The Heuristic Evolutionary Rule Optimization System (HEROS) is a supervised rule-based machine learning algorithm designed to agnostically model diverse 'structured' data problems and yield compact human interpretable solutions. This implementation is scikit-learn compatible.
Project description
Table of contents:
- Introduction
- Installation
- Input Data
- Using HEROS
- Hyperparameters
- Algorithm History
- Citing HEROS
- Futher Documentation
- License
- Contact
- Acknowledgements
Introduction
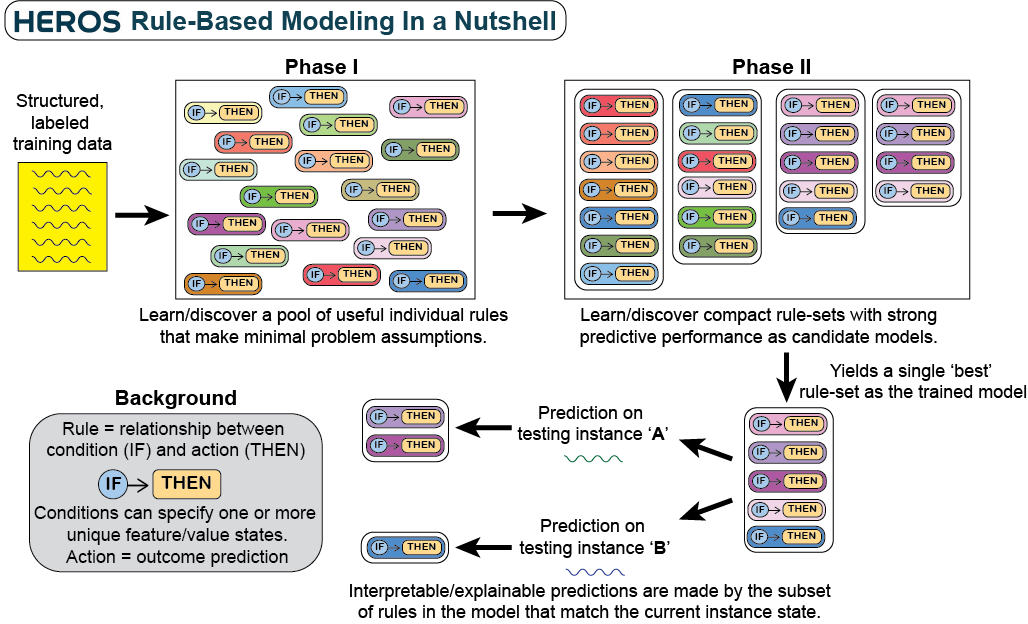
HEROS (Heuristic Evolutionary Rule Optimization System) is an evolutionary rule-based machine learning (ERBML) algorithm framework for supervised learning. It is designed to agnostically modeling simple/complex and/or clean/noisy problems (without hyperparameter optimization) and yield maximally human interpretable models. HEROS adopts a two-phase approach separating rule optimization, and rule-set (i.e. model) optimization, each with distinct multi-objective Pareto-front-based optimization. Rules are optimized based on maximizing rule-accuracy and instance coverage using a Pareto-inspired rule fitness function. Differently, models are optimized based on maximizing balanced accuracy and minimizing rule-set size using an NSGA-II-inspired evolutionary algorithm. This package is scikit-learn compatible. A simple visual summary of HEROS rule-based modeling is given below:
To date, HEROS functionality has been validated on binary classification problems, and has also passed bug checks on multiclass outcomes and data with a mix of categorical and quantitative features. This project is under active development with a number of improvements/expansions planned or in progress. For example, we will be expanding HEROS to support regression, and survival outcomes in future releases.
