smarts.bio bioinformatics AI agent — JupyterLab extension
Project description
AI Bioinformatics for JupyterLab (smarts.bio)
The bioinformatics AI agent that lives in your notebook. Design proteins, run GPU pipelines, search every major database, and visualize results — without leaving JupyterLab.
What you can do
Ask the agent anything in plain English and watch it work:
"Use Boltz to design a nanobody that binds the RBD of SARS-CoV-2 spike protein — give me 5 candidates ranked by predicted affinity."
"Run RFdiffusion to generate 10 novel enzyme scaffolds around this active site geometry, then score them with ProteinMPNN."
"My RNA-seq experiment has 6 samples (3 treated, 3 control) in my workspace. Run DESeq2 and tell me which pathways are enriched in the differentially expressed genes."
"Search PubMed, UniProt, and STRING DB for everything known about the interaction between BRCA1 and RAD51 — summarize the key structural contacts."
The agent chooses tools, runs them, and streams the answer back in real time. Long-running GPU jobs (RFdiffusion, Boltz, GATK, etc.) are dispatched to the cloud and tracked in the Processes panel — you keep working while they run.
Jupyter advantage: Insert any code block the agent writes directly as a new notebook cell with one click. Attach your active cell as context so the agent knows exactly what you're working on.
Features
AI agent chat
A persistent chat panel lives in the left sidebar. It understands biological context, remembers your conversation history, and can attach files from your workspace directly.
Example workflows:
- "BLAST this sequence against NCBI nt, then fetch the top 3 hits from GenBank and align them"
- "Explain the clinical significance of the variants in
cohort.vcf— focus on BRCA1 and TP53" - "Fetch the crystal structure of human HBB from PDB, run a stability analysis on the sickle-cell mutation E6V, and visualize the result"
Type / to browse all available tools and pipelines with autocomplete. Type @ to attach a file from your workspace directly into the message.
Notebook integration
smarts.bio is built for Jupyter workflows:
- Insert as cell — when the agent writes a code block, click Insert to create a new notebook cell below your current position
