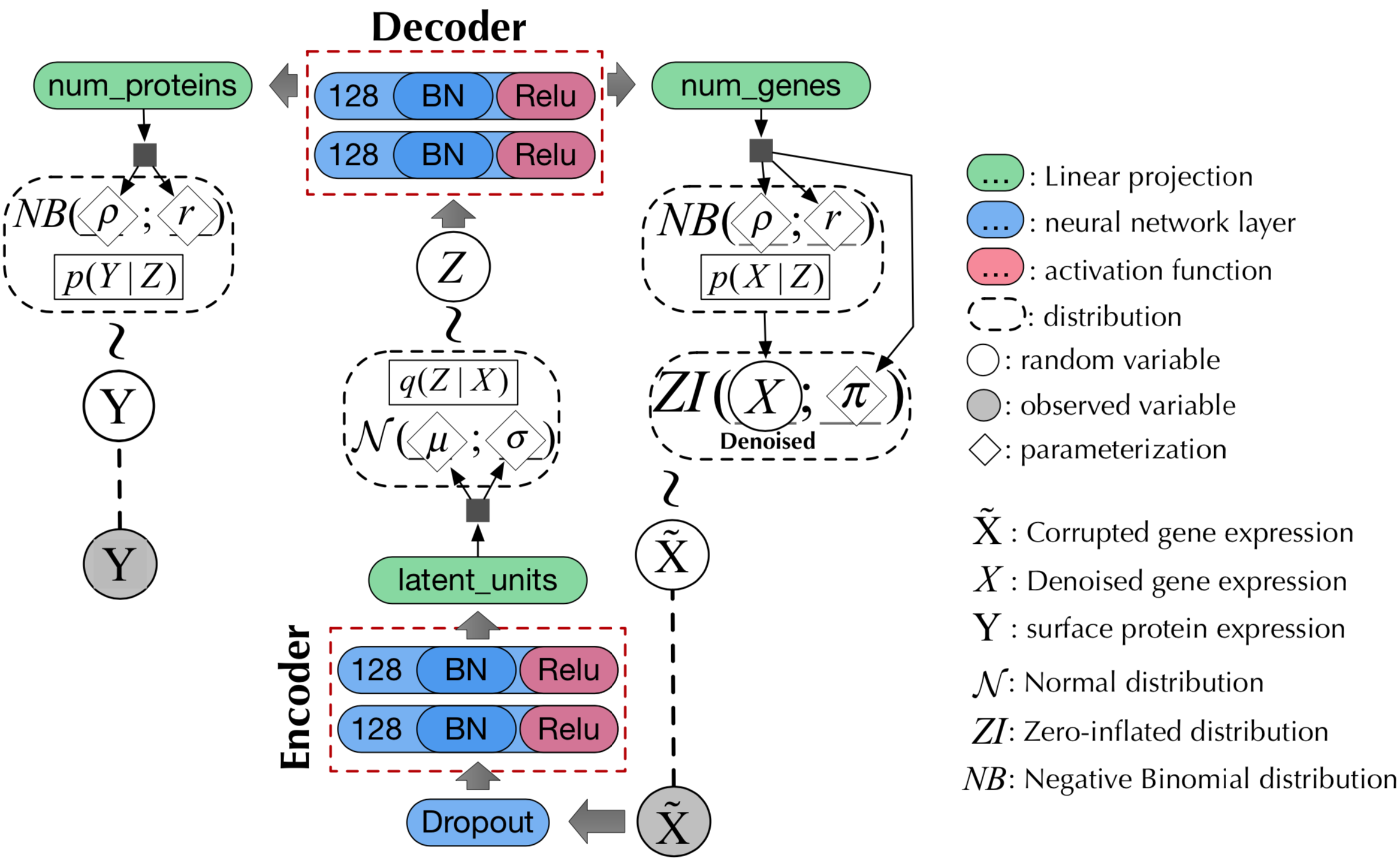
SemI-SUpervised generative Autoencoder for single cell data
Project description
Semi-supervised Single-cell modeling:
Free software: MIT license
Documentation: https://github.com/trungnt13/sisua/tree/master/docs.
