Spatial phenotype analysis of crisp screens (SpaCr)
Project description
Badges
SpaCr
Spatial phenotype analysis of CRISPR-Cas9 screens (SpaCr).
The spatial organization of organelles and proteins within cells constitutes a key level of functional regulation. In the context of infectious disease, the spatial relationships between host cell structures and intracellular pathogens are critical to understanding host clearance mechanisms and how pathogens evade them. SpaCr is a Python-based software package for generating single-cell image data for deep-learning sub-cellular/cellular phenotypic classification from pooled genetic CRISPR-Cas9 screens. SpaCr provides a flexible toolset to extract single-cell images and measurements from high-content cell painting experiments, train deep-learning models to classify cellular/subcellular phenotypes, simulate, and analyze pooled CRISPR-Cas9 imaging screens.
Features
Generate Masks: Generate cellpose masks of cell, nuclei, and pathogen objects.
Object Measurements: Measurements for each object including scikit-image regionprops, intensity percentiles, shannon-entropy, Pearson’s and Manders’ correlations, homogeneity, and radial distribution. Measurements are saved to a SQL database in object-level tables.
Crop Images: Save objects (cells, nuclei, pathogen, cytoplasm) as images. Object image paths are saved in a SQL database.
Train CNNs or Transformers: Train Torch models to classify single object images.
Manual Annotation: Supports manual annotation of single-cell images and segmentation to refine training datasets for training CNNs/Transformers or cellpose, respectively.
Finetune Cellpose Models: Adjust pre-existing Cellpose models to your specific dataset for improved performance.
Timelapse Data Support: Track objects in timelapse image data.
Simulations: Simulate spatial phenotype screens.
Sequencing: Map FASTQ reads to barcode and gRNA barcode metadata.
Misc: Analyze Ca oscillation, recruitment, infection rate, plaque size/count.
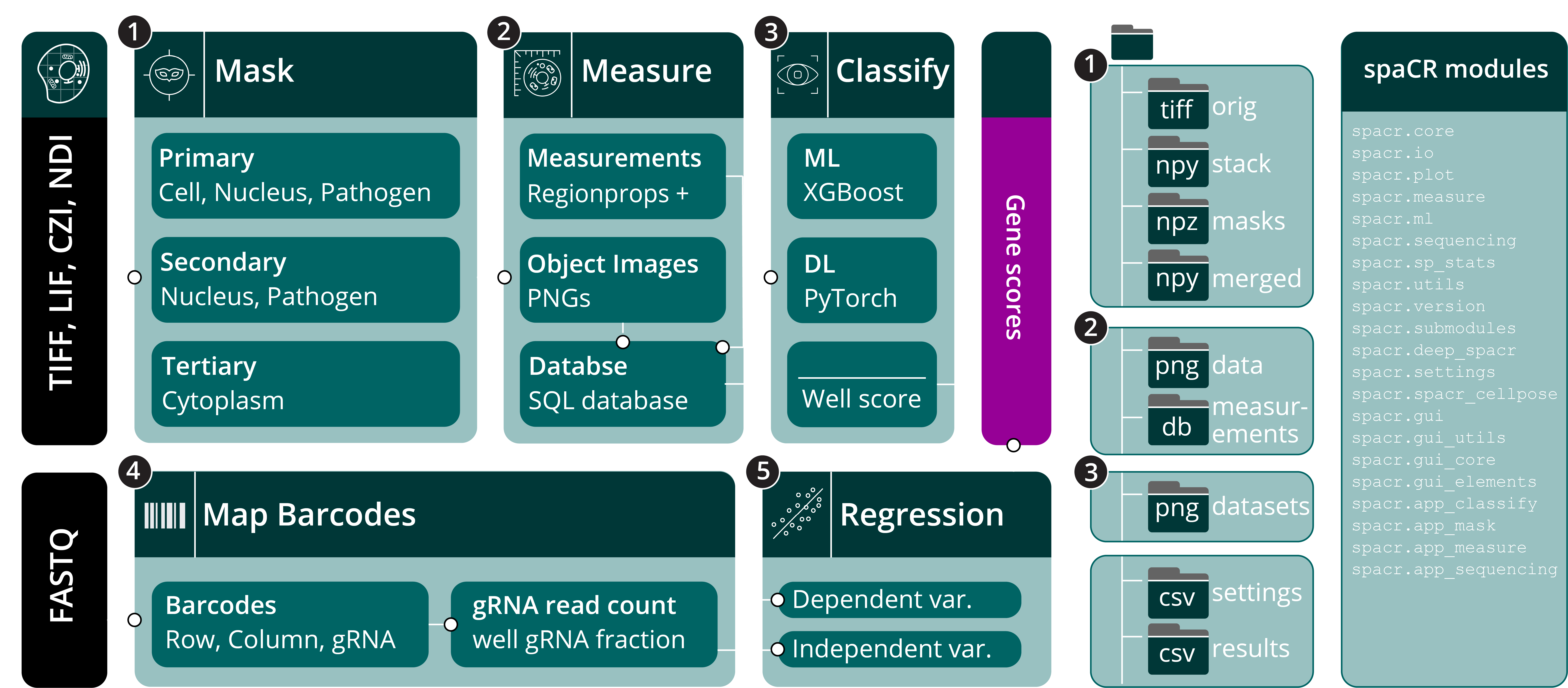
Overview and data organization of spaCR.
