BioTuring SpatialX Connector
Project description
BioTuring SpatialX Connector
Version Update: April 2025
This update introduces several enhancements to data management and analysis capabilities:
-
Enhanced Cell Filtering: Users can now refine cell selection by adjusting filtering parameters based on both the number of genes and transcripts. Check 1.7.5. Parsing Submission Information for Multiple Samples
-
Modular Study Creation: Studies can be created and data elements submitted independently, providing greater flexibility in data organization. [Check 1.7.2. Submitting a New Study with a Single Sample and Data and Check 1.7.6. Data Details and Element Management]
-
Improved Study Information Access: Enhanced features for accessing and reviewing comprehensive study information. Check 1.7.6. Data Details and Element Management
-
Metadata and Embeddings Upload Functionality: Users can now upload new metadata files, enriching their datasets with additional contextual information. Check 1.7.6. Data Details and Element Management
-
Flexible Data Structure Customization: Users gain increased control over the data structure of submissions, allowing for tailored data organization. Check 1.7.7. Data Details and Element Management
-
Expanded Visualization Options: The analysis section now includes a wider range of visualization tools for improved data exploration. Check 1.8.5. Interact with elements
IMPORTANT: To utilize all new functionalities, please re-execute the installation step: 1.1. Installation SpatialX Connector.
1.1. Installation SpatialX Connector
!pip uninstall spatialx_connector -y -q
!pip install spatialx_connector -U -q
1.2. Import the related packages
import warnings
warnings.filterwarnings("ignore")
import os
import pandas as pd
from matplotlib import pyplot as plt
import spatialx_connector
from spatialx_connector import SpatialXConnector
from spatialx_connector import Technologies
from spatialx_connector import DefaultGroup
from spatialx_connector import Species
from spatialx_connector import SpatialAttrs
from spatialx_connector import ConnectorKeys
from spatialx_connector import SubmissionElementKeys
from spatialx_connector import ImagesSubmission
from spatialx_connector import SegmentationSubmission
from spatialx_connector import ExpressionSubmission
1.3. Domain and Token
To obtain your domain URL and personal token, navigate to "SpatialX SDK" in the left panel of your SpatialX interface. Then, enter the information in the fields below. For example:
DOMAIN = "https://example.bioturing.com/spatialx/"
TOKEN = "000000000000000000000000NM"
DOMAIN = ""
TOKEN = ""
1.4. Explore Your Account
With your domain and token added, you can now connect to your SpatialX account and workspace and explore your account details.
1.4.1. User's Information:
connector = SpatialXConnector(DOMAIN, TOKEN)
spatialx_connector.format_print(connector.info)
{
email: techsupport@bioturing.com
sub_dir: 649e75177eb042a6638f319e35e4959f
name: ub-techsu-332f83917f3b7d3
app_base_url: ...
routing_table: {
services: {}
}
}
1.4.2. Groups:
spatialx_connector.format_print(connector.groups)
{
Personal workspace: 40d094152b98ba25151936c18becb267
All members: GLOBAL_GROUP
My submitted data: MY_SUBMITTED_GROUP
}
1.5. List of Storages
If you have configured your cloud storages or would like to check your data list in SpatialX, the list of functions below can help you to get the information:
1.5.1. AWS buckets
spatialx_connector.format_print(connector.s3)
{
bioturingpublic: <USER_PATH>/public_cloud/bioturingpublic
}
1.5.2. Personal and Shared Folders
spatialx_connector.format_print(connector.folders)
{
Upload: <USER_PATH>/upload
Submitted: <USER_PATH>/study
Converted: <USER_PATH>/converted
}
1.5.3. Browsing Storage
connector.listdir(connector.s3["bioturingpublic"])
['SpatialX_datasets', 'mount']
connector.listdir(os.path.join(connector.s3["bioturingpublic"], "SpatialX_datasets"))
['COSMX_VER1', 'Human_Colon_Cancer_P2']
connector.listdir(os.path.join(connector.s3["bioturingpublic"], "SpatialX_datasets/COSMX_VER1"))
['Lung6', 'Lung9_Rep1', 'Lung9_Rep2']
1.6. Uploading files
To upload files to your personal folders within your SpatialX account, execute the code below. Be sure to replace the placeholder file_path with the complete path to the file you wish to upload.