Reference:
Trung Ngo Trong, Roger Kramer, Juha Mehtonen, Gerardo González, Ville Hautamäki, Merja Heinäniemi. “SISUA: SemI-SUpervised Generative Autoencoder for Single Cell Data”, ICML Workshop on Computational Biology, 2019. [pdf]
Installation
You only need Python 3.6, the stable version of SISUA installed via pip:
pip install sisua
Install the nightly version on github:
pip install git+https://github.com/trungnt13/sisua@master
For developers, we create a conda environment for SISUA contribution sisua_env
conda env create -f=sisua_env.yml
Getting started
- The basics:
Models specification
- Single-cell analysis:
Latent space
Imputation of genes expression
Prediction of protein markers
- Advanced technical topics:
Hierarchical modeling (coming soon)
Causal analysis (coming soon)
Cross datasets analysis (coming soon)
- Benchmarks:
Fine-tuning networks
Data normalization
Toolkits
We provide binary toolkits for fast and efficient analyzing single-cell datasets:
sisua-train: train single-cell modeling algorithms, support training multiple systems in parallel.
sisua-analyze: evaluate, compare, and interpret trained model.
sisua-embed: probabilistic embedding for semi-supervised training.
sisua-data: coming soon
Some important arguments:
- -model
name of function declared in models
scvi: single-cell Variational Inference model
dca: Deep Count Autoencoder
vae: single-cell Variational Autoencoder
movae: SISUA
- -ds
name of dataset declared in data.
Description of all predefined datasets is in docs.
Some good datasets for practicing:
pbmc8k_ly
cortex
pbmcecc_ly
pbmcscvi
pbmcscvae
Configuration
By default, the data will be saved at your home folder at ~/bio_data, and the experiments’ outputs will be stored at ~/bio_log
You can customize these two paths using the environment variables:
For storing downloaded and preprocessed data: SISUA_DATA
For the experiments: SISUA_EXP
For example:
import os
os.environ['SISUA_DATA'] = '/tmp/bio_data'
os.environ['SISUA_EXP'] = '/tmp/bio_log'
from sisua.data import EXP_DIR, DATA_DIR
print(DATA_DIR) # /tmp/bio_data
print(EXP_DIR) # /tmp/bio_logor you could set the variables in advance:
export SISUA_DATA=/tmp/bio_data
export SISUA_EXP=/tmp/bio_log
python sisua/train.py
# or using the provided toolkit: sisua-trainProject details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sisua-0.4.4.tar.gz.
File metadata
- Download URL: sisua-0.4.4.tar.gz
- Upload date:
- Size: 114.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.15.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.0.1 requests-toolbelt/0.9.1 tqdm/4.32.2 CPython/3.6.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
500eeda3795a439fc7c8e31ba4654dd6b5b480e3bbb3c76b516ce35f9814c446
|
|
| MD5 |
66b79233f9f3a22c96022f8d0023e12c
|
|
| BLAKE2b-256 |
1b590a51a8e3811892819cfc4f90509f0c32e8f8bf14462699995a5616d4f254
|
File details
Details for the file sisua-0.4.4-py3.6.egg.
File metadata
- Download URL: sisua-0.4.4-py3.6.egg
- Upload date:
- Size: 342.3 kB
- Tags: Egg
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.15.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.0.1 requests-toolbelt/0.9.1 tqdm/4.32.2 CPython/3.6.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
812be2c55077d40aea9e254377f0a7e47d75d3236f4e1a1ae7cb3f1e020b4d4c
|
|
| MD5 |
104408251f88243e75ff728e465c3c4b
|
|
| BLAKE2b-256 |
887d49e476fac4d658b2c328caf6237fa78e869c63c8adbf9555a349e37c0fb5
|