Efficient implementations of Needleman-Wunsch and other sequence alignment algorithms in Rust with Python bindings.
Project description
sequence_align
Efficient implementations of Needleman-Wunsch and other sequence alignment algorithms written in Rust with Python bindings via PyO3. Supports both binary match/mismatch scoring and custom pairwise scoring functions for applications like OCR text alignment, spatial matching, and other domains where continuous similarity measures are needed.
Installation
sequence_align is distributed via PyPi for Python 3.10 - 3.14, making installation as simple as the following --
no special setup required for cross-platform compatibility, Rust installation, etc.!
pip install sequence_align
Alternatively, if one wishes to develop for sequence_align, first ensure that both
Python and Rust
are installed on your system. Then, install Maturin and run
maturin develop (optionally with the -r flag to compile a release build, instead of an unoptimized debug build)
from the root of your cloned repo to build and install sequence_align in your active Python environment.
Quick Start
Pairwise sequence algorithms are available in sequence_align.pairwise.
The following algorithms are implemented:
- Needleman-Wunsch: Global sequence alignment with
O(M*N)time and space. - Needleman-Wunsch with custom scores: A variant that accepts a custom pairwise scoring function
score_fn(a, b) -> floatinstead of flat match/mismatch scores. This is useful when alignment quality depends on continuous similarity measures rather than binary element equality. - Hirschberg: A modification of Needleman-Wunsch with the same
O(M*N)time complexity but onlyO(min{M, N})space, making it an appealing option for memory-limited applications or extremely large sequences.
One may also compute the Needleman-Wunsch alignment score for alignments produced by any of the above algorithms
using alignment_score.
Using these algorithms is straightforward:
from sequence_align.pairwise import (
alignment_score,
hirschberg,
needleman_wunsch,
needleman_wunsch_with_scores,
)
# See https://en.wikipedia.org/wiki/Needleman%E2%80%93Wunsch_algorithm#/media/File:Needleman-Wunsch_pairwise_sequence_alignment.png
# Use Needleman-Wunsch default scores (match=1, mismatch=-1, indel=-1)
seq_a = ["G", "A", "T", "T", "A", "C", "A"]
seq_b = ["G", "C", "A", "T", "G", "C", "G"]
aligned_seq_a, aligned_seq_b = needleman_wunsch(
seq_a,
seq_b,
"_", # Represent gaps with this value
match_score=1.0,
mismatch_score=-1.0,
indel_score=-1.0,
)
# Expects ["G", "_", "A", "T", "T", "A", "C", "A"]
print(aligned_seq_a)
# Expects ["G", "C", "A", "_", "T", "G", "C", "G"]
print(aligned_seq_b)
# Expects 0
score = alignment_score(
aligned_seq_a,
aligned_seq_b,
"_",
match_score=1.0,
mismatch_score=-1.0,
indel_score=-1.0,
)
print(score)
# See https://en.wikipedia.org/wiki/Hirschberg%27s_algorithm#Example
seq_a = ["A", "G", "T", "A", "C", "G", "C", "A"]
seq_b = ["T", "A", "T", "G", "C"]
aligned_seq_a, aligned_seq_b = hirschberg(
seq_a,
seq_b,
"_",
match_score=2.0,
mismatch_score=-1.0,
indel_score=-2.0,
)
# Expects ["A", "G", "T", "A", "C", "G", "C", "A"]
print(aligned_seq_a)
# Expects ["_", "_", "T", "A", "T", "G", "C", "_"]
print(aligned_seq_b)
# Expects 1
score = alignment_score(
aligned_seq_a,
aligned_seq_b,
"_",
match_score=2.0,
mismatch_score=-1.0,
indel_score=-2.0,
)
print(score)
# Custom pairwise scoring: align words using character overlap similarity
words_a = ["hello", "world", "foo"]
words_b = ["hallo", "welt", "baz", "foo"]
def char_overlap_score(a: str, b: str) -> float:
"""Score based on character-level overlap between two words."""