1.6.1. Create data for testing upload:
os.makedirs("./upload_folder", exist_ok=True)
with open("./upload_folder/upload_file", "w") as f:
f.write("This is file for upload")
with open("./upload_folder/upload_big_file", "w") as f:
f.write("This is big file for upload")
1.6.2. Upload files and folders:
uploading_results = connector.upload_file(file_path="./upload_folder/upload_file")
spatialx_connector.format_print(uploading_results)
{
folder_name:
unique_id: 01JRVQJQZMGJA377FPFZHZZZSC
path: <USER_PATH>/upload/upload_file
url: <USER_URL>/upload/upload_file
}
uploading_results = connector.upload_big_file(file_path="./upload_folder/upload_big_file", debug_mode=True)
spatialx_connector.format_print(uploading_results)
{
folder_name:
unique_id: 01JRVQJV5BKGF3MDSBY3DH57VH
path: <USER_PATH>/upload/upload_big_file
url: <USER_URL>/upload/upload_big_file
}
uploading_results = connector.upload_folder(dir_path="./upload_folder", debug_mode=True)
spatialx_connector.format_print(uploading_results)
<USER_PATH>/upload/upload_folder.v0
1.7. Submission
1.7.1. Parsing Data Information for Submission:
data_name: Name of the dataset.technology: Technology used for the dataset.data_path: Path to the dataset.
Visium_V2_Human_Colon_Cancer_P2_submission_information = connector.parse_data_information(
data_name="Visium_V2_Human_Colon_Cancer_P2",
technology=Technologies.VISIUM.value,
data_path=os.path.join(
connector.s3["bioturingpublic"],
"SpatialX_datasets/Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2"
)
)
spatialx_connector.format_print(Visium_V2_Human_Colon_Cancer_P2_submission_information)
[
{
name: Visium_V2_Human_Colon_Cancer_P2
submission_type: SUBMIT_SPATIAL_BULK
technology: VISIUM
files: [
{
key: images
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2_tissue_image.btf
}
{
key: matrix
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2_raw_feature_bc_matrix.h5
}
{
key: tissue_positions
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2/spatial/tissue_positions.csv
}
{
key: scalefactors
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Visium_V2_Human_Colon_Cancer_P2/spatial/scalefactors_json.json
}
]
folders: []
args: []
kwargs: [
{
key: min_genes
value: 1
}
{
key: min_counts
value: 1
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
]
Xenium_V1_Human_Colon_Cancer_P2_submission_information = connector.parse_data_information(
data_name="Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE",
technology=Technologies.XENIUM.value,
data_path=os.path.join(
connector.s3["bioturingpublic"],
"SpatialX_datasets/Human_Colon_Cancer_P2/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE"
)
)
spatialx_connector.format_print(Xenium_V1_Human_Colon_Cancer_P2_submission_information)
[
{
name: Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE
submission_type: SUBMIT_SPATIAL_TRANSCRIPTOMICS
technology: XENIUM
files: [
{
key: experiment
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE/experiment.xenium
}
{
key: images
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE/morphology.ome.tif
}
{
key: alignment
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE_he_imagealignment.csv
}
{
key: segmentation
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE/cell_boundaries.parquet
}
{
key: transcripts
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/Human_Colon_Cancer_P2/Xenium_V1_Human_Colon_Cancer_P2_CRC_Add_on_FFPE/transcripts.parquet
}
]
folders: []
args: []
kwargs: [
{
key: min_genes
value: 1
}
{
key: min_counts
value: 1
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
]
1.7.2. Submitting a New Study with a Single Sample and Data:
group: User's group.species: Species of the dataset.title: Title of the new study.sample_name: Name of the new sample.sample_data: Data information, obtained fromconnector.parse_data_informationor a combination of its results.
submission_results = connector.submit(
group=DefaultGroup.PERSONAL_WORKSPACE.value,
species=Species.HUMAN.value,
title="Human Colon Cancer - 10xgenomics",
sample_name="Human_Colon_Cancer_P2",
sample_data=Xenium_V1_Human_Colon_Cancer_P2_submission_information + Visium_V2_Human_Colon_Cancer_P2_submission_information,
)
spatialx_connector.format_print(submission_results)
1.7.3. Adding a New Sample to an Existing Study:
study_id: ID of the study to which the sample is added.name: Name of the new sample.sample_data: Data information, obtained fromconnector.parse_data_informationor a combination of its results.
adding_sample_results = connector.add_sample(
study_id=submission_results[ConnectorKeys.STUDY_ID.value],
sample_name="Human_Colon_Cancer_P2 - New Sample",
sample_data=Visium_V2_Human_Colon_Cancer_P2_submission_information,
)
spatialx_connector.format_print(adding_sample_results)
1.7.4. Adding New Data to an Existing Sample:
study_id: ID of the study containing the sample.sample_id: ID of the existing sample.sample_data: Data information, obtained fromconnector.parse_data_informationor a combination of its results.
