Your package description
Project description
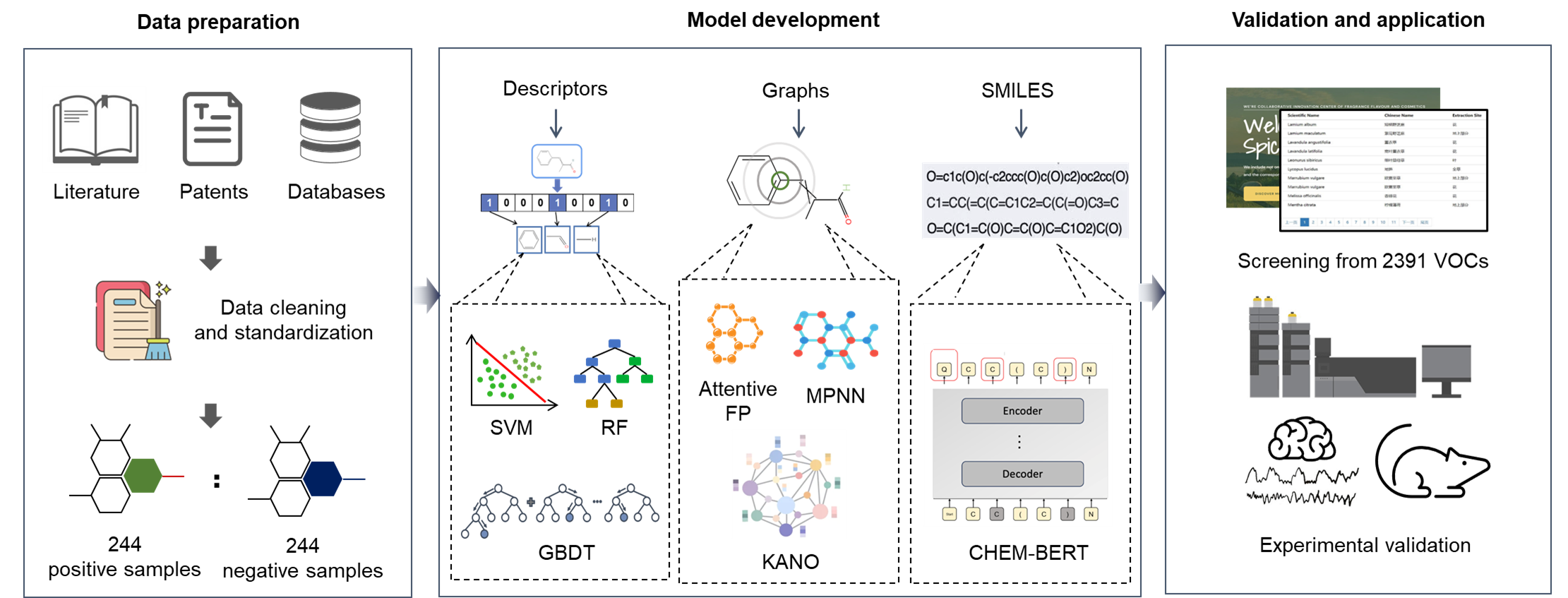
Machine Learning-Guided Mapping Sleep-Promoting Volatiles in Aromatic Plants
Project Description
This repository provides a machine-learning pipeline for identifying sleep-promoting volatile organic compounds (VOCs) from aromatic plants, including pretrained base models and a stacking predictor for quick inference.
Dependency
The code has been tested in the following environment:
| Package | Version |
|---|---|
| Python | 3.8.16 |
| Conda | 23.5.0 |
| RDKit | 2023.3.1 |
| Scikit-learn | 1.0.2 |
How to Use
Installation
Option A: From PyPI (simplest)
pip install sleep-model
Option B: Conda environment (recommended for RDKit/DeepChem)
conda env create -f environment.yaml -n sleep_model
conda activate sleep_model
# install this project from source (editable or regular)
python -m pip install -e .
# python -m pip install .
Option C: uv (fast installer; lockfile included)
python -m pip install uv
py -m venv .venv
.\.venv\Scripts\Activate.ps1 # Windows PowerShell
uv sync
File Structure
├── data/ # Input data files
├── data_analysis/ # Data processing and analysis
├── models/ # Pretrained base model files for Stacking model training
│ ├── RF/
│ │ ├── rf_MACCSkeys_random_0.ipynb
│ │ ├── rf_RDkit_random_0.ipynb
│ ├── SVM/
│ │ ├── svm_MACCSkeys_random_3.ipynb
│ ├── XGB/
│ │ ├── xgb_MACCSkeys_random_0.ipynb
│ │── stacking_predict.ipynb
├── predict_smiles.py
└── README.md
These four models (rf_MACCSkeys, rf_RDkit, svm_MACCSkeys, xgb_MACCSkeys) are the base models that we use to train the final stacking model.
Predicting
Command-line (console script)
After installation, a console command is available:
sleep-predict --smiles "CC(=O)OC1=CC=CC=C1C(=O)O"
As a Python module
python -m predict_smiles --smiles "CC(=O)OC1=CC=CC=C1C(=O)O"
Batch prediction from CSV
Predict for a CSV file containing a SMILES column (default column name: smiles):
python predict_smiles.py --csv example/input.csv --out example/preds.csv
Customize the SMILES column name and encoding when needed (e.g., column SMILES):
python predict_smiles.py --csv example/input.csv --out example/preds.csv --smiles-column SMILES --input-encoding utf-8
Notes
- Models and training data are loaded from the installed package resources (project
models/anddata/GABAA.csv). Ensure they are present if running from source. - If the console command is not found on Windows, re-activate your environment or run the module form.
Troubleshooting
- RDKit/DeepChem wheels can be environment-specific. If installation via
pipfails, prefer the Conda-based installation. - If you see a file-not-found error for
models/ordata/GABAA.csv, run from the project root or install the project so resources are available in the environment.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file sleep_model-0.1.3-py3-none-any.whl.
File metadata
- Download URL: sleep_model-0.1.3-py3-none-any.whl
- Upload date:
- Size: 102.3 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b5b802457e52bad4450673794ab8f04f0157fe2e883f7b74777c99cbac44b32b
|
|
| MD5 |
b569bfd86fa93658dd549b5f1c02de59
|
|
| BLAKE2b-256 |
137b0c386c7c6fbe66af452a74d71d8d43a6e4dfdab584d6f59932709611d13c
|