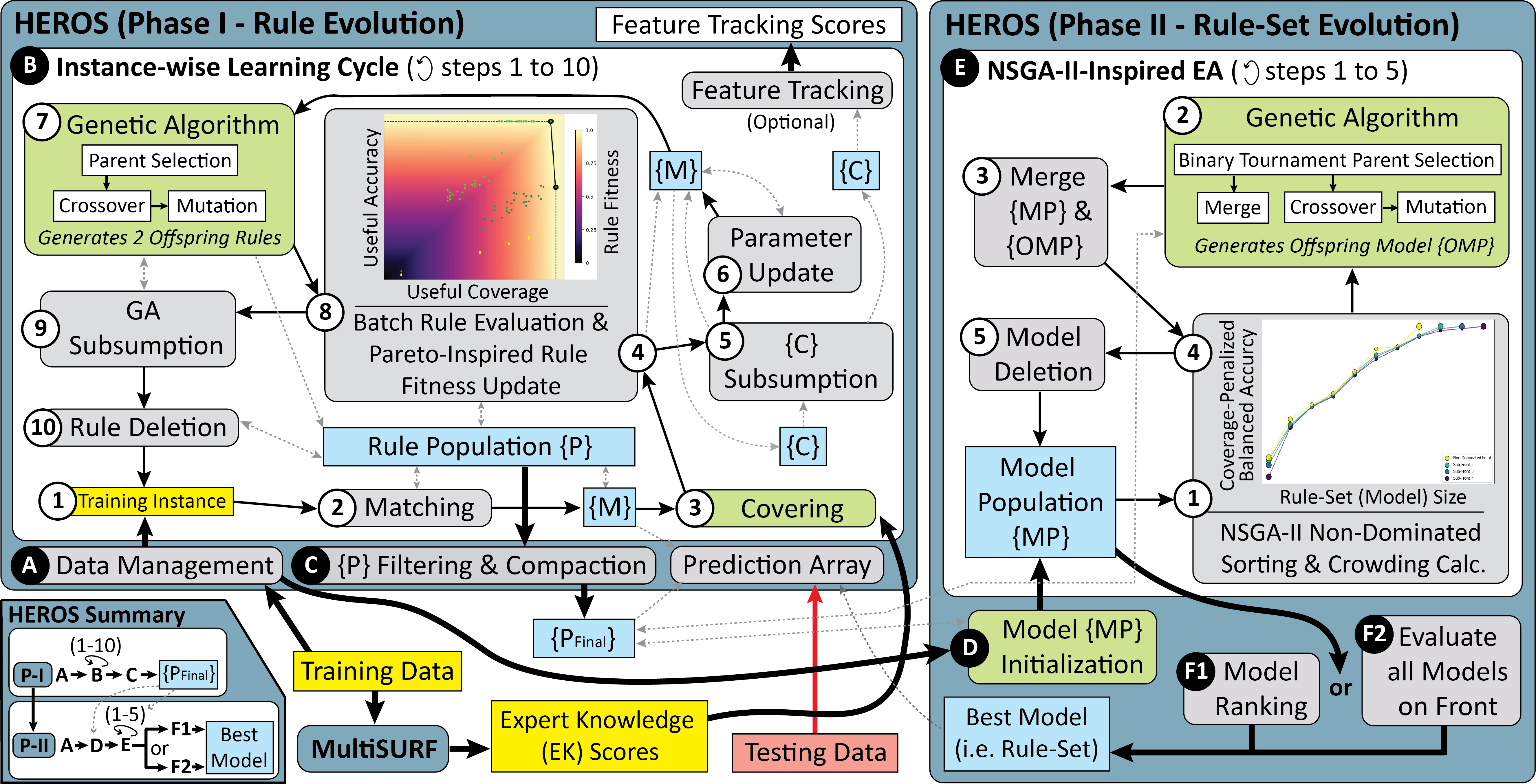
A schematic detailing how the HEROS algorithm works is given below:
Installation
HEROS can be installed with pip or by cloning this repository.
Pip
HEROS can most easily be installed using the following pip command:
pip install skheros
In order to run the HEROS_Demo_Notebook, download it and make sure to set the following notebook parameter to False in order to import HEROS from the above pip installation.
load_from_cloned_repo = False
Clone Respository
To install/run HEROS from this cloned repository, run the following commands from the desired folder:
git clone --single-branch https://github.com/UrbsLab/heros
cd heros
pip install -r requirements.txt
Input Data
HEROS's fit() method takes 'X', an array-like {n_samples, n_features} object of training instances, as well as 'y', an array-like {n_samples} object of training labels, like other standard scikit-learn classification algorithms.
