JAX implementation of the Skala neural exchange-correlation functional
Project description
skalax

pypi: https://pypi.org/project/skalax/
JAX/Equinox implementation of the Skala neural exchange-correlation functional for density functional theory (DFT) calculations.
Overview
skalax is a pure JAX port of the Skala neural XC functional. It reproduces the PyTorch reference to within ~2 kcal/mol (due to custom JAX PySCF wrapper) NPE on the tested systems and exposes the usual JAX machinery: jax.grad, jax.jit, jax.vmap, with XLA compilation.
The goal is to make this functional usable, finetunable and modifiable from JAX-based DFT codes.
[!WARNING] Work in progress, tested on CPU only so far. The PySCF JAX wrapper I wrote is not optimal and is therefore slower than the original PySCF Skala in Torch, but the model itself is comparable in training and inference performance.
Performance
JAX JIT (XLA) matches PyTorch on tested grid sizes. All variants use radius_cutoff=5.0 and are benchmarked in steady state (post-compilation, CPU). GPU validation is the next step.
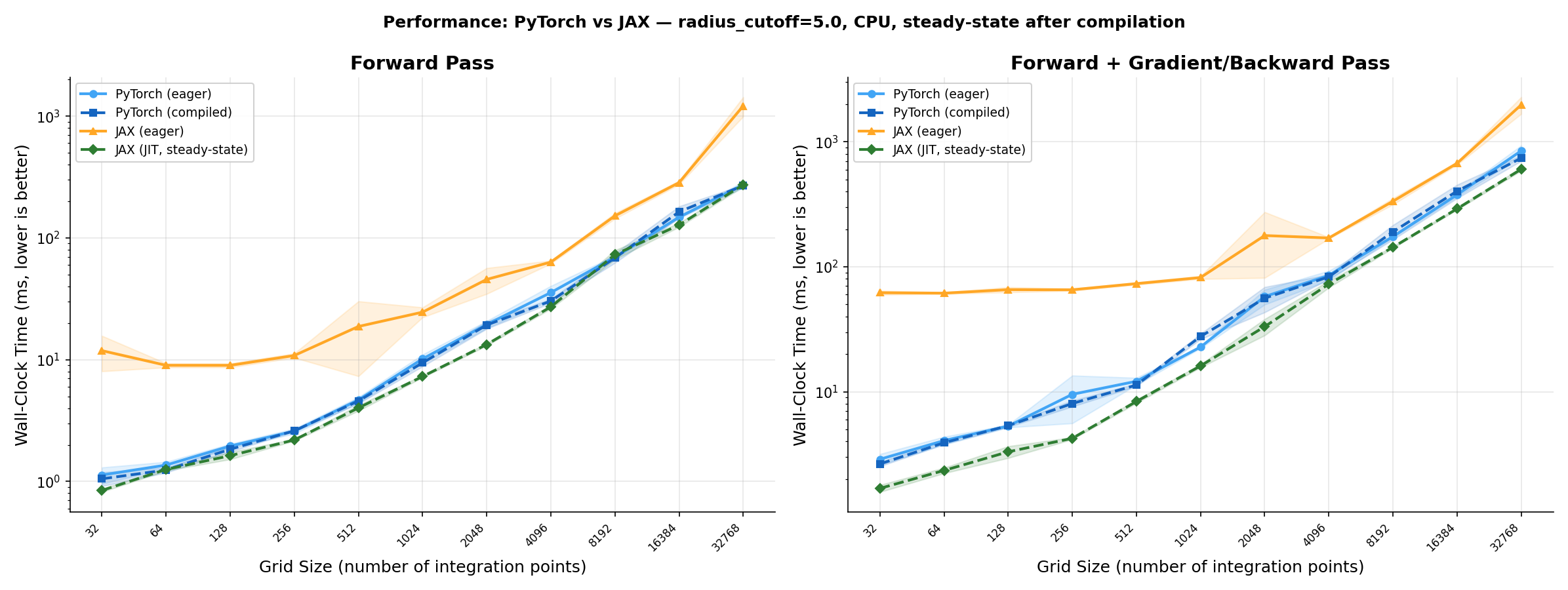
Left: forward pass only. Right: forward + backward. Eager: op-by-op execution (no compilation). JIT / traced: compiled graph; steady state = timed after a warm-up call, so compile cost is excluded. At 32k grid points on CPU, JAX JIT forward is ~1.4× faster than PyTorch traced, and JAX JIT fwd+grad is ~1.6× faster than PyTorch traced fwd+backward.
