Pan-cancer detection of T-cell clonal expansion from single-cell RNA sequencing (CUDA-enabled)
Project description

scXpand: Pan-cancer Detection of T-cell Clonal Expansion
Detect T-cell clonal expansion from single-cell RNA sequencing data without paired TCR sequencing
Documentation • Installation • Quick Start • Examples • Citation
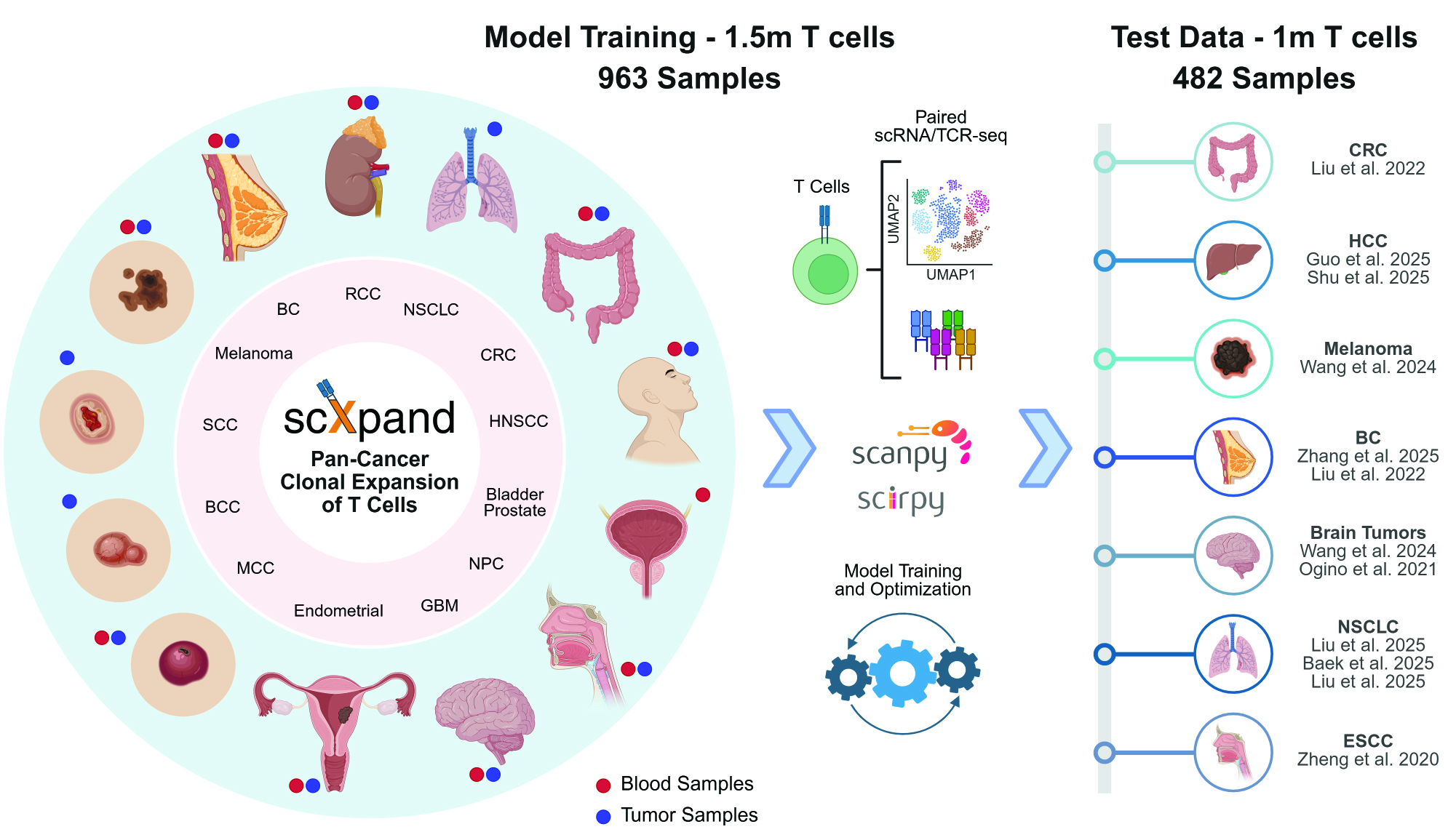
A framework for predicting T-cell clonal expansion from single-cell RNA sequencing data.