Specifying Feature Types (Categorical vs. Quantiative)
The fit() method can (and should) also be passed 'cat_feat_indexes', an array-like max({n_features}) object of feature indexes in 'X' that are to be treated as categorical variables (where all others will be treated as quantitative by default).
Instance Identifiers
The fit() method can optionally be passed 'row_id', an array-like {n_samples} object of instance lables to link internal feature tracking scores with specific training instances.
Loading Expert Knowledge Scores for (Phase I) Rule-Covering (i.e. Initialization)
The fit() method can optionally be passed 'ek', an array-like {n_features} object of feature weights that probabilistically influence rule covering (i.e. rule-initialization), such that features with higher weights are more likely to be 'specified' within initialized rules.
Loading a Previously Trained (Phase I) Rule-Population
Lastly, the fit() method can optionally be passed 'pop_df', a dataframe object, including a previously trained HEROS-formatted rule population. This allows users to reboot progress from a previous run, or to manually add their own custom rules to the initial population.
Using HEROS
Demonstration Notebook
A Jupyter Notebooks has been included to demonstrate how HEROS (and it's functions) can be applied to train, evaluate, and apply models with a wide variety of saved outputs, visualizations and model prediction explanations. We strongly recommend exploring this demonstration notebook to get familiar with HEROS and its capabilities.
This notebook is currently set up to run by cloning this repository and running the included notebook.
Basic Run Command Walk-Through
As a simple example of HEROS data preparation and training:
# Data Preparation
train_data = pd.read_csv('evaluation/datasets/partitioned/gametes/A_uni_4add_CV_Train_1.txt', sep="\t")
outcome_label = 'Class'
X = train_data.drop(outcome_label, axis=1)
cat_feat_indexes = list(range(X.shape[1])) #all feature are categorical
X = X.values
y = train_df[outcome_label].values
# HEROS Initialization and Training
heros = HEROS(iterations=10000, pop_size=500, nu=1, model_iterations=100, model_pop_size=100)
heros = heros_trained.fit(X, y, cat_feat_indexes=cat_feat_indexes)
Once trained, HEROS can be applied to make predictions on testing data. Users have the option to choose the model to use; either (1) the top Phase II model from the model-pareto front (selected based on maximizing testing performance, maximizing instance coverage, and, if possible, minimizing rule-count) - RECOMMENDED, (2) the default top Phase II model (automatically selected based on training performance), or (3) the entire Phase I rule population.
Below is an example of the first option (RECOMMENDED):
# Data Preparation
test_data = pd.read_csv('evaluation/datasets/partitioned/gametes/A_uni_4add_CV_Test_1.txt', sep="\t")
X_test = test_data.drop(outcome_label, axis=1)
X_test = X_test.values
y_test = test_data[outcome_label].values
# HEROS Prediction (Model selection via Phase II Model Testing Evaluation) and Performance Report
best_model_index = heros.auto_select_top_model(X_test,y_test)
predictions = heros.predict(X_test, target_model=best_model_index)
print(classification_report(predictions, y_test, digits=8))
To get predicitions with the second option, after preparing the data we would run the following:
# HEROS Prediction (Model selection via Phase II Default Model Selection) and Performance Report
predictions = heros.predict(X_test)
print(classification_report(predictions, y_test, digits=8))
To get predictions with the third option, after preparing the data we would run the following:
# HEROS Prediction (Whole Phase I Rule Population Applied as Model) and Performance Report
predictions = heros.predict(X_test,whole_rule_pop=True)
print(classification_report(predictions, y_test, digits=8))
Differently, HEROS will return prediction probabilities using the following:
predictions = heros.predict_proba(X_test, target_model=best_model_index)
HEROS can also return whether each instance is covered (i.e. at least one rule matches it in the given 'model') using the following:
predictions = heros.predict_covered(X_test, target_model=best_model_index)
Lastly, HEROS can give direct explanations of individual model predictions using the following:
testing_instance = X_test[0] # Testing instance index 0 arbitrarily chosen here
heros.predict_explanation(testing_instance, feature_names, target_model=best_model_index)
The parameter, feature_names, is the ordered list of original feature names from the training dataset.
Below is a simple example prediction explanation for a HEROS model trained on the 6-bit multiplexer problem:
PREDICTION REPORT ------------------------------------------------------------------
Outcome Prediction: 0
Model Prediction Probabilities: {0: 1.0, 1: 0.0}
Instance Covered by Model: Yes
Number of Matching Rules: 1
PREDICTION EXPLANATION -------------------------------------------------------------
Supporting Rules: --------------------
6 rule copies assert that IF: (A_0 = 0) AND (A_1 = 0) AND (R_0 = 0) THEN: predict outcome '0' with 100.0% confidence based on 68 matching training instances (15.11% of training instances)
Contradictory Rules: -----------------
No contraditory rules matched.
In the case that multiple rules match an instance, they will all be displayed in a similar human-readable format.
Hyperparameters
Key Hyperparameters
While HEROS has a number of available hyperparameters only a few are expected to have a significant impact on algorithm performance (see first table below). In general, setting iterations and pop_size to larger integers is expected to improve training performance, but will require longer Phase I run times, and the same is true for model_iterations and model_pop_size with respect to Phase II. The nu parameter should always be set to 1 unless the user is confident that they are modeling a problem that can achieve 100% testing accuracy (i.e. a problem with no signal noise).
| Hyperparameter | Description | Type/Options | Default Value |
|---|---|---|---|
| iterations | Number of (rule population) learning iterations (Phase I) | int | 100000 |
| pop_size | Maximum 'micro' rule-population size (Phase I) | int | 1000 |
| model_iterations | Number of (model population) learning iterations (Phase II) | int | 500 |
| model_pop_size | Maximum model-population size (Phase II) | int | 100 |
| nu | Power parameter used to determine the importance of high rule-accuracy when calculating fitness (Phases I & II) | int | 1 |
Other Hyperparameters
The table below gives other HEROS hyperparameters that should generally be left as-is, unless experimenting with general algorithm configuration optimization.
| Hyperparameter | Description | Type/Options | Default Value |
|---|---|---|---|
| beta | Learning parameter - used in calculating average match set size (Phase I) | float | 0.2 |
| theta_sel | The fraction of the correct set to be included in tournament selection (Phases I & II) | float | 0.5 |
| cross_prob | The probability of applying crossover in rule discovery with the genetic algorithm (Phases I & II) | float | 0.8 |
| mut_prob | The probability of mutating a position within an offspring rule (Phases I & II) | float | 0.04 |
| merge_prob | The probablity of the merge operator being used during model offspring generation (Phase II) | float | 0.1 |
| new_gen | Proportion of maximum pop size used to generate an model offspring population each generation (Phase II) | float | 1.0 |
| model_pop_init | Model population initialization method (Phase II) | 'random', 'probabilistic', 'bootstrap', or 'target_acc' | 'target_acc' |
| subsumption | Specify subsumption strategy(s) to apply i.e. genetic algorithm, correct set, both or None (Phase I) | 'ga', 'c', 'both', or None | 'both' |
| rsl | Rule specificity limit (automatically determined when 0) (Phase I) | int | 0 |
| compaction | Specifies type of rule-compaciton to apply at end of rule population training (if any) (Phase I) | 'sub' or None | 'sub' |
| random_state | The seed value needed to generate a random number (Phases I & II) for reproducibility | int or None | None |
Performance Tracking Hyperparameters
This next table gives optional hyperparameters that control HEROS performance tracking options (over the course of learning). These are useful when interested in analyzing rule/model learning across iterations.
| Hyperparameter | Description | Type/Options | Default Value |
|---|---|---|---|
| track_performance | (For detailed algorithm evaluation) Activates performance tracking when > 0. Value indicates how many iteration steps to wait to gather tracking data (Phase I) | int | 0 |
| model_tracking | (For detailed algorithm evaluation) Activates tracking of top model performance across training iterations (Phase II) | True or False | False |
| stored_rule_iterations | (For detailed algorithm evaluation) Specifies iterations where a copy of the rule population is stored (Phase I) | comma-separated string of ints (e.g. 500,1000,5000) | None |
| stored_model_iterations | (For detailed algorithm evaluation) Specifies iterations where a copy of the model population is stored (Phase II) | comma-separated string of ints (e.g. 50,100,500) | None |
| verbose | Boolean flag to run in 'verbose' mode - display run details | True or False | False |
In-Development Hyperparameters
This last table gives hyperparameters that are 'in-development' and should be left as-is. Currently we have only tested HEROS for binary classification. While it should work for multi-class classification it has not been fully tested and may yield bugs.
