Inference of ploidy and heterozygosity structure using whole genome sequencing data
Project description
Smudgeplot
Version: 0.5.3 Skylight
Authors: Sam Ebdon, Gene W Myers and Kamil S. Jaron, Tianyi Ma.
Installation
This version of smudgeplot operates on FastK k-mer databases. The smudgeplot installation consists of a python package and C-backend to search for all the k-mer pairs (smudgeplot hetmers) and extract sequences of k-mer pairs (smudgeplot extract).
We recommend installing smudgeplot within a conda environment.
#optional conda environment setup
conda create -n smudgeplot && conda activate smudgeplot
conda install pip
# install via pypi
pip install smudgeplot
# or download and install directly. See below if you need to compile the C dependencies.
git clone https://github.com/KamilSJaron/smudgeplot.git
cd smudgeplot && pip install .
smudgeplot -h # check installation succeeded
That should do everything necesarry to make smudgeplot fully operational. You can run smudgeplot --help to see if it worked. If you activated a virtual environment prior to installation (either conda or any other) then smudgeplot installed within the environment.
Note the smudgeplot version downloadable from conda itself is not currently up to date.
Compiling the C code
The process above should install everything including compilation of the C backend. If you need or would like to know how to compile the code yourself you can simply run
make
This will not, however, install the smudgeplot python package.
Pypi installation [EXPERIMENTAL]
We are working on packaging smudgeplot for pypi. You are welcome to try installing from pypi if you are interested and please open an issue if you have problems. If it fails please follow the main instructions above to install for now.
pip install smudgeplot
Example run on Saccharomyces data
Requires ~2.1GB of space and FastK and smudgeplot installed.
# download data
wget ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR326/001/SRR3265401/SRR3265401_1.fastq.gz
wget ftp://ftp.sra.ebi.ac.uk/vol1/fastq/SRR326/001/SRR3265401/SRR3265401_2.fastq.gz
# sort them in a reasonable place
mkdir data/Scer
mv *fastq.gz data/Scer/
# run FastK to create a k-mer database
FastK -v -t4 -k31 -M16 -T4 data/Scer/SRR3265401_[12].fastq.gz -Ndata/Scer/FastK_Table
# Find all k-mer pairs in the dataset using hetmer module
smudgeplot hetmers -L 12 -t 4 -o data/Scer/kmerpairs --verbose data/Scer/FastK_Table
# this now generated `data/Scer/kmerpairs_text.smu` file;
# it's a flat file with three columns; covB, covA and freq (the number of k-mer pairs with these respective coverages)
# use the .smu file to infer ploidy and create smudgeplot
smudgeplot all -o data/Scer/trial_run data/Scer/kmerpairs.smu
# check that bunch files are generated (3 pdfs; some summary tables and logs)
ls data/Scer/trial_run_*
The y-axis scaling is by default 100, one can spcify argument ylim to scale it differently
smudgeplot all -o data/Scer/trial_run_ylim70 data/Scer/kmerpairs.smu -ylim 70
There is also a plotting module that requires the coverage, a list of smudges, and the smudge sizes listed in a tabular file. This plotting module does not perform the inference and should be used only if you know the right answers already.
How smudgeplot works
This tool extracts heterozygous kmer pairs from kmer count databases and performs gymnastics with them. We are able to disentangle genome structure by comparing the sum of kmer pair coverages (CovA + CovB) to their relative coverage (CovB / (CovA + CovB)). Such an approach also allows us to analyze obscure genomes with duplications, various ploidy levels, etc.
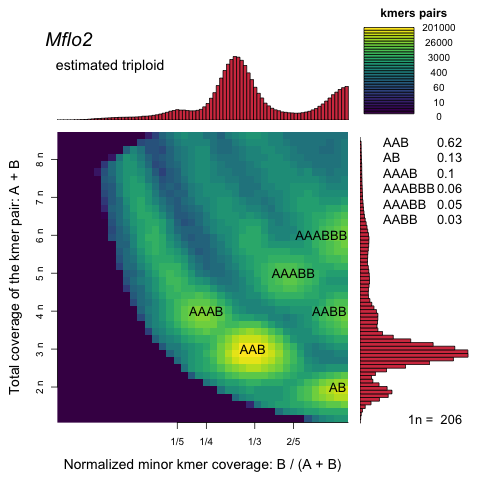
Smudgeplots are computed from raw or even better from trimmed reads and show the haplotype structure using heterozygous kmer pairs. For example (of an older version):
Every haplotype structure has a unique smudge on the graph and the heat of the smudge indicates how frequently the haplotype structure is represented in the genome compared to the other structures. The image above is an ideal case, where the sequencing coverage is sufficient to beautifully separate all the smudges, providing very strong and clear evidence of triploidy.