a. Schematic workflow of the spaCR pipeline for pooled image-based CRISPR screens. Microscopy images (TIFF, LIF, CZI, NDI) and sequencing reads (FASTQ) are used as inputs (black). The main modules (teal) are: (1) Mask: generates object masks for cells, nuclei, pathogens, and cytoplasm; (2) Measure: extracts object-level features and crops object images, storing quantitative data in an SQL database; (3) Classify—applies machine learning (ML, e.g., XGBoost) or deep learning (DL, e.g., PyTorch) models to classify objects, summarizing results as well-level classification scores; (4) Map Barcodes: extracts and maps row, column, and gRNA barcodes from sequencing data to corresponding wells; (5) Regression: estimates gRNA effect sizes and gene scores via multiple linear regression using well-level summary statistics. b. Downstream submodules available for extended analyses at each stage. c. Output folder structure for each module, including locations for raw and processed images, masks, object-level measurements, datasets, and results. d. List of all spaCR package modules.
Installation
Linux recommended. If using Windows, switch to Linux—it’s free, open-source, and better.
macOS prerequisites (before install):
brew install libomp hdf5 cmake openssl
Linux GUI requirement: SpaCr GUI requires Tkinter.
sudo apt-get install python3-tk
Installation:
pip install spacr
Run SpaCr GUI:
spacr
Example Notebooks
The following example Jupyter notebooks illustrate common workflows using spaCR.
Generate masks Generate cell, nuclei, and pathogen segmentation masks from microscopy images using Cellpose.
Capture single cell images and measurements Extract object-level measurements and crop single-cell images for downstream analysis.
Machine learning based object classification Train traditional machine learning models (e.g., XGBoost) to classify cell phenotypes based on extracted features.
Computer vision based object classification Train and evaluate deep learning models (PyTorch CNNs/Transformers) on cropped object images.
Map sequencing barcodes Map sequencing reads to row, column, and gRNA barcodes for CRISPR screen genotype-phenotype mapping.
Finetune cellpose models Finetune Cellpose models using your own annotated training data for improved segmentation accuracy.
Interactive Tutorial (under construction)
Click below to explore the step-by-step GUI and Notebook tutorials for spaCR:
spaCRPower
Power analasys of pooled perterbation spaCR screens.
Data Availability
Full microscopy image dataset: EMBL-EBI BioStudies S-BIAD2135
Testing dataset: Hugging Face toxo_mito
Sequencing data: NCBI BioProject PRJNA1261935
License
spaCR is distributed under the terms of the MIT License. See the LICENSE file for details.
How to Cite
If you use spaCR in your research, please cite: Olafsson EB, et al. SpaCr: Spatial phenotype analysis of CRISPR-Cas9 screens. Manuscript in preparation.
Papers Using spaCR
Below are selected publications that have used or cited spaCR:
Olafsson EB, et al. SpaCr: Spatial phenotype analysis of CRISPR-Cas9 screens. Manuscript in preparation.
IRE1α promotes phagosomal calcium flux to enhance macrophage fungicidal activity
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file spacr-1.3.4.tar.gz.
File metadata
- Download URL: spacr-1.3.4.tar.gz
- Upload date:
- Size: 64.4 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6e885ec8150c1941f196cb3bbe15b6edc4af99a96158817755e2d7196e5fba5e
|
|
| MD5 |
9babfa3ff6b908f9fc41c7e4f52651f7
|
|
| BLAKE2b-256 |
dce4a6978ceeb4dddf2d53f8eac361e03bbddc2a5836af793b8b017fe6f81029
|
File details
Details for the file spacr-1.3.4-py3-none-any.whl.
File metadata
- Download URL: spacr-1.3.4-py3-none-any.whl
- Upload date:
- Size: 28.3 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
715f24c4499dba92b5ace5a7ef2d0b9372e3da6942853fb0893ba58c775fd36f
|
|
| MD5 |
dbe061095d1593a7e3f3fb2521568e67
|
|
| BLAKE2b-256 |
4268fa3748740be9bb22a9e1d5779f74386fd51d0dc9dedfecfff733e510ae65
|

