| Hyperparameter | Description | Type/Options | Default Value |
|---|---|---|---|
| outcome_type | Defines the type of outcome in the dataset | 'class','quant' | 'class' |
| fitness_function | Defines the Phase I fitness function used by HEROS. The 'accuracy' option should only be used for clean-signal problems. | 'accuracy','pareto' | 'pareto' |
| feat_track | Feature tracking strategy applied | None, 'add','wh','end' | None |
| rule_pop_init | Specifies rule population pre-initialization method | None, 'load','dt' | None |
fit() Parameters
In addition to the typical X and y parameters for HEROS's fit function users can utilize the following fit() parameters:
| Parameter | Description | Type/Options | Default Value |
|---|---|---|---|
| row_id | List of unique row/instance identifiers that can be included for instance-related outputs | None, array-like {n_samples} | None |
| cat_feat_indexes | List of feature indexes to be treated as categorical (vs. quantitative) | None, array-like max({n_features}) | None |
| pop_df | HEROS-formatted rule population dataframe (to manually initialize rule-population) | None, HEROS {P} dataframe | None |
| ek | List of expert knowledge (EK) scores which activats HEROS EK covering and mutation | None, array-like {n_features} | None |
Algorithm History
HEROS directly descends from a lineage of "Michigan-Syle" Learning Classifier System (LCS) algorithms including XCS, UCS, and ExSTraCS.
External Research
XCS is, to date, the best-known and most popular LCS algorithm, having introduced the accuracy-based rule-fitness (1995). XCS simplified and refined the original LCS algorithm concept described by John Holland (1975) in "Adaptation in natural and artificial systems". Like earlier LCS algorithms, XCS was designed as a reinforcement learning algorithm that could also easily be applied to supervised learning problems. Later, UCS was introduced, adapting XCS to the more specific task of supervised learning.
In the years 2000 and 2013, two Fuzzy LCS algorithms were proposed pioneering the hybridization of Michigan and Pittsburgh-style LCSs with distinct fitness functions for rule vs. rule-set discovery. In 2022-2023, the 'SupRB' LCS algorithm was proposed for regression tasks that adopted this rule vs. rule-set optimization concept, relying on distinct, weighted, multi-objective fitness functions.
Internal Research
In 2012, our lab developed a number of algorithmic improvements for the UCS algorithm framework. We developed statistical and visualization-guided knowledge discovery stratagies for global interpretation of an LCS rule-population, and introduced the novel mechanisms of expert knowledge covering and attribute (a.k.a. feature) tracking and feedback to improve algorithm performance in complex/noisy problem domains. Later in 2013, we developed a simple rule compaction/filtering strategy for noisy problems.
In 2014, we introduced ExSTraCS, which combined several proposed LCS advacements up to that point: (1) expert knowledge covering, (2) feature tracking and feedback, (3) rule filtering/compaction for noisy problems, and (4) a mixed discrete-continuous attribute-list rule representation (similar to ALKR introduced by Jaume Bacardit).
In 2015, we released ExSTraCS 2.0 which further extended ExSTraCS to better deal with problem/dataset scalability, by introducing (1) an automatic rule-specificity limit, (2) updates to expert knowledge covering and (3) utilization of the TuRF wrapper algorithm in combination with Relief-algorithms to generate statistically-derrived expert knowledge scores. Later that year, ExSTraCS 2.0 also extended with a novel protype strategy to extend it to quantiative outcome modeling and a multi-objective fitness function seeking to optimize both the accuracy and correct coverage of rules to improve performance in noisy domains (yielding mixed reliability).