Manuscript in preparation - detailed methodology and benchmarks coming soon.
View full documentation for comprehensive guides and API reference.
Features
- Multiple Model Architectures:
- Autoencoder-based: Encoder-decoder with reconstruction and classification heads
- MLP: Multi-layer perceptron
- LightGBM: Gradient boosted decision trees
- Linear Models: Logistic regression and support vector machines
- Scalable Processing: Handles millions of cells with memory-efficient data streaming from disk during training
- Automated Hyperparameter Optimization: Built-in Optuna integration for model tuning
Installation
scXpand is available in two variants to match your hardware:
If you have an NVIDIA GPU with CUDA support:
With plain pip (add CUDA index):
pip install --upgrade scxpand-cuda --extra-index-url https://download.pytorch.org/whl/cu128
With uv:
uv pip install --upgrade scxpand-cuda --extra-index-url https://download.pytorch.org/whl/cu128 --index-strategy unsafe-best-match
Otherwise (CPU, Apple Silicon, or non-CUDA GPUs):
With plain pip:
pip install --upgrade scxpand
With uv:
uv pip install --upgrade scxpand
See the full installation guide for detailed setup instructions.
Quick Start
import scxpand
# Make sure that "your_data.h5ad" includes only T cells for the results to be meaningful
# Ensure that "your_data.var_names" are provided as Ensembl IDs (as the pre-trained models were trained using this gene representation)
# Please refer to our documentation for more information
# List available pre-trained models
scxpand.list_pretrained_models()
# Run inference with automatic model download
results = scxpand.run_inference(
model_name="pan_cancer_autoencoder", # default model
data_path="your_data.h5ad"
)
Documentation
See our Tutorial Notebook for a complete example with data preprocessing, T-cell filtering, gene ID conversion, and model application using a real breast cancer dataset.
Getting Started:
- Development Guide - Setup for local development of scXpand
- Usage Examples - Detailed tutorials and CLI examples
- Data Format - Input data requirements and specifications
Model Training & Optimization:
- Model Architectures - Detailed architecture descriptions and configurations
- Hyperparameter Optimization - Automated model tuning with Optuna
- Data Pipeline - Preprocessing and normalization details
Analysis & Evaluation:
- Evaluation Metrics - Performance assessment and stratified analysis
- Output Format - Understanding model outputs and results
📖 Full Documentation - Complete guides, API reference, and interactive tutorials
License
This project is licensed under the MIT License – see the LICENSE file for details.
Citation
If you use scXpand in your research, please cite:
@article{scxpand2025,
title={scXpand: Pan-cancer detection of T-cell clonal expansion from single-cell RNA sequencing without paired single-cell TCR sequencing},
author={[Your Name]},
journal={[Journal Name]},
year={2025},
doi={[DOI]}
}
This project was created in favor of the scientific community worldwide, with a special dedication to the cancer research community. We hope you’ll find this repository helpful, and we warmly welcome any requests or suggestions - please don’t hesitate to reach out!
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file scxpand_cuda-0.1.56.tar.gz.
File metadata
- Download URL: scxpand_cuda-0.1.56.tar.gz
- Upload date:
- Size: 126.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.8.17
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cc08188f6caa0b880b63fa535ce405733749b8c64f6d3e4ee71351d3a24c9247
|
|
| MD5 |
e9599983a0ed51cbb11b4653c74c3a7c
|
|
| BLAKE2b-256 |
78df3679066377109869ea741f5d1641e4e23e458e9803d7aefc63ca25c15378
|
File details
Details for the file scxpand_cuda-0.1.56-py3-none-any.whl.
File metadata
- Download URL: scxpand_cuda-0.1.56-py3-none-any.whl
- Upload date:
- Size: 138.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: uv/0.8.17
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
111128b2766f19ae595ba0b5ee1dea3e0c1f299f287fdb3c147ec56952c954b1
|
|
| MD5 |
285a0b2f30e0b8c9ccda47064ff2f6bd
|
|
| BLAKE2b-256 |
40b2bd6b6aadd4b3c3813d149ccf7ca5fd5501ece603f7f9f423391586a76b95
|