- Analyze active cell — right-click any cell or use the toolbar button to send it to the agent as context
- Attach kernel variables — share the current state of your kernel with the agent so it understands your data
3D structure viewer
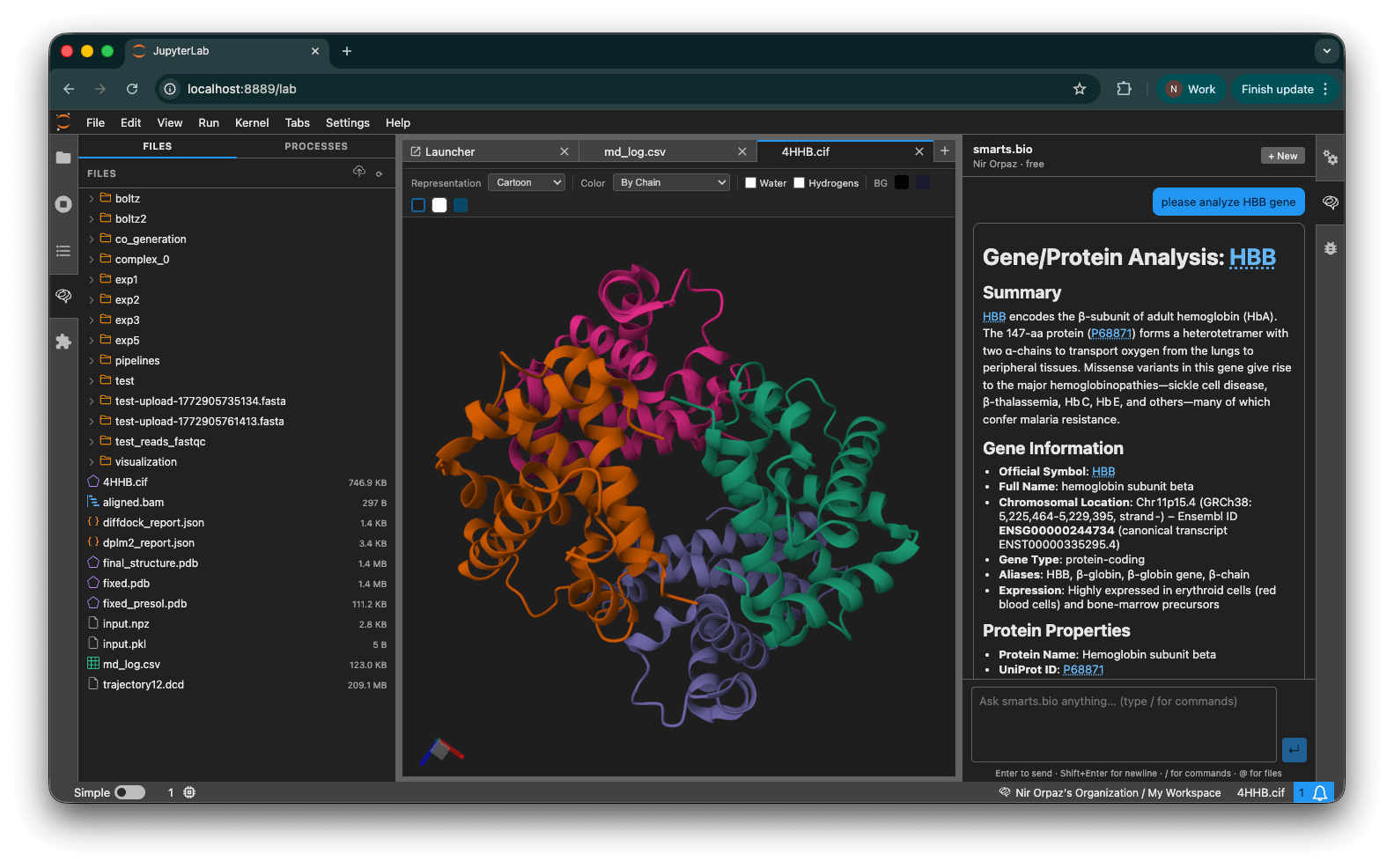
Open any .pdb or .mmcif file directly in JupyterLab. A full interactive Molstar viewer renders the structure with a toolbar for instant control.
Controls:
- Representation — Cartoon, Ball & Stick, Surface, Spacefill, Backbone
- Color — By Chain, By Element, By Residue, Secondary Structure, Uniform
- Visibility — Toggle water molecules and hydrogens
- Background — Black, dark blue, white, teal
Click Analyze with smarts.bio on any structure in the Files panel to immediately send it to the chat for AI analysis.
Sequence & genomics viewers
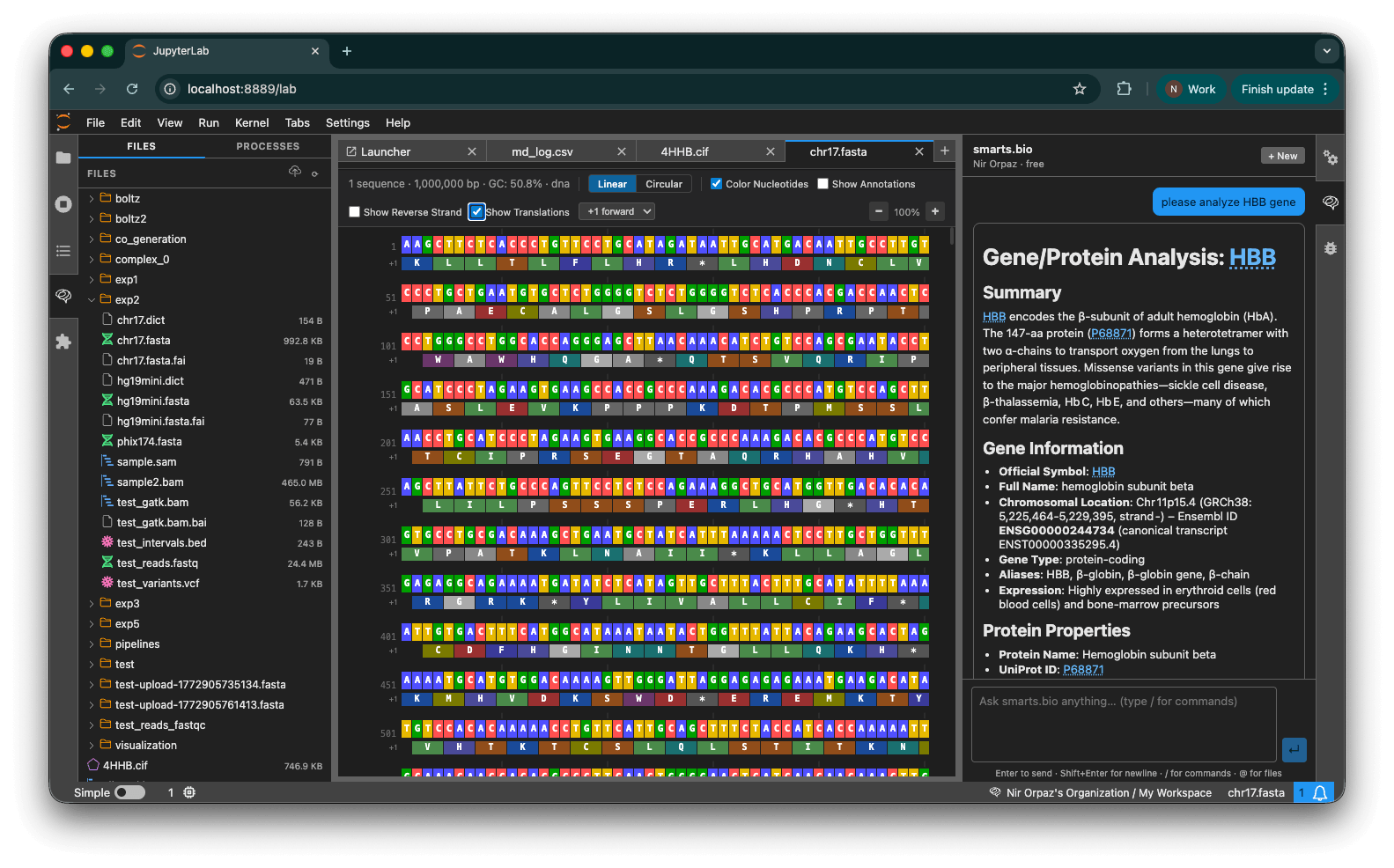
Open .fasta, .fastq, .bam, .sam, .vcf, .bed, .csv, and .tsv files natively in JupyterLab. smarts.bio registers as the default viewer for all standard bioinformatics formats, with custom icons in the JupyterLab file browser for every recognized type.
