Scikit-learn models hyperparameters tuning and features selection, using evolutionary algorithms
Project description

Sklearn-genetic-opt
scikit-learn models hyperparameters tuning and feature selection, using evolutionary algorithms.
This is meant to be an alternative to popular methods inside scikit-learn such as Grid Search and Randomized Grid Search for hyperparameters tuning, and from RFE (Recursive Feature Elimination), Select From Model for feature selection.
Table of Contents
Sklearn-genetic-opt Overview - Main Features - Demos on Features
Installation - Basic Installation - Full Installation with Extras
Usage - Hyperparameters Tuning - Feature Selection
Documentation - Stable - Latest - Development
Changelog
Important Links
Source Code
Contributing
Testing
Sklearn-genetic-opt uses evolutionary algorithms from the DEAP (Distributed Evolutionary Algorithms in Python) package to choose the set of hyperparameters that optimizes (max or min) the cross-validation scores, it can be used for both regression and classification problems.
Documentation is available here
Main Features:
GASearchCV: Main class of the package for hyperparameters tuning, holds the evolutionary cross-validation optimization routine.
GAFeatureSelectionCV: Main class of the package for feature selection.
Algorithms: Set of different evolutionary algorithms to use as an optimization procedure.
Callbacks: Custom evaluation strategies to generate early stopping rules, logging (into TensorBoard, .pkl files, etc) or your custom logic.
Schedulers: Adaptive methods to control learning parameters.
Plots: Generate pre-defined plots to understand the optimization process.
MLflow: Build-in integration with mlflow to log all the hyperparameters, cv-scores and the fitted models.
Demos on Features:
Visualize the progress of your training:
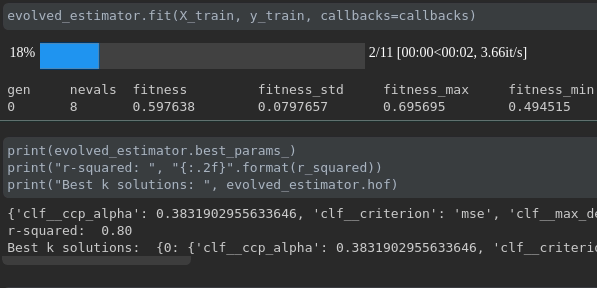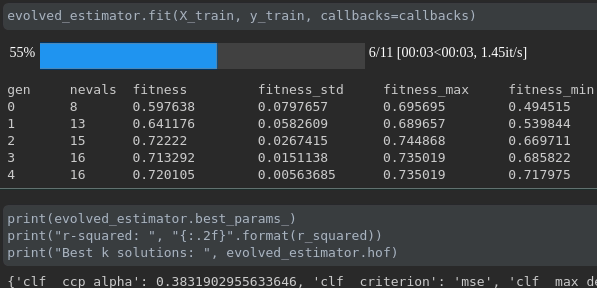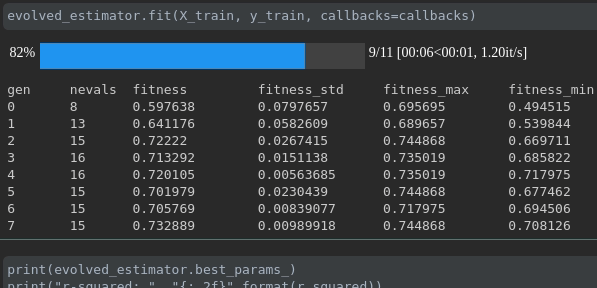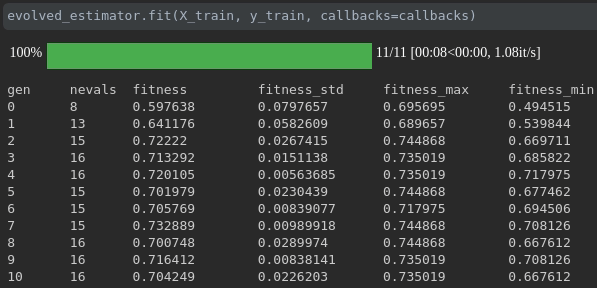
Real-time metrics visualization and comparison across runs:
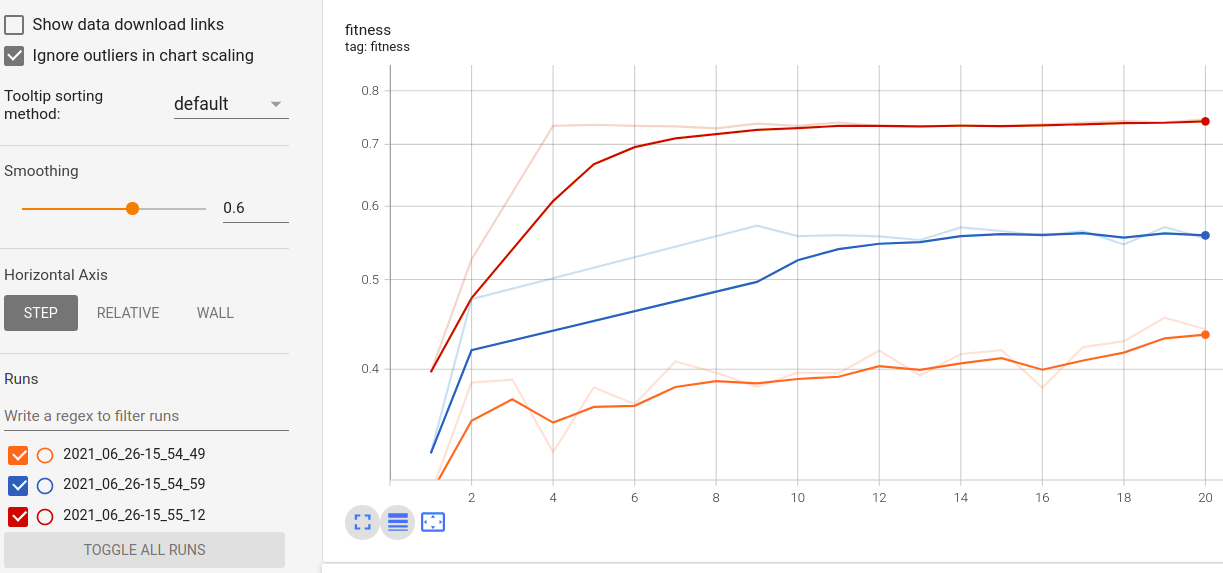
Sampled distribution of hyperparameters:
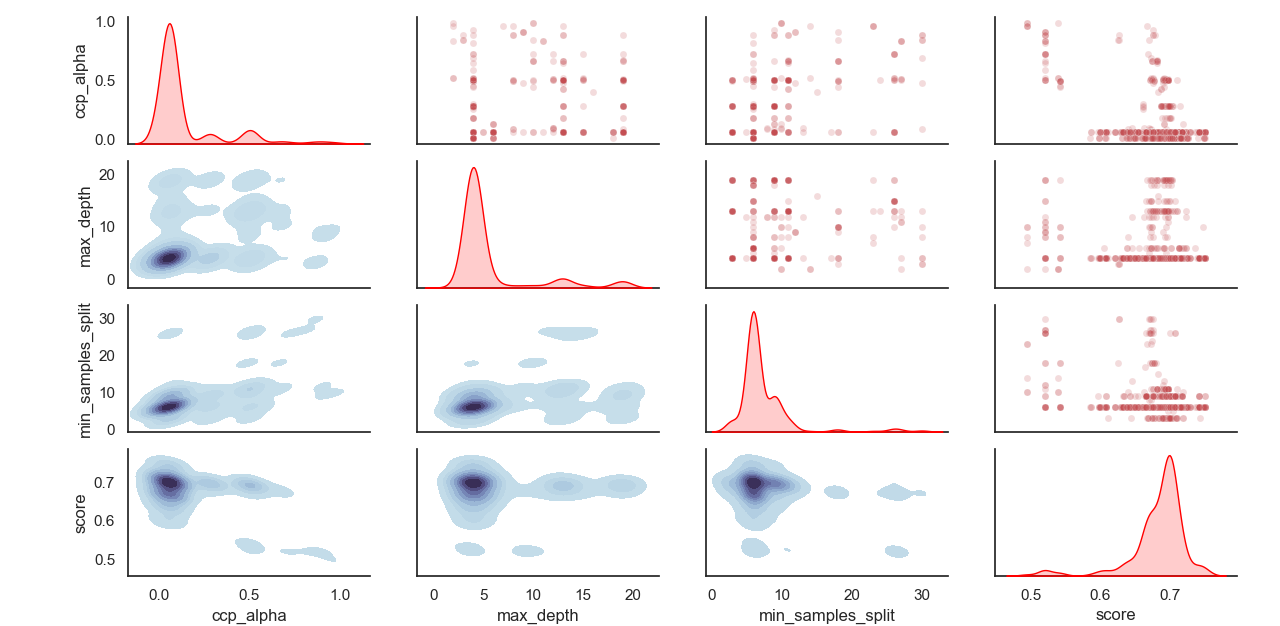
Artifacts logging:
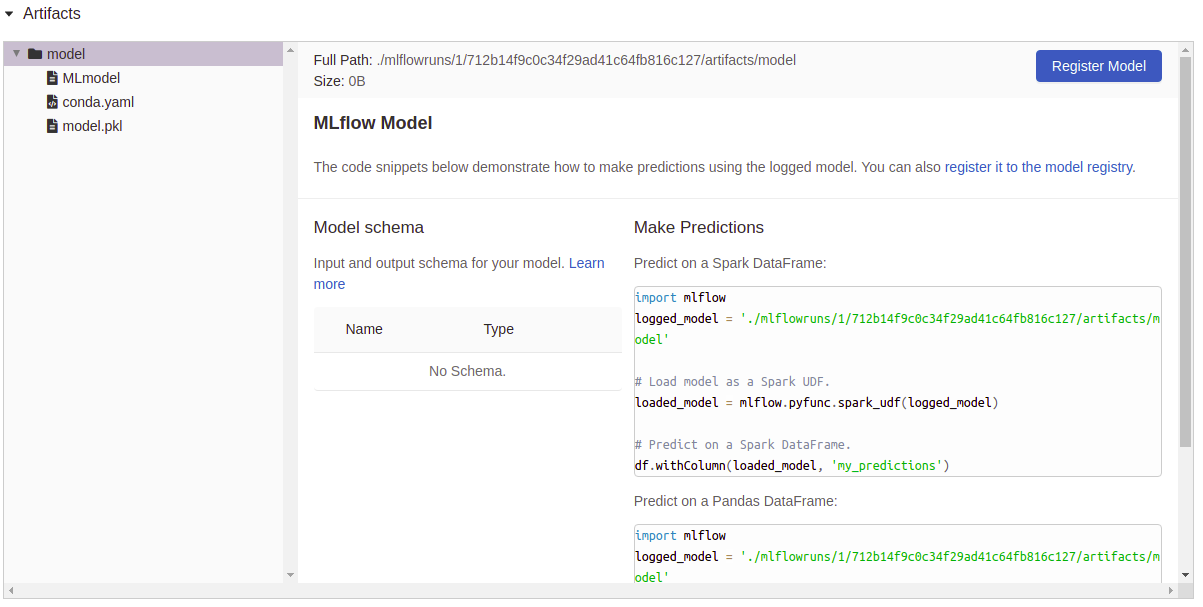
Usage:
Install sklearn-genetic-opt
It’s advised to install sklearn-genetic using a virtual env, inside the env use:
pip install sklearn-genetic-opt
If you want to get all the features, including plotting, tensorboard and mlflow logging capabilities, install all the extra packages:
pip install sklearn-genetic-opt[all]
Example: Hyperparameters Tuning
Quick Start (Minimal Example)
Here is a basic example of how to run GASearchCV on a scikit-learn model:
from sklearn_genetic import GASearchCV
from sklearn_genetic.space import Continuous, Categorical, Integer
from sklearn.ensemble import RandomForestClassifier
from sklearn.datasets import load_iris
X, y = load_iris(return_X_y=True)
# Defines the possible values to search
param_grid = {'min_weight_fraction_leaf': Continuous(0.01, 0.5, distribution='log-uniform'),
'bootstrap': Categorical([True, False]),
'max_depth': Integer(2, 30)}
evolved_estimator = GASearchCV(estimator=RandomForestClassifier(),
cv=3,
scoring="accuracy",
population_size=10,
generations=5,
param_grid=param_grid)
evolved_estimator.fit(X, y)
print(evolved_estimator.best_params_)
print(evolved_estimator.best_score_)from sklearn_genetic import GASearchCV
from sklearn_genetic.space import Continuous, Categorical, Integer
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split, StratifiedKFold
from sklearn.datasets import load_digits
from sklearn.metrics import accuracy_score
data = load_digits()
n_samples = len(data.images)
X = data.images.reshape((n_samples, -1))
y = data['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
clf = RandomForestClassifier()
# Defines the possible values to search
param_grid = {'min_weight_fraction_leaf': Continuous(0.01, 0.5, distribution='log-uniform'),
'bootstrap': Categorical([True, False]),
'max_depth': Integer(2, 30),
'max_leaf_nodes': Integer(2, 35),
'n_estimators': Integer(100, 300)}
# Seed solutions
warm_start_configs = [
{"min_weight_fraction_leaf": 0.02, "bootstrap": True, "max_depth": None, "n_estimators": 100},
