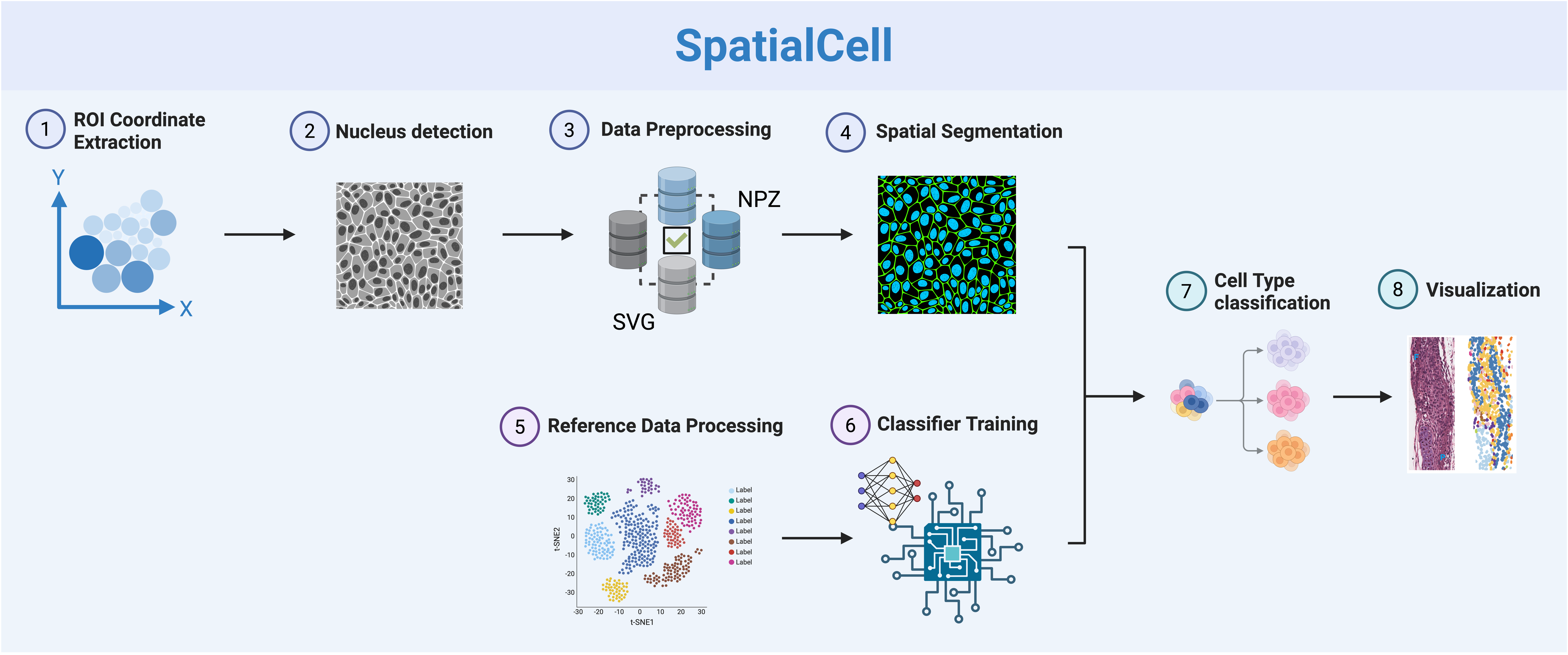
Integrated pipeline for spatial transcriptomics cell segmentation and annotation using QuPath, Bin2cell, and TopAct
Project description
SpatialCell: Integrated Spatial Transcriptomics Analysis Pipeline
SpatialCell is an integrated computational pipeline for spatial transcriptomics analysis that combines cell segmentation and automated cell type annotation. It seamlessly integrates Stardist (applied as QuPath plugin for cell detection) for histological image analysis, Bin2cell for spatial cell segmentation, and TopAct for machine learning-based cell classification.