Installation
Requires gfortran and cmake (for dftd3/pyscf via skala).
pip install skalax
GPU Support
pip install skalax
pip install --upgrade "jax[cuda12]"
Development (no Fortran compiler needed)
git clone https://github.com/Brogis1/skalax
cd skalax
pip install -e .[dev]
pip install --no-deps microsoft-skala
Quick Start
Basic Usage
import jax
import jax.numpy as jnp
jax.config.update("jax_enable_x64", True)
from skalax import SkalaFunctional, load_weights_from_npz, load_config, get_default_weights_dir
# Load pretrained weights (bundled with package)
weights_dir = get_default_weights_dir()
config = load_config(weights_dir)
key = jax.random.PRNGKey(0)
model = SkalaFunctional(
lmax=config["lmax"],
non_local=config["non_local"],
non_local_hidden_nf=config["non_local_hidden_nf"],
radius_cutoff=config["radius_cutoff"],
key=key,
)
model = load_weights_from_npz(model, weights_dir)
n_points, n_atoms = 100, 3
mol = {
"density": jnp.ones((2, n_points)) * 0.1,
"grad": jnp.zeros((2, 3, n_points)),
"kin": jnp.ones((2, n_points)) * 0.05,
"grid_coords": jnp.zeros((n_points, 3)),
"grid_weights": jnp.ones(n_points) * 0.01,
"coarse_0_atomic_coords": jnp.zeros((n_atoms, 3)),
}
E_xc = model.get_exc(mol)
print(f"E_xc = {E_xc:.10f} Ha")
JAX Autodiff
# Gradient of E_xc with respect to all inputs in one line
grads = jax.grad(model.get_exc)(mol)
print(f"dE/d(density): {grads['density'].shape}")
JIT Compilation
import equinox as eqx
@eqx.filter_jit
def get_exc_jit(m, mol):
return m.get_exc(mol)
# First call compiles; subsequent calls are fast
E = get_exc_jit(model, mol)
PySCF Integration
import jax
jax.config.update("jax_enable_x64", True)
from pyscf import gto
from skalax import SkalaFunctional, load_weights_from_npz, load_config, get_default_weights_dir
from skalax.pyscf import JaxSkalaKS
weights_dir = get_default_weights_dir()
config = load_config(weights_dir)
key = jax.random.PRNGKey(0)
model = SkalaFunctional(
lmax=config["lmax"],
non_local=config["non_local"],
non_local_hidden_nf=config["non_local_hidden_nf"],
radius_cutoff=config["radius_cutoff"],
key=key,
)
model = load_weights_from_npz(model, weights_dir)
mol = gto.M(
atom="O 0 0 0; H 0.757 0.586 0; H -0.757 0.586 0",
basis="sto-3g",
verbose=0,
)
ks = JaxSkalaKS(mol, xc=model)
energy = ks.kernel()
print(f"Total energy: {energy:.8f} Ha")
Input/Output Specification
Input Features
| Feature | Shape | Description |
|---|---|---|
density |
(2, n_points) |
Spin densities [α, β] |
grad |
(2, 3, n_points) |
Density gradients [spin, xyz, points] |
kin |
(2, n_points) |
Kinetic energy densities |
grid_coords |
(n_points, 3) |
Grid coordinates (Bohr) |
grid_weights |
(n_points,) |
Integration weights |
coarse_0_atomic_coords |
(n_atoms, 3) |
Atomic positions (Bohr) |
Outputs
| Method | Shape | Description |
|---|---|---|
model.get_exc(mol) |
() |
Scalar E_xc (Hartree) |
model.get_exc_density(mol) |
(n_points,) |
Energy density per grid point |
Model Architecture
Roughly 276k parameters, in three stages:
-
Input MLP. Per grid point, the 7 scalar features (spin densities, gradient norms, kinetic densities, and the α+β gradient norm) go through
Linear(7→256) → SiLU → Linear(256→256) → SiLU. Spin-swapped features are pushed through the same MLP and averaged, so the model is symmetric under α↔β. -
Non-local branch (optional,
lmax=3,radius_cutoff≈5 Bohr). The 256-dim scalar features are squeezed to 16 channels (pre_down_linear), then a fine→coarse tensor product (tp_down) aggregates to atomic centers, a coarse→fine tensor product (tp_up) broadcasts back to the grid, and a finalpost_up_linear(SiLU) mixes channels. Edge features use an exponential radial basis and spherical harmonics up tolmax. The non-local output is damped byexp(-ρ)and concatenated to the scalar features. -
Output MLP. The 256+16 dim features go through three
Linear(→256) → SiLUlayers, a finalLinear(→1), and aScaledSigmoid(scale=2.0). The scalar output is an enhancement factor multiplied against the LDA exchange density to give the per-point XC energy density.