- Sequence viewer — Colorized nucleotide / amino acid display for FASTA and FASTQ, with linear and circular views, reverse strand, and translation tracks
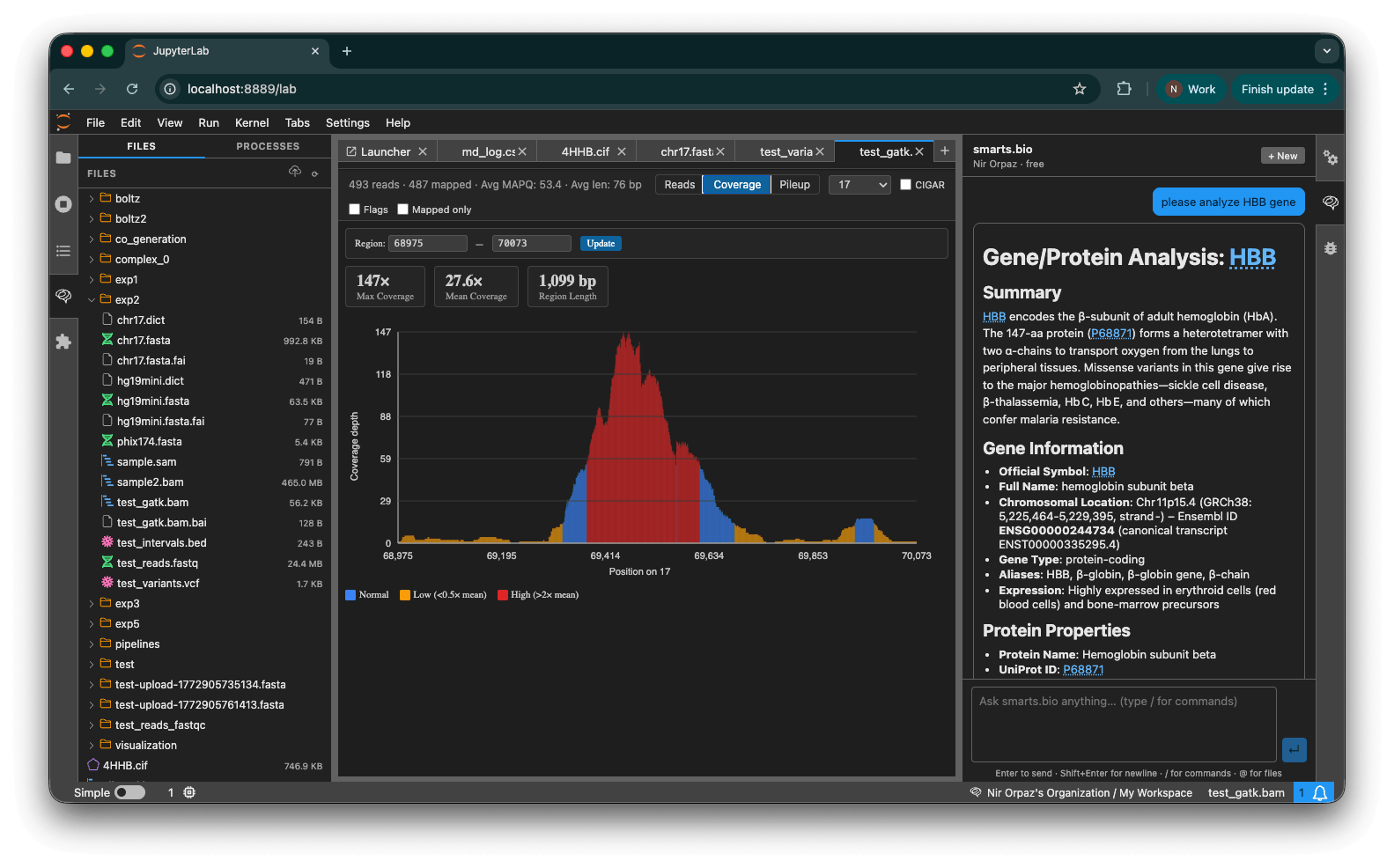
- Alignment viewer — BAM/SAM coverage histogram with per-base depth, mapped/unmapped read filtering, and region navigation
- Variant viewer — VCF/BED annotation with clinical context
- Table viewer — CSV/TSV with sortable columns — useful for reviewing DESeq2 output or BLAST hit tables
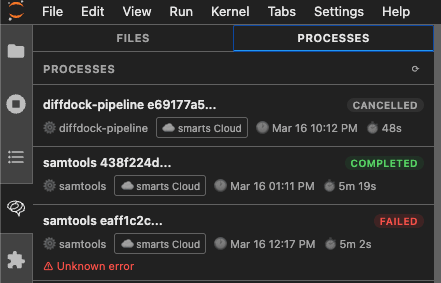
Processes panel — track long-running jobs
Submit GPU-heavy pipelines (RFdiffusion, Boltz, GATK, samtools, and more) and watch them run without blocking your notebook. The Processes panel shows live status, execution time, and error details.