🚀 Key Features
- Multi-scale Cell Segmentation: Stardist-enabled QuPath cell detection with Bin2cell spatial segmentation
- Automated Cell Annotation: TopAct-based machine learning classification
- ROI-aware Processing: Region-of-interest focused analysis for large datasets
- Scalable Pipeline: Support for multiple developmental time points (e.g., E14.5, E18.5, P3) and samples
- Visualization Tools: Comprehensive plotting and export capabilities
- Modular Design: Easy to customize and extend for specific research needs
🔧 Installation
Prerequisites
- Python 3.10 or higher
- QuPath (for histological image analysis)
- Git
- Operating Systems tested: Ubuntu 22.04.03, MacOS 15.5
- Hardware: Standard desktop CPU; GPU not required but optional for accelerated image processing
- Additional Python dependencies are listed in
requirements.txt
Typical installation time
Installation usually completes within 5 minutes on a stable internet connection and a typical desktop computer.
Quick Install (Recommended)
To enable full functionality including TopAct classification, please install TopAct separately:
pip install spatialcell
pip install git+https://gitlab.com/kfbenjamin/topact.git
Alternative: Install from Source
# Clone the repository
git clone https://github.com/Xinyan-C/Spatialcell.git
cd Spatialcell
# Install dependencies
pip install -r requirements.txt
# Install the package in editable mode
pip install -e .
📋 Demo Data and tutorial notebook
The examples/ directory contains the tutorial notebook to quickly test and understand SpatialCell.
Demo datasets for E14.5, E18.5, and P3 are archived on Zenodo (https://zenodo.org/records/16400171)
Expected output
-
ROI coordinates saved as a
.txtfile- e.g.
examples/demo_data/E18.5_ranges.txt
- e.g.
-
Binary segmentation masks saved as
.npzfiles- e.g.
examples/demo_data/E18.5_qupath.npz
- e.g.
-
Spatial segmentation results under
examples/demo_data/demo_output/(more information at https://github.com/Teichlab/bin2cell.git):- Data/
E18.5_2um.h5ad— AnnData containing 2 μm‐bin counts and coordinates for the entire sampleE18.5_b2c.h5ad— Bin2cell‐reconstructed cell‐level AnnData for the entire sample
- ROI_Data/ (one subfolder per ROI: CS1, CS2, WT1)
{ROI}_adata.h5ad— Spot‐level AnnData extracted for that specific region{ROI}_cdata.h5ad— Cell‐level AnnData (Bin2cell output) for that region
- destripe/, expanded_labels/, gex_labels/, joint_labels/, joint_labels_all/, npz_labels/, render_gex/, render_labels/, segmentation/
- PDF reports (quality‐control and visualization overlays) for each processing step
- Log file
spatial_processing.log— Records parameters (e.g.prob_thresh,nms_thresh), runtime info, and warnings
- Data/
-
Cell annotation outputs under
examples/demo_data/demo_output/cell_annotation/:outfile_<sample>_<sample>_-_<ROI>.npy- NumPy arrays of per-cell feature matrices (e.g. classification probabilities or aggregated counts) for each ROI (CS1, CS2, WT1)
sd_<sample>_<sample>_-_<ROI>.joblib- Serialized TopACT classifier models saved after training or calibration on each ROI
spatial_data_<sample>_roi.joblib- Serialized AnnData object containing spatially indexed spot‐level and cell‐level data passed into TopACT for classification
-
Visualization outputs under
examples/demo_data/demo_output/visualizations/:
For each ROI (CS1, CS2, WT1):Spatial_Classification_<sample>_-_<ROI>_overlay.pdf- Cell type predictions overlaid directly on the high‐resolution tissue image
Spatial_Classification_<sample>_-_<ROI>_side_by_side.pdf- Side-by-side panels showing (left) raw segmentation mask and (right) classification overlay for comparison
Spatial_Classification_<sample>_-_<ROI>.pdf- High-resolution, publication-ready map of predicted cell types (colored segmentation)
Runtime estimate
Approximately 30-45 minutes on a standard desktop for the demo dataset.
📖 Usage Instructions
The easiest way to accomplish the pipeline is with our Jupyter notebook tutorial, the tutorial covers the complete workflow from ROI extraction to visualization.