if a == b:
return 2.0
shared = len(set(a) & set(b))
total = len(set(a) | set(b))
return (2.0 * shared / total) - 1.0 if total > 0 else -1.0
aligned_words_a, aligned_words_b = needleman_wunsch_with_scores(
words_a,
words_b,
"_",
score_fn=char_overlap_score,
indel_score=-1.0,
)
# Expects ["hello", "world", "_", "foo"]
print(aligned_words_a)
# Expects ["hallo", "welt", "baz", "foo"]
print(aligned_words_b)
Development
To set up a local development environment, ensure that both Python and Rust are installed, then:
maturin develop -r # build and install in the active Python environment
./scripts/test.sh # run tests via pytest
./scripts/lint.sh # run all linters (ruff, mypy, cargo fmt, cargo clippy)
./scripts/lint.sh --fix # auto-fix where possible
Performance Benchmarks
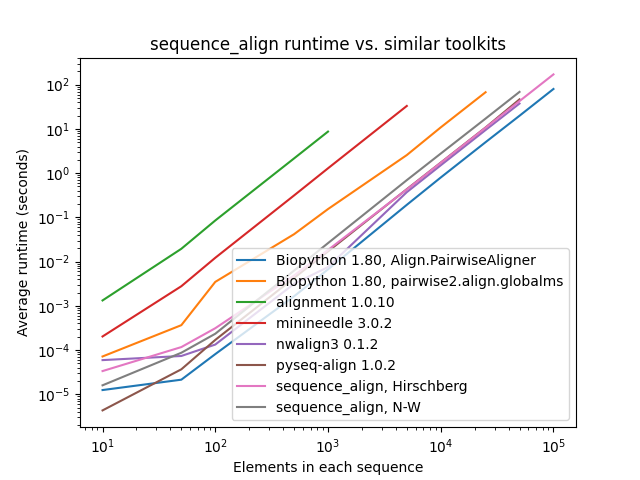
All tests below were conducted sequentially on a AWS R5.4 instance with 16 cores and 128 GB of memory. The pair of sequences for alignment consist of a character sequence of randomly selected A/C/G/T nucleotide bases along with another that is identical, except with 10% of the characters randomly perturbed by deletion, insertion of another randomly-selected character after the entry, or replacement with a different randomly-selected character.
As one can see, while sequence_align is comparable to some other toolkits in terms of speed, its memory performance
is best-in-class, even when compared to toolkits using the same algorithm, such as Needleman-Wunsch being used in
pyseq-align.
(Please note that some lines terminate early, as some toolkits took prohibitively long and/or ran out of memory at higher scales.)
Changelog
See CHANGELOG.md for a full list of changes across versions.
License
Licensed under the Apache 2.0 License. Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
Copyright 2023-present Kensho Technologies, LLC. The present date is determined by the timestamp of the most recent commit in the repository.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sequence_align-0.4.0-cp310-abi3-win_amd64.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-win_amd64.whl
- Upload date:
- Size: 111.2 kB
- Tags: CPython 3.10+, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
988e1413ba1a241ec4df7ed1a62f0b98afacf63dd13dc85efc54fff15a1c8177
|
|
| MD5 |
6df90fdc4ecf8e54cf87ebf22e7344c3
|
|
| BLAKE2b-256 |
2ddccf40fe93d70f08db7e7665e28d55b4d8c3f60a2dcf89244c847f247cf750
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-win32.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-win32.whl
- Upload date:
- Size: 107.2 kB
- Tags: CPython 3.10+, Windows x86
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d376fa8adac14f575957c14a56d8185538883680e0094081336b45b046dbc115
|
|
| MD5 |
5e08366a6359b6c48292013f101a8c19
|
|
| BLAKE2b-256 |
acb0902df73c42671e4d9eaf69686744662a9256380b6e403d3455e5f47febdb
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 226.6 kB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c608d953403477a1de01f65274dedc043636f97cc5084cacf48f5eab19018025
|
|
| MD5 |
9c6a389ff9ed82ffa14c6d8a6fda4c2a
|
|
| BLAKE2b-256 |
cb2cce874d9ab02c2e12805483c7a4ca6bbecab1f06cece5a50ead8eed2dbc08
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-manylinux_2_17_s390x.manylinux2014_s390x.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-manylinux_2_17_s390x.manylinux2014_s390x.whl
- Upload date:
- Size: 256.2 kB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ s390x
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d062d0f0c18b28703dfa9fc77923289c912479b5ce85b51511dda22793232160
|
|
| MD5 |
1863a59a222047c70a3ad232f6a222c5
|
|
| BLAKE2b-256 |
2b4253a1a59eee2027a26a0324c4e19ce4298294f4f58fef9b9eff4ac2b49ea7
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-manylinux_2_17_ppc64le.manylinux2014_ppc64le.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-manylinux_2_17_ppc64le.manylinux2014_ppc64le.whl
- Upload date:
- Size: 252.6 kB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ ppc64le
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
26d10d5823a04ecf284b8e8513f1a8709ed4db49e969122557fb17c1385b8987
|
|
| MD5 |
9eed1f4dd118dd03f2b04790310e1088
|
|
| BLAKE2b-256 |
0163754587630e166b05f00261b263fb4643e7348316e6d89bdbebc8b41a77a5
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-manylinux_2_17_armv7l.manylinux2014_armv7l.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-manylinux_2_17_armv7l.manylinux2014_armv7l.whl
- Upload date:
- Size: 232.8 kB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ ARMv7l
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b73a638f5fdbaf1b99697bc13882304b6125eed42cd70d2d18af20bd02a19bc1
|
|
| MD5 |
cc5c5753a82910878e5704ddc6f6078f
|
|
| BLAKE2b-256 |
1abff7c18fb9357fa39563c0bc2a0c88679eed91c31f0c94b2f56442cf04ce98
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-manylinux_2_17_aarch64.manylinux2014_aarch64.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-manylinux_2_17_aarch64.manylinux2014_aarch64.whl
- Upload date:
- Size: 224.3 kB
- Tags: CPython 3.10+, manylinux: glibc 2.17+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4e19bb3a70cc2256beff220a84a478a27ac2dd1ffd6df0760d6d0d1b7258c1db
|
|
| MD5 |
b9e8babb0673a570e263e58cc8bbce33
|
|
| BLAKE2b-256 |
7ab52fd8b7d901410a060458b09dfa124fb2423b2ae60964d5aa38485609b00d
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-manylinux_2_5_i686.manylinux1_i686.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-manylinux_2_5_i686.manylinux1_i686.whl
- Upload date:
- Size: 241.6 kB
- Tags: CPython 3.10+, manylinux: glibc 2.5+ i686
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f2997b8f5694981ef39a924c282e752873cf11b2187abfa18771829096861329
|
|
| MD5 |
3ccf8ea581cdd59652605db7d12b9e97
|
|
| BLAKE2b-256 |
7e9ce96d416eb82b3b86d16fd9aab40222796a5392e8074b01aff8a20f7d5640
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-macosx_11_0_arm64.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-macosx_11_0_arm64.whl
- Upload date:
- Size: 205.9 kB
- Tags: CPython 3.10+, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
757d34f36ac096b749ddffdbb1b4046c13f93f2414136c244ba8a1b2fa9305bd
|
|
| MD5 |
eab2a86730b324f26c4fd868317bd14a
|
|
| BLAKE2b-256 |
1f2eaae0eea098cc4155d1be38083dac2bdbb1cf516510992ef38dc09e4bfd9c
|
File details
Details for the file sequence_align-0.4.0-cp310-abi3-macosx_10_12_x86_64.whl.
File metadata
- Download URL: sequence_align-0.4.0-cp310-abi3-macosx_10_12_x86_64.whl
- Upload date:
- Size: 216.9 kB
- Tags: CPython 3.10+, macOS 10.12+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.0
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
522268ec87a4608cd9dd1f1b068de2c4fe919e073583ad71afa5e058b778ca2c
|
|
| MD5 |
e526cbe0c77641e78536c014da6fb3b6
|
|
| BLAKE2b-256 |
1a63b72bff868b7890aaa1d2dbc88f2e1acd3d31f2dc8a188d3082d70e02297b
|