Numerical Equivalence
On a handful of test cases the JAX implementation matches the PyTorch reference to machine precision:
| Test | Max |ΔE| |
|---|---|
get_exc (local only) |
0.00e+00 Ha |
get_exc (with non-local) |
1.14e-13 Ha |
get_exc_density |
1.17e-13 Ha |
More comprehensive benchmarks follow below.
Benchmarks
A few tests to check the correctness of the implementation. Note that results can be affected (positively or negatively) by the JAX PySCF wrapper I included, which explains the imperfect match with the PyTorch reference.
Forward pass equivalence
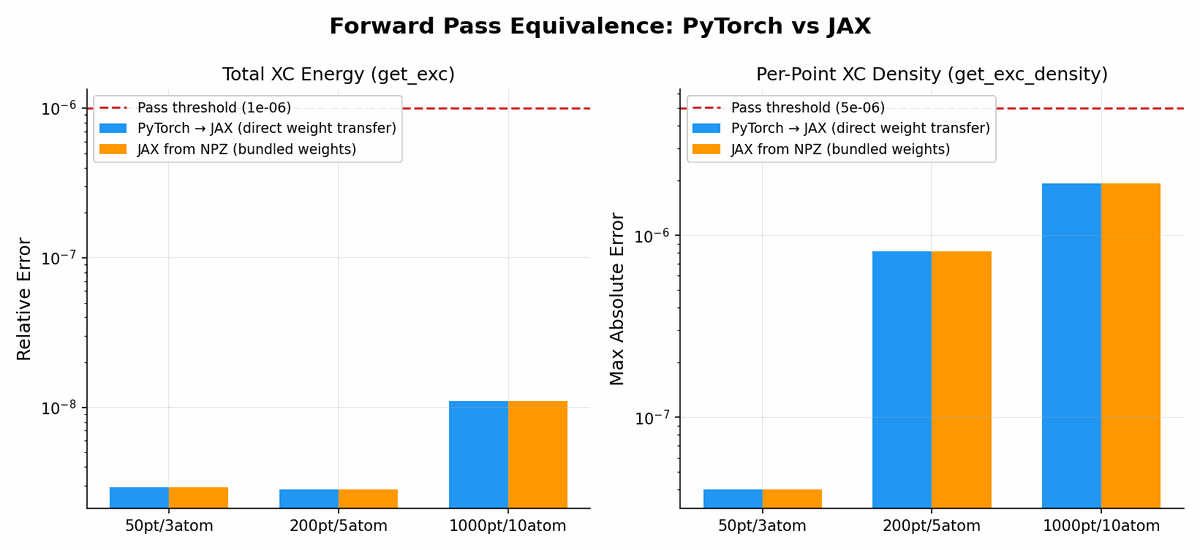
Energy profiles
The non-parallelity error (NPE) is crucial for the correctness of a prediction