adding_sample_data_results = connector.add_sample_data(
study_id=adding_sample_results[ConnectorKeys.STUDY_ID.value],
sample_id=adding_sample_results[ConnectorKeys.SAMPLE_ID.value],
sample_data=Xenium_V1_Human_Colon_Cancer_P2_submission_information,
)
spatialx_connector.format_print(adding_sample_data_results)
1.7.5. Parsing Submission Information for Multiple Samples:
technology: Technology used for all samples (supports a single technology).data_path: Path to the directory containing multiple dataset subfolders (each subfolder represents a dataset).sample_name_mapping: Mapping of subfolder names to sample names.min_genes: Filtering cells by number of genes.min_counts: Filtering cells by number of transcripts.
multiple_cosmx_samples_submission_information = connector.parse_multiple_samples_information(
technology=Technologies.COSMX_VER1.value,
data_path=os.path.join(connector.s3["bioturingpublic"], "SpatialX_datasets/COSMX_VER1"),
sample_name_mapping={
"Lung6": "Human Lung Cancer - Sample 6",
"Lung9_Rep1": "Human Lung Cancer - Sample 9 Rep 1",
"Lung9_Rep2": "Human Lung Cancer - Sample 9 Rep 2",
},
data_name_mapping={
"Lung6": "Sample 6",
"Lung9_Rep1": "Sample 9 Rep 1",
"Lung9_Rep2": "Sample 9 Rep 2",
},
min_genes=10,
min_counts=10,
)
spatialx_connector.format_print(multiple_cosmx_samples_submission_information)
[
{
sample_name: Human Lung Cancer - Sample 6
data: [
{
name: Sample 6
submission_type: SUBMIT_SPATIAL_TRANSCRIPTOMICS
technology: COSMX_VER1
files: [
{
key: fov_positions
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung6/Lung6-Flat_files_and_images/Lung6_fov_positions_file.csv
}
{
key: transcripts
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung6/Lung6-Flat_files_and_images/Lung6_tx_file.csv
}
]
folders: [
{
key: images
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung6/Lung6-RawMorphologyImages
}
{
key: segmentation
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung6/Lung6-Flat_files_and_images/CellLabels
}
]
args: [
{
key: mpp
value: 0.18
}
]
kwargs: [
{
key: min_genes
value: 10
}
{
key: min_counts
value: 10
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
]
}
{
sample_name: Human Lung Cancer - Sample 9 Rep 1
data: [
{
name: Sample 9 Rep 1
submission_type: SUBMIT_SPATIAL_TRANSCRIPTOMICS
technology: COSMX_VER1
files: [
{
key: fov_positions
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep1/Lung9_Rep1-Flat_files_and_images/Lung9_Rep1_fov_positions_file.csv
}
{
key: transcripts
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep1/Lung9_Rep1-Flat_files_and_images/Lung9_Rep1_tx_file.csv
}
]
folders: [
{
key: images
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep1/Lung9_Rep1-RawMorphologyImages
}
{
key: segmentation
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep1/Lung9_Rep1-Flat_files_and_images/CellLabels
}
]
args: [
{
key: mpp
value: 0.18
}
]
kwargs: [
{
key: min_genes
value: 10
}
{
key: min_counts
value: 10
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
]
}
{
sample_name: Human Lung Cancer - Sample 9 Rep 2
data: [
{
name: Sample 9 Rep 2
submission_type: SUBMIT_SPATIAL_TRANSCRIPTOMICS
technology: COSMX_VER1
files: [
{
key: fov_positions
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep2/Lung9_Rep2-Flat_files_and_images/Lung9_Rep2_fov_positions_file.csv
}
{
key: transcripts
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep2/Lung9_Rep2-Flat_files_and_images/Lung9_Rep2_tx_file.csv
}
]
folders: [
{
key: images
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep2/Lung9_Rep2-RawMorphologyImages
}
{
key: segmentation
value: <USER_PATH>/public_cloud/bioturingpublic/SpatialX_datasets/COSMX_VER1/Lung9_Rep2/Lung9_Rep2-Flat_files_and_images/CellLabels
}
]
args: [
{
key: mpp
value: 0.18
}
]
kwargs: [
{
key: min_genes
value: 10
}
{
key: min_counts
value: 10
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
]
}
]
multiple_samples_submission_results = connector.submit_multiple_samples(
group=DefaultGroup.PERSONAL_WORKSPACE.value,
species=Species.HUMAN.value,
title="Multiple Human Lung Cancer - CosMX Ver1",
sample_data=multiple_cosmx_samples_submission_information,
)
spatialx_connector.format_print(multiple_samples_submission_results)
1.7.6. Data Details and Element Management
This section provides tools for accessing and modifying data within existing studies. Users can perform the following actions:
- Retrieve Data:
- Fetch data identifiers (IDs).
- List existing elements associated with a dataset.
- Download recently submitted AnnData objects.
- Append Data Elements:
- Add individual data components to a dataset, including:
- Protein images.
- Segmentation masks.
- Expression matrices.
- Metadata.