This tool is planned to be a part of GenomeScope in the near future.
More usage information
The input is a set of whole genome sequencing reads, the higher the coverage the better. The method is designed to process big datasets, don't hesitate to pull all single-end/pair-end libraries together.
The workflow is automatic, but it's not fool-proof. It requires some decisions. Use this tool jointly with GenomeScope.
Smudgeplot generates two plots, one with coloration on a log scale and the other on a linear scale. The legend indicates approximate kmer pairs per tile densities. Note that a single polymorphism generates multiple heterozygous kmers. As such, the reported numbers do not directly correspond to the number of variants. Instead, the actual number is approximately 1/k times the reported numbers, where k is the kmer size (in summary already recalculated). It's important to note that this process does not exhaustively attempt to find all of the heterozygous kmers from the genome. Instead, only a sufficient sample is obtained in order to identify relative genome structure. You can also report the minimal number of loci that are heterozygous if the inference is correct.
GenomeScope
You can feed the kmer coverage histogram to GenomeScope. (Either run the genomescope or use the web server)
This tool estimates the size, heterozygosity, and repetitive fraction of the genome. By inspecting the fitted model you can determine the location of the smallest peak after the error tail. Then, you can decide the low end cutoff below which all kmers will be discarded as errors (cca 0.5 times the haploid kmer coverage), and the high end cutoff above which all kmers will be discarded (cca 8.5 times the haploid kmer coverage).
Frequently Asked Questions
Are collected on our wiki. Smudgeplot does not demand much computational resources, but make sure you check memory requirements before you extract kmer pairs (hetkmers task). If you don't find an answer for your question in FAQ, open an issue or drop us an email.
Check projects to see how the development goes.
Contributions
This is definitely an open project, contributions are welcome. You can check some of the ideas for the future in projects and in the development dev branch. The file playground/DEVELOPMENT.md contains some development notes. The directory playground contains some snippets, attempts, and other items of interest.
Reference
Ranallo-Benavidez, T.R., Jaron, K.S. & Schatz, M.C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nature Communications 11, 1432 (2020). https://doi.org/10.1038/s41467-020-14998-3
Acknowledgements
This blogpost by Myles Harrison has largely inspired the visual output of smudgeplots. The colourblind friendly colour theme was suggested by @ggrimes. Grateful for helpful comments of beta testers and pre-release chatters!
## Changelog
#### 0.5
- experimental feature to extract sequences of the kmers in the pair; this functionality will hopefully restore at some point together with functionality to assess the quality of assembly.
- histograms are back
#### 0.4
- the search for the kmer pair will be within both canonical and non-canonical k-mer sets (Gene demonstrated it makes a difference)
- the tool will be supporting FastK kmer counter only
- the backend by Gene is paralelized and massively faster
- the intermediate file will be a flat file with the 2d histogram with cov1, cov2, freq columns (as opposed to list of coverages of pairs cov1 cov2);
- completelly revamped plot showing how all individual kmer pairs insead of agregating them into squares
- new smudge detection algorithm based on grid projection on the smudge plane (working, but under revisions at the moment)
- R package smudgeplot was retired and is no longer used
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file smudgeplot-0.5.3.tar.gz.
File metadata
- Download URL: smudgeplot-0.5.3.tar.gz
- Upload date:
- Size: 59.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f551a812cca280e4cd5341fe4167aa50039d5dae7ae64ca5d00c41c0fa0aca3b
|
|
| MD5 |
34b5f4db1198e9e27bfdf278f21fd187
|
|
| BLAKE2b-256 |
ff08ece11dae68c4f069fcc5f109a2aaba5c2945aabea51ae589e14e5e47452c
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3.tar.gz:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3.tar.gz -
Subject digest:
f551a812cca280e4cd5341fe4167aa50039d5dae7ae64ca5d00c41c0fa0aca3b - Sigstore transparency entry: 757587157
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp312-cp312-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp312-cp312-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 121.7 kB
- Tags: CPython 3.12, manylinux: glibc 2.12+ x86-64, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