in chemistry. Here I compare the JAX implementation against the PyTorch reference:
Simple system:
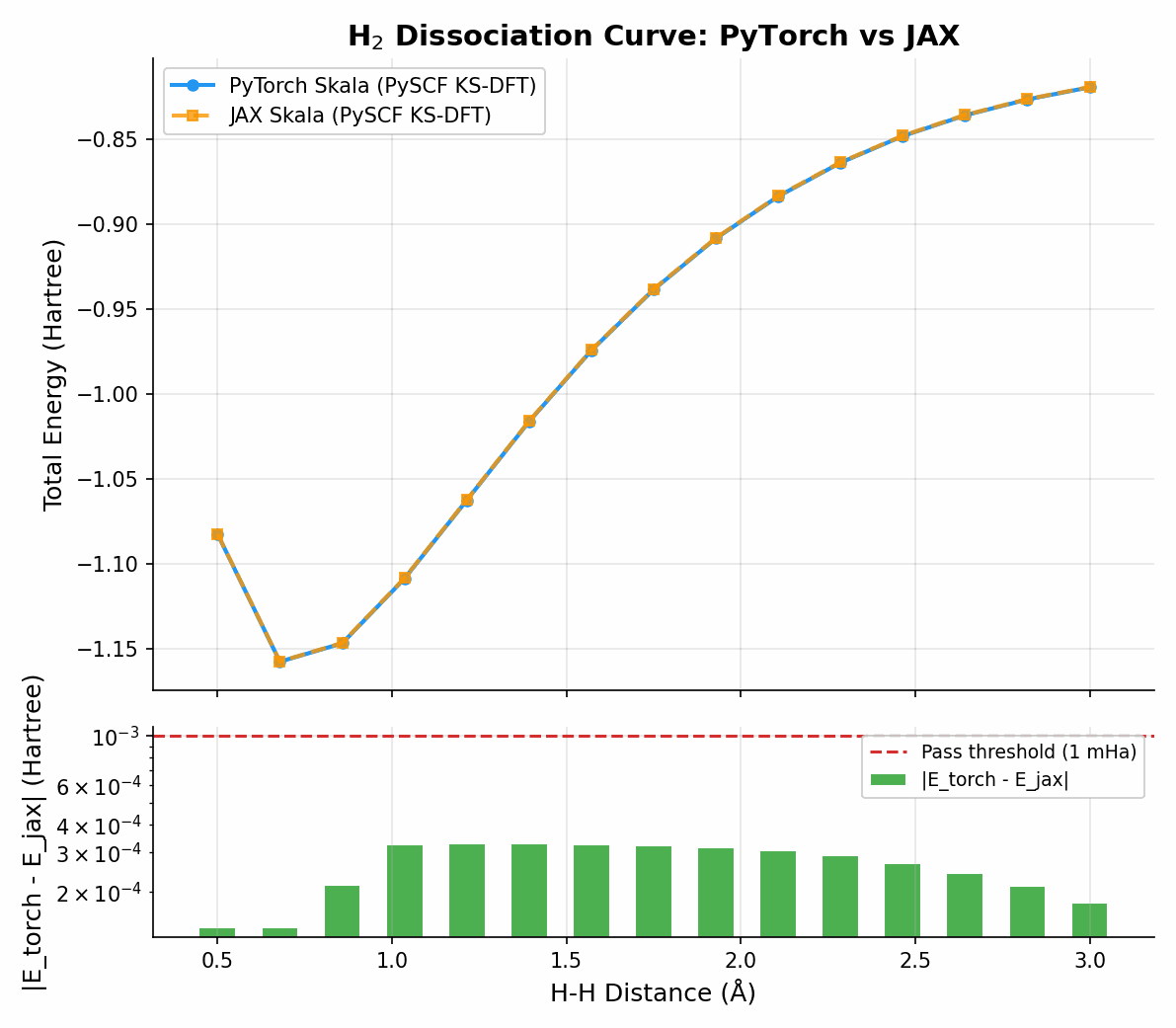
And more challenging:
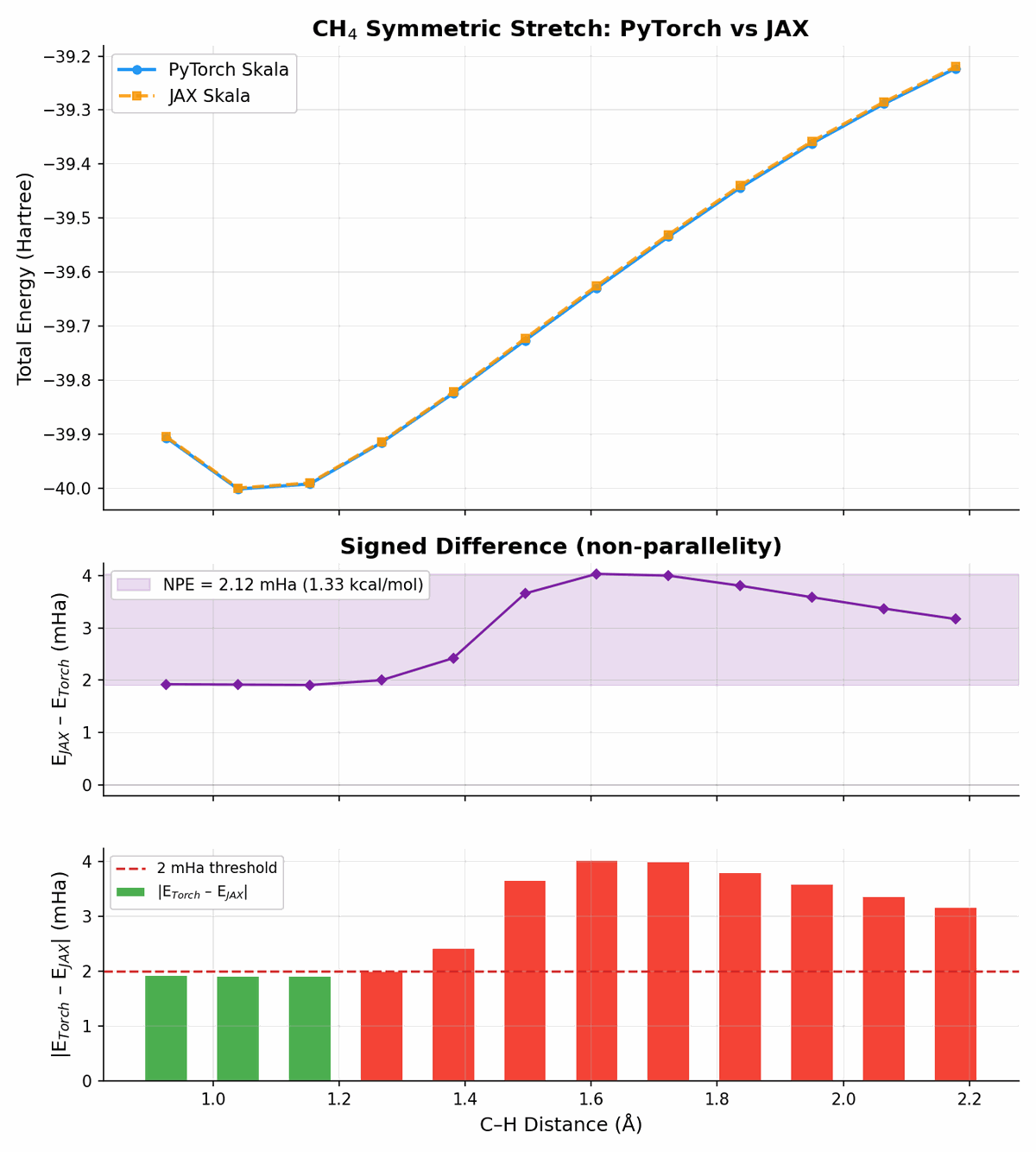
The curves agree well given that the two implementations share the same parameters but run on completely different backends (PyTorch vs JAX).
Dependencies
Tested versions
The following versions are known to work together (tested on CPU, Python 3.12):
| Package | Tested version | Role |
|---|---|---|
jax |
0.9.2 | Core |
jaxlib |
0.9.2 | Core |
equinox |
0.13.6 | Core |
e3nn-jax |
0.20.8 | Core |
numpy |
2.4.3 | Core |
skala (microsoft-skala) |
1.1.1 | Full install |
torch |
2.10.0 | Full install (via skala) |
pyscf |
2.12.1 | Full install (via skala) |
e3nn |
0.6.0 | Full install (via skala) |
dftd3 |
— | Full install (via skala, requires gfortran) |
huggingface_hub |
1.7.1 | Full install (via skala) |
opt_einsum_fx |
0.1.4 | Full install (via skala) |
ase |
— | Full install (via skala) |
qcelemental |
— | Full install (via skala) |
Cluster / custom installs
To install only the JAX core without PyTorch or Fortran dependencies:
pip install --no-deps skalax
pip install "jax>=0.4.0" "jaxlib>=0.4.0" "equinox>=0.11.0" "e3nn-jax>=0.20.0" "numpy>=1.21.0"
To pin specific versions for reproducibility (e.g. on a cluster):
pip install --no-deps skalax
pip install jax==0.9.2 jaxlib==0.9.2 equinox==0.13.6 "e3nn-jax==0.20.8" numpy==2.4.3
For a full install with PyTorch reference and PySCF integration, see Installation.
Tests
pytest tests/ -v
License
MIT License, see LICENSE.txt.
The pretrained weights bundled with this package are derived from the Skala model originally released by Microsoft Corporation under the MIT License (github.com/microsoft/skala).
Citation
If you use skalax, please cite the Skala paper:
@misc{luise2025skala,
title={Accurate and scalable exchange-correlation with deep learning},
author={Giulia Luise and Chin-Wei Huang and Thijs Vogels and Derk P. Kooi
and Sebastian Ehlert and Stephanie Lanius and Klaas J. H. Giesbertz
and Amir Karton and Deniz Gunceler and Megan Stanley
and Wessel P. Bruinsma and Lin Huang and Xinran Wei
and Jose Garrido Torres and Abylay Katbashev and Rodrigo Chavez Zavaleta
and B{\'a}lint M{\'a}t{\'e} and Roberto Sordillo and Yingrong Chen
and David B. Williams-Young and Christopher M. Bishop
and Jan Hermann and Rianne van den Berg and Paola Gori-Giorgi},
year={2025},
eprint={2506.14665},
archivePrefix={arXiv},
url={https://arxiv.org/abs/2506.14665}
}
If you additionally use this JAX implementation, please also cite:
@software{sokolov2025skalax,
author = {Sokolov, Igor O.},
title = {skalax: {JAX} implementation of the {Skala} neural exchange-correlation functional},
year = {2025},
url = {https://github.com/Brogis1/skalax},
license = {MIT}
}
Related Projects
- Skala (PyTorch): original PyTorch implementation
- e3nn-jax: equivariant neural networks for JAX
- Equinox: JAX neural network library
- PySCF: quantum chemistry in Python
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file skalax-1.0.5.tar.gz.
File metadata
- Download URL: skalax-1.0.5.tar.gz
- Upload date:
- Size: 1.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5dbe6f748b75ff9c19d6c6b349642467e38b098a005453f4621ef59d4627d2a3
|
|
| MD5 |
7d80b6561e81b543a434d271094f71d0
|
|
| BLAKE2b-256 |
a3cc2d6c1d8b0cb3daab20d6162add730925b36807b4cec0c63fd6b224679c8c
|
File details
Details for the file skalax-1.0.5-py3-none-any.whl.
File metadata
- Download URL: skalax-1.0.5-py3-none-any.whl
- Upload date:
- Size: 1.3 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a5d283d224db602e34fe1201dd7288a778cccbc1b96f26039d06a386a182f865
|
|
| MD5 |
a569c001c4e3d0a0007b6433d264f7f0
|
|
| BLAKE2b-256 |
c03d1cc06a9e108a6e1b322622b6ddb8add9fba767bcaa951ed5be35748aa89e
|