- Real-time status updates (queued → running → completed / failed)
- Job duration and timestamps
- Cancel any running job directly from the panel
Files panel
A full file browser panel gives you access to your smarts.bio workspace — browse folders, upload, download, rename, move, and delete files without leaving JupyterLab.
Right-click any file to:
- Open in the built-in viewer
- Analyze with smarts.bio (sends the file as context to chat)
- Download, rename, move to a folder, or delete
Right-click any folder to:
- Upload files here
- Create a subfolder
- Rename or delete
Getting started
1. Install
pip install smartsbio-jupyterlab
Then start (or restart) JupyterLab:
jupyter lab
2. Sign in
Click the smarts.bio icon in the left sidebar → Sign In. A popup window opens, you authenticate, and JupyterLab resumes automatically.
No account? Get started at chat.smarts.bio — includes compute credits to try GPU pipelines.
3. Ask something
Use Boltz to design a nanobody targeting human PD-L1.
Use my workspace file antigen.pdb as the target.
Give me 3 candidates ranked by pLDDT score.
Requirements
- JupyterLab 4.x
- Python 3.8 or later
- A smarts.bio account (get started at chat.smarts.bio)
- Internet connection (GPU pipelines and AI run on smarts.bio cloud infrastructure)
Commands
All commands are available via the JupyterLab command palette (Ctrl+Shift+C / Cmd+Shift+C):
| Command | Description |
|---|---|
smarts-bio:open-chat |
Open the AI chat panel |
smarts-bio:new-chat |
Start a new conversation |
smarts-bio:analyze-cell |
Send the active notebook cell to chat |
smarts-bio:insert-cell-below |
Insert agent output as a new code cell |
smarts-bio:attach-kernel-context |
Share kernel variables with the agent |
smarts-bio:open-explorer |
Open the Files panel |
smarts-bio:open-processes |
Open the Processes panel |
smarts-bio:upload-file |
Upload files to your workspace |
smarts-bio:select-workspace |
Switch active workspace |
smarts-bio:sign-in |
Sign in to smarts.bio |
smarts-bio:sign-out |
Sign out |
Settings
Open Settings → Advanced Settings Editor → smarts.bio to configure:
| Setting | Default | Description |
|---|---|---|
sendOnEnter |
true |
Send message on Enter. Shift+Enter inserts a newline. |
defaultWorkspaceId |
"" |
Pin a workspace ID for this JupyterLab session. |
enableKernelContext |
true |
Allow the agent to read kernel variable names for context. |
apiBaseUrl |
https://api.smarts.bio |
API endpoint (advanced). |
Privacy
Files and text you send to the agent are transmitted to smarts.bio servers for processing. The extension shows a one-time confirmation before uploading any file. Do not send files containing patient identifiers (PHI) unless your organization has a BAA with smarts.bio.
Feedback & support
- Issues & feature requests: github.com/smartsbio/smarts-bio-jupyterlab
- Documentation: smarts.bio/docs
- Website: smarts.bio
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file smartsbio_jupyterlab-0.1.10.tar.gz.
File metadata
- Download URL: smartsbio_jupyterlab-0.1.10.tar.gz
- Upload date:
- Size: 891.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e71136a00c73185616de2c29a75f92cbf757e85740ac9b80192b4e2c3f845324
|
|
| MD5 |
f0a1dfd527d3444f1bc99308e9c07322
|
|
| BLAKE2b-256 |
f617b389f3061cc46e9a2e5a61ba252397afecd452197a086f063deca6ff98e2
|
File details
Details for the file smartsbio_jupyterlab-0.1.10-py3-none-any.whl.
File metadata
- Download URL: smartsbio_jupyterlab-0.1.10-py3-none-any.whl
- Upload date:
- Size: 773.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3acf3f8f285c3ed4b41687cfd1ac0d18fe0c154eff247f4db6f3ded5679c477a
|
|
| MD5 |
ec947213c49ac05e1e81d056acb1dbe7
|
|
| BLAKE2b-256 |
8bd3148587fdd9188b43914d6ddc38c999557b672391865445816ed07eda0c29
|