425d756828117fc0b5684982f0b8ed761c2697d76989d3615858a1e9e7b457da
|
|
| MD5 |
08fc97e702e5e6139d29839f521a6b19
|
|
| BLAKE2b-256 |
9be1f28e3e8b33f4f23b209c0fc1fa40fd43de0ff700bba8abb121ee236a7d76
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp312-cp312-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp312-cp312-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl -
Subject digest:
425d756828117fc0b5684982f0b8ed761c2697d76989d3615858a1e9e7b457da - Sigstore transparency entry: 757587209
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp312-cp312-macosx_11_0_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp312-cp312-macosx_11_0_x86_64.whl
- Upload date:
- Size: 126.0 kB
- Tags: CPython 3.12, macOS 11.0+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3a8679d95e55ba58b1c1b366b61136a02007e94b5363c778f36f67e99142b86e
|
|
| MD5 |
aca5a7d639e1ad25591d266db07a8236
|
|
| BLAKE2b-256 |
9944ba942ca19acb4262100ea53f28da9ac38318ca9b14696acd8c41a9ecf2ee
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp312-cp312-macosx_11_0_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp312-cp312-macosx_11_0_x86_64.whl -
Subject digest:
3a8679d95e55ba58b1c1b366b61136a02007e94b5363c778f36f67e99142b86e - Sigstore transparency entry: 757587172
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp312-cp312-macosx_11_0_arm64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp312-cp312-macosx_11_0_arm64.whl
- Upload date:
- Size: 120.5 kB
- Tags: CPython 3.12, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ed453bcd0df270d78517a298973afec0f91e3b8434ecfbb3a721eba69ac60b89
|
|
| MD5 |
4ef151c3f67777c971a5c46498aee354
|
|
| BLAKE2b-256 |
1e0987d1f1e13f168fe6cf977d804659425d6e9c70b71e4e0a4f26dd833f652a
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp312-cp312-macosx_11_0_arm64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp312-cp312-macosx_11_0_arm64.whl -
Subject digest:
ed453bcd0df270d78517a298973afec0f91e3b8434ecfbb3a721eba69ac60b89 - Sigstore transparency entry: 757587191
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp311-cp311-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp311-cp311-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 121.7 kB
- Tags: CPython 3.11, manylinux: glibc 2.12+ x86-64, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
850edfb34ee8548b9b2bf9d0f7acc63a58c9e7193c2eee64e18c05ddeb957d41
|
|
| MD5 |
f602ad891fdd9c0e2c3d28b26183edc9
|
|
| BLAKE2b-256 |
a8dbf72d4f9fce55c7d508daf15ef4fe66141bf005122d54dc44e9e57d461b8e
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp311-cp311-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp311-cp311-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl -
Subject digest:
850edfb34ee8548b9b2bf9d0f7acc63a58c9e7193c2eee64e18c05ddeb957d41 - Sigstore transparency entry: 757587199
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp311-cp311-macosx_11_0_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp311-cp311-macosx_11_0_x86_64.whl
- Upload date:
- Size: 126.0 kB
- Tags: CPython 3.11, macOS 11.0+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e7a95444dd99afea9f7882e362305b894b001ebd8ddb76dcacb9a4a20b86456e
|
|
| MD5 |
db2ae25b48dbf29524e0961c4218f7e9
|
|
| BLAKE2b-256 |
d0b438505d1621a44f3674319b22b500740d58a1a4331273487cf1322c220e70
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp311-cp311-macosx_11_0_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp311-cp311-macosx_11_0_x86_64.whl -
Subject digest:
e7a95444dd99afea9f7882e362305b894b001ebd8ddb76dcacb9a4a20b86456e - Sigstore transparency entry: 757587182
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp311-cp311-macosx_11_0_arm64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp311-cp311-macosx_11_0_arm64.whl
- Upload date:
- Size: 120.5 kB
- Tags: CPython 3.11, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f11de8649e911e3dee2d6afb5924f3b4d8c775b8374b505b57cbd268b9c95843
|
|
| MD5 |
97bc90647c86648fd18bc680ba776572
|
|
| BLAKE2b-256 |
33c63802059334376bf6c0fc92aef8b272a921b258c19e270fa5b06dd3f8c51b
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp311-cp311-macosx_11_0_arm64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp311-cp311-macosx_11_0_arm64.whl -
Subject digest:
f11de8649e911e3dee2d6afb5924f3b4d8c775b8374b505b57cbd268b9c95843 - Sigstore transparency entry: 757587221
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp310-cp310-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp310-cp310-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 121.7 kB
- Tags: CPython 3.10, manylinux: glibc 2.12+ x86-64, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3080345cd207e1250ce015acd388213227ec5d25dc71b0d273c61d7e1175eac7
|
|
| MD5 |
027774bef7fcd2a8c1cc2ab6937e3d55
|