🗂️ Project Structure
Spatialcell/
├── spatialcell/ # Main package
│ ├── qupath_scripts/ # QuPath-Stardist integration scripts
│ ├── preprocessing/ # Data preprocessing modules
│ ├── spatial_segmentation/ # Bin2cell integration
│ ├── cell_annotation/ # TopAct classification
│ └── utils/ # Utility functions
├── examples/ # Tutorial notebook
│ └── SpatialCell_Demo.ipynb # Jupyter notebooks for tutorial and article reproducibility
├── requirements.txt # Python dependencies
├── setup.py # Package installation script
└── README.md # This file
🔬 Workflow Overview
- ROI Coordinate Extraction: Extract region-of-interest coordinates from Loupe Browser exports
- Nucleus detection: StarDist-based nucleus detection via QuPath with SVG export
- Data Preprocessing: SVG to NPZ conversion and label mask generation
- Spatial Segmentation: Bin2cell integration with nucleus boundaries and label expansion
- Reference Data Processing: Extract training data from Seurat RDS files
- Classifier Training: Train TopAct machine learning models for cell type annotation
- Cell Type Classification: Apply TopAct classifiers for spatial cell type prediction
- Comprehensive Visualization: Multi-scale plotting, overlay generation, and result export
📝 License
SpatialCell is licensed under the Apache License 2.0, which includes patent protection and allows commercial use.
Dependency Licenses:
- bin2cell: MIT License (automatically installed)
- TopAct: GPL v3 License (optional, user installs separately)
Note: Users should be aware of GPL license requirements when installing TopAct.
For full license text, see the LICENSE file.
📚 Article reproducibility
Jupyter notebooks (e.g. examples/SpatialCell_Demo.ipynb) needed to reproduce our analyses in the article Spatiotemporal Single-Cell Atlas of Suture Stem Cell Dynamics in Craniosynostosis are included in the examples/ directory. A minimal example dataset for E14.5, E18.5, and P3 is archived on Zenodo (https://zenodo.org/records/16400171).
📄 Citation
If you use SpatialCell in your research, please cite:
@software{spatialcell2025,
author = {Xinyan},
title = {SpatialCell: Integrated Spatial Transcriptomics Analysis Pipeline},
url = {https://github.com/Xinyan-C/Spatialcell},
year = {2025}
}
📧 Contact
- Author: Xinyan
- Email: keepandon@gmail.com
- GitHub: @Xinyan-C
🔗 References
- QuPath: Bankhead P, Loughrey MB, Fernández JA, et al. QuPath: Open source software for digital pathology image analysis. Sci Rep. 2017;7(1):16878. doi:10.1038/s41598-017-17204-5
- Stardist: Schmidt U, Weigert M, Broaddus C, Myers G. Cell detection with star-convex polygons. MICCAI 2018: 265-273. doi:10.1007/978-3-030-00934-2_30
- Bin2cell: Polański K, Bartolomé-Casado R, Sarropoulos I, et al. Bin2cell reconstructs cells from high resolution visium HD data. Bioinformatics. 2024;40(9):btae546. doi:10.1093/bioinformatics/btae546
- TopAct: Benjamin K, Bhandari A, Kepple JD, et al. Multiscale topology classifies cells in subcellular spatial transcriptomics. Nature. 2024;630(8018):943-949. doi:10.1038/s41586-024-07563-1
- Scanpy: Wolf FA, Angerer P, Theis FJ. SCANPY: large-scale single-cell gene expression data analysis. Genome Biology. 2018;19(1):15. doi:10.1186/s13059-017-1382-0
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file spatialcell-1.1.1.tar.gz.
File metadata
- Download URL: spatialcell-1.1.1.tar.gz
- Upload date:
- Size: 74.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2093f599fd956db23b7f3f29879a37f35c03b335e3babf171bba9ec66010b749
|
|
| MD5 |
07402db6d72935e2f3165668e9a6479b
|
|
| BLAKE2b-256 |
13e34e2f22d5519552df9dd68ebbcc39b75fced3faf650b35f5e3302925fb0d0
|
File details
Details for the file spatialcell-1.1.1-py3-none-any.whl.
File metadata
- Download URL: spatialcell-1.1.1-py3-none-any.whl
- Upload date:
- Size: 76.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
882994973478fa68069544635e5b1f33cb28dbb68e93ccda327182744753299d
|
|
| MD5 |
88e80bb74b8582bc2155245a43f2e8a9
|
|
| BLAKE2b-256 |
1c7072f41a7310fc3eada9c405386d002fa780d8e7d0046b4ed9a6630e986252
|