In 2016, we expanded on this multi-objective fitness function work, introducing a novel pareto-inspired multi-objective rule fitness strategy, yielding much more promising results. Later in 2018, we revisited the topic of feature tracking, and how to best update these scores during LCS training.
In 2017 Drs. Will Browne and Ryan Urbanowicz published an Introductory Textbook on Learning Classifier Systems, which was paired with a very simple "Educational Learning Classifier System" (eLCS).
In 2020, toward making LCS algorithms more accessible to users, we re-implemented eLCS as a scikit learn compatible package (scikit-eLCS).
In 2021, we combined our earlier work with (1) ExSTraCS 2.0, (2) developing a statistical and visualization-guided knowledge discovery strategies, and (3) attribute tracking, into LCS-DIVE, an automated rule-based machine learning pipeline for characterizing complex associations in classification problems. That paper also introduced a scikit-learn implementation of ExSTraCS 2.0 (scikit-ExSTraCS).
In 2023, we released an automated machine learning analysis pipeline called STREAMLINE, which included a variety of well-known machine learning modeling algorithms along side of scikit-ExStraCS, scikit-eLCS, and a new scikit-learn compatible implementation of XCS called scikit-XCS
Most recently, in 2024, we released Survival-LCS an LCS algorithm adapted to the task of survival-data analysis (with censoring), built based on the ExSTraCS 2.0 algorithm.
Citing HEROS
If you use HEROS in a scientific publication, cite the following paper:
Gabe Lipschutz-Villa, Harsh Bandhey, Ruonan Yin, Malek Kamoun, Ryan J. Urbanowicz. [Rule-based Machine Learning: Separating Rule and Rule-Set Pareto-Optimization for Interpretable Noise-Agnostic Modeling] 2025. (In Press)
BibTeX entry:
Not yet available
Futher Documentation:
Further code documentation regarding the HEROS API is under development
License:
HEROS adopts a custom academic software license. Please review 'LICENSE' for details.
Contact
Please email Ryan.Urbanowicz@cshs.org for any application or collaboration inquiries related to HEROS.
Commercial entities or for commercial use of the Software: please contact CSTechTransfer@cshs.org for licensing opportunities.
Acknowledgements
The study was supported by Cedars Sinai Medical Center and NIH grants R01 AI173095, U01 AG066833 and P30 AG0373105. We thank Drs. John Holmes and Jason Moore for their mentorship and and research insights regarding rule-based machine learning for biomedicine, and Robert Zhang, who implemented scikit-ExSTraCS and prototyped an early batch-learning version of ExSTraCS.
Code Contributors
- Ryan Urbanowicz - Developed algorithm concepts, implemented algorithm, led debugging and evaluation
- Gabriel Lipschutz-Villa - Prototyped implementation of Phase II, model initialization strategies, and phase alternation (in development)
- Harsh Bandhey - Prototyped implementation of random forest rule initialization (in development)
- Khoi Dinh - Prototyped implementation of model interpretation visualization
- Ruonan Yin - Developed strategy for calculating distance from the rule-pareto front for Phase I rule fitness
- Robert Zhang - Prototyped strategy for rule batch-learning (adapted for HEROS)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file skheros-0.2.4.tar.gz.
File metadata
- Download URL: skheros-0.2.4.tar.gz
- Upload date:
- Size: 22.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.17
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7dcfd8c657c92f7373695322e6986c2abe820ae9683fc289faaf919172236b4f
|
|
| MD5 |
c07236221def6755ae7b4d25b46eef9d
|
|
| BLAKE2b-256 |
9425ef2b1c8b5384b015cdc12eca4a392621055601a43eea4da6621e65be78ce
|
File details
Details for the file skheros-0.2.4-py3-none-any.whl.
File metadata
- Download URL: skheros-0.2.4-py3-none-any.whl
- Upload date:
- Size: 12.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.17
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
78a5f9dab654ba655e9bae469021f71b16c49f9241ab763632edfb2b28fb7d88
|
|
| MD5 |
874bcf824be9488fc1609988da762470
|
|
| BLAKE2b-256 |
b208238be404efaf400635f3a74a517b25e9c70acbdc168c1da9b3cdbd49c94a
|