|
| BLAKE2b-256 |
8e5963eaf7f6856d962e90332b5aea04a7f39b0da00d2e1a806d22ff6014773f
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp310-cp310-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp310-cp310-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl -
Subject digest:
3080345cd207e1250ce015acd388213227ec5d25dc71b0d273c61d7e1175eac7 - Sigstore transparency entry: 757587243
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp310-cp310-macosx_11_0_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp310-cp310-macosx_11_0_x86_64.whl
- Upload date:
- Size: 126.0 kB
- Tags: CPython 3.10, macOS 11.0+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
971f91ae95548bbdb6bad6b109b8aecdfb1d21473f3603d33630687c8ae16569
|
|
| MD5 |
0efe6dfe6a0678892779f462371ba6c5
|
|
| BLAKE2b-256 |
21470bfbb5cab5e27fcd98b852fb7407e548cf3d25b5b1e3e744cd78cedf3d42
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp310-cp310-macosx_11_0_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp310-cp310-macosx_11_0_x86_64.whl -
Subject digest:
971f91ae95548bbdb6bad6b109b8aecdfb1d21473f3603d33630687c8ae16569 - Sigstore transparency entry: 757587241
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp310-cp310-macosx_11_0_arm64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp310-cp310-macosx_11_0_arm64.whl
- Upload date:
- Size: 120.5 kB
- Tags: CPython 3.10, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dccb1e7817a495d0f8de1f30c17e701c623bf50a976d1e5360b05240bdcdd79b
|
|
| MD5 |
889ea206369d827419a7ad1d47915bc8
|
|
| BLAKE2b-256 |
cbdbe6d5e2111909a6403e4985df7527a8a3b1d6a43fc838a7fad56f6bcb7349
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp310-cp310-macosx_11_0_arm64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp310-cp310-macosx_11_0_arm64.whl -
Subject digest:
dccb1e7817a495d0f8de1f30c17e701c623bf50a976d1e5360b05240bdcdd79b - Sigstore transparency entry: 757587217
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp39-cp39-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp39-cp39-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 121.7 kB
- Tags: CPython 3.9, manylinux: glibc 2.12+ x86-64, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
56a1c7ac1d8270c2de26c032bf261feb8d7c0882c8c2c509829250689f5a6911
|
|
| MD5 |
56853e6d18c869d4bfe8f666eb7b89de
|
|
| BLAKE2b-256 |
a9e619ee2fd84fdcad5f8a07969f8dfabcdc13130d0457381b1b06cc25fea758
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp39-cp39-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp39-cp39-manylinux_2_12_x86_64.manylinux2010_x86_64.manylinux_2_17_x86_64.manylinux2014_x86_64.whl -
Subject digest:
56a1c7ac1d8270c2de26c032bf261feb8d7c0882c8c2c509829250689f5a6911 - Sigstore transparency entry: 757587228
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp39-cp39-macosx_11_0_x86_64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp39-cp39-macosx_11_0_x86_64.whl
- Upload date:
- Size: 126.0 kB
- Tags: CPython 3.9, macOS 11.0+ x86-64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fc2370cb98d1b49df28c9ff23716d2ff2a4254b0a4cc0118f3f5e81c79d5da72
|
|
| MD5 |
63529c81605d4517f097d13925daacd8
|
|
| BLAKE2b-256 |
1805042cae187d133b08596d62f2d7791a0f2152f5c8c9eb38cdc83db0b5a1b7
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp39-cp39-macosx_11_0_x86_64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp39-cp39-macosx_11_0_x86_64.whl -
Subject digest:
fc2370cb98d1b49df28c9ff23716d2ff2a4254b0a4cc0118f3f5e81c79d5da72 - Sigstore transparency entry: 757587165
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type:
File details
Details for the file smudgeplot-0.5.3-cp39-cp39-macosx_11_0_arm64.whl.
File metadata
- Download URL: smudgeplot-0.5.3-cp39-cp39-macosx_11_0_arm64.whl
- Upload date:
- Size: 120.5 kB
- Tags: CPython 3.9, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bdef93b7f2ee368436944f2bba0b0824f1515ac187a9da7a99f9a50f4895e64a
|
|
| MD5 |
c0e7922d7720560f0a2824a0bd3ebd8c
|
|
| BLAKE2b-256 |
6c0d58997e3bc65a9b7e9a458392159412699f9f56fa3bc033fca5eae2cb86b3
|
Provenance
The following attestation bundles were made for smudgeplot-0.5.3-cp39-cp39-macosx_11_0_arm64.whl:
Publisher:
publish.yml on KamilSJaron/smudgeplot
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
smudgeplot-0.5.3-cp39-cp39-macosx_11_0_arm64.whl -
Subject digest:
bdef93b7f2ee368436944f2bba0b0824f1515ac187a9da7a99f9a50f4895e64a - Sigstore transparency entry: 757587236
- Sigstore integration time:
-
Permalink:
KamilSJaron/smudgeplot@9049082c20363a5a464a88202193ddb04a28d5eb -
Branch / Tag:
refs/tags/v0.5.3 - Owner: https://github.com/KamilSJaron
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@9049082c20363a5a464a88202193ddb04a28d5eb -
Trigger Event:
release
-
Statement type: