2.4-py3-none-any.whl
- Upload date:
- Size: 7.5 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.11.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f738589ec7e0ad511a536249fc9e74834872ec371f8b4f1e24c67f9018b6745c
|
|
| MD5 |
ab697698cc77a8ed45f9f93b55985048
|
|
| BLAKE2b-256 |
4bc6c0fe2819019fb3a8f68d1efaae9a211baa4c3618fe61703a2c5a1adbaddd
|