{"min_weight_fraction_leaf": 0.4, "bootstrap": True, "max_depth": 5, "n_estimators": 200},
]
cv = StratifiedKFold(n_splits=3, shuffle=True)
evolved_estimator = GASearchCV(estimator=clf,
cv=cv,
scoring='accuracy',
population_size=20,
generations=35,
param_grid=param_grid,
n_jobs=-1,
verbose=True,
use_cache=True,
warm_start_configs=warm_start_configs,
keep_top_k=4)
# Train and optimize the estimator
evolved_estimator.fit(X_train, y_train)
# Best parameters found
print(evolved_estimator.best_params_)
# Use the model fitted with the best parameters
y_predict_ga = evolved_estimator.predict(X_test)
print(accuracy_score(y_test, y_predict_ga))
# Saved metadata for further analysis
print("Stats achieved in each generation: ", evolved_estimator.history)
print("Best k solutions: ", evolved_estimator.hof)Example: Feature Selection
from sklearn_genetic import GAFeatureSelectionCV, ExponentialAdapter
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
import numpy as np
data = load_iris()
X, y = data["data"], data["target"]
# Add random non-important features
noise = np.random.uniform(5, 10, size=(X.shape[0], 5))
X = np.hstack((X, noise))
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=0)
clf = SVC(gamma='auto')
mutation_scheduler = ExponentialAdapter(0.8, 0.2, 0.01)
crossover_scheduler = ExponentialAdapter(0.2, 0.8, 0.01)
evolved_estimator = GAFeatureSelectionCV(
estimator=clf,
scoring="accuracy",
population_size=30,
generations=20,
mutation_probability=mutation_scheduler,
crossover_probability=crossover_scheduler,
n_jobs=-1)
# Train and select the features
evolved_estimator.fit(X_train, y_train)
# Features selected by the algorithm
features = evolved_estimator.support_
print(features)
# Predict only with the subset of selected features
y_predict_ga = evolved_estimator.predict(X_test)
print(accuracy_score(y_test, y_predict_ga))
# Transform the original data to the selected features
X_reduced = evolved_estimator.transform(X_test)Changelog
See the changelog for notes on the changes of Sklearn-genetic-opt
Important links
Official source code repo: https://github.com/rodrigo-arenas/Sklearn-genetic-opt/
Download releases: https://pypi.org/project/sklearn-genetic-opt/
Issue tracker: https://github.com/rodrigo-arenas/Sklearn-genetic-opt/issues
Stable documentation: https://sklearn-genetic-opt.readthedocs.io/en/stable/
Source code
You can check the latest development version with the command:
git clone https://github.com/rodrigo-arenas/Sklearn-genetic-opt.git
Install the development dependencies:
pip install -r dev-requirements.txt
Check the latest in-development documentation: https://sklearn-genetic-opt.readthedocs.io/en/latest/
Contributing
Contributions are more than welcome! There are several opportunities on the ongoing project, so please get in touch if you would like to help out. Make sure to check the current issues and also the Contribution guide.
Big thanks to the people who are helping with this project!
Testing
After installation, you can launch the test suite from outside the source directory:
pytest sklearn_genetic
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sklearn_genetic_opt-0.12.0.tar.gz.
File metadata
- Download URL: sklearn_genetic_opt-0.12.0.tar.gz
- Upload date:
- Size: 34.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.11.1 requests/2.32.3 setuptools/72.1.0 requests-toolbelt/1.0.0 tqdm/4.66.5 CPython/3.10.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4e526cc62b424706d6158048a0b50ecb0162fe8015d0c8889aaec1a2d40c0ac6
|
|
| MD5 |
de3b9b001b4478c47ffbda614b5ce4f9
|
|
| BLAKE2b-256 |
41394b3ff7d5ef107bda7cbf155a376188074c0a50c5295dacc9c487bb36185a
|
File details
Details for the file sklearn_genetic_opt-0.12.0-py3-none-any.whl.
File metadata
- Download URL: sklearn_genetic_opt-0.12.0-py3-none-any.whl
- Upload date:
- Size: 37.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.3.0 pkginfo/1.11.1 requests/2.32.3 setuptools/72.1.0 requests-toolbelt/1.0.0 tqdm/4.66.5 CPython/3.10.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7e0b9b5c0db51b822bf731a05f14fbf2a6a79b1f6b8418fdb077ed76cf134365
|
|
| MD5 |
4d95d8083f8efc9e71d7dedad33113c8
|
|
| BLAKE2b-256 |
5c1015c3c2e30b4393a04469761c99ffbee7a00e1eb255b418c08197a7261312
|

