- Embeddings.
- Add individual data components to a dataset, including:
- Submit Protein Images: Submit protein images as a study.
human_pancreas_codex_information = connector.parse_data_information(
data_name="human_pancreas_codex",
technology=Technologies.PROTEIN_QPTIFF.value,
data_path=os.path.join(connector.s3["bioturingpublic"], "mount/examples/spatialx/human_pancreas_codex"),
args={SubmissionElementKeys.MPP.value: 1},
)
spatialx_connector.format_print(human_pancreas_codex_information)
[
{
name: human_pancreas_codex
submission_type: SUBMIT_SPATIAL_PROTEOMICS
technology: PROTEIN_QPTIFF
files: [
{
key: protein_images
value: <USER_PATH>/public_cloud/bioturingpublic/mount/examples/spatialx/human_pancreas_codex/human_pancreas_codex.qptiff
}
]
folders: []
args: [
{
key: mpp
value: None
}
{
key: mpp
value: 1
}
]
kwargs: [
{
key: min_genes
value: 1
}
{
key: min_counts
value: 1
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
]
proteomics_submission_results = connector.submit(
group=DefaultGroup.PERSONAL_WORKSPACE.value,
species=Species.HUMAN.value,
title="Human Pancreas CODEX",
sample_name="Human Pancreas CODEX",
sample_data=human_pancreas_codex_information,
tracking=True,
)
spatialx_connector.format_print(proteomics_submission_results)
[2025-04-15 03:24:41] [3992384] [INFO] SUBMISSION JOB IS WAITING AT ID 15.
[2025-04-15 03:24:41] [3991797] [INFO] RUN SUBMISSION JOB ID 15: {
study_path: <USER_PATH>/study/ST-01JRVQMFQVB2JYPM27C331SXKB
sample_id: SP-01JRVQMG6SD4EGW2VNYJ6RVP0E
submit_id: SB-01JRVQMG6SD4EGW2VNYHDJ6GRG
data_id: DA-01JRVQMG70A1G37MZBB6E538C8
submission_name: human_pancreas_codex
submission_info: {
submission_type: SUBMIT_SPATIAL_PROTEOMICS
technology: PROTEIN_QPTIFF
files: [
{
key: protein_images
value: <USER_PATH>/public_cloud/bioturingpublic/mount/examples/spatialx/human_pancreas_codex/human_pancreas_codex.qptiff
}
]
folders: []
args: [
{
key: mpp
value: None
}
{
key: mpp
value: 1
}
]
kwargs: [
{
key: min_genes
value: 1
}
{
key: min_counts
value: 1
}
{
key: mito_controls_percentage
value: 0.25
}
]
identities: []
}
}.
[2025-04-15 03:24:41] [3991797] [INFO] SUBMIT: protein_images
[2025-04-15 03:24:41] [3991797] [INFO] Fail to extracting channel information of OME-Tiff: 'NoneType' object has no attribute 'encode'
[2025-04-15 03:24:41] [3991797] [INFO] Write protein_images.
[2025-04-15 03:24:41] [3991797] [INFO] Process channel ACTA2
[2025-04-15 03:24:41] [3991797] [INFO] Write image channel
[2025-04-15 03:25:05] [3991797] [INFO] - progress: 100%
[2025-04-15 03:25:05] [3991797] [INFO] Process channel BETA-CATENIN
[2025-04-15 03:25:05] [3991797] [INFO] Write image channel
[2025-04-15 03:25:24] [3991797] [INFO] - progress: 100%
[2025-04-15 03:25:24] [3991797] [INFO] Process channel CGC
[2025-04-15 03:25:24] [3991797] [INFO] Write image channel
[2025-04-15 03:25:43] [3991797] [INFO] - progress: 100%
[2025-04-15 03:25:43] [3991797] [INFO] Process channel CHGA
[2025-04-15 03:25:43] [3991797] [INFO] Write image channel
[2025-04-15 03:26:04] [3991797] [INFO] - progress: 100%
[2025-04-15 03:26:04] [3991797] [INFO] Process channel CPEP
[2025-04-15 03:26:04] [3991797] [INFO] Write image channel
[2025-04-15 03:26:22] [3991797] [INFO] - progress: 100%
[2025-04-15 03:26:22] [3991797] [INFO] Process channel E-CADHERIN
[2025-04-15 03:26:22] [3991797] [INFO] Write image channel
[2025-04-15 03:26:40] [3991797] [INFO] - progress: 100%
[2025-04-15 03:26:40] [3991797] [INFO] Process channel HOECHST
[2025-04-15 03:26:40] [3991797] [INFO] Write image channel
[2025-04-15 03:27:02] [3991797] [INFO] - progress: 100%
[2025-04-15 03:27:02] [3991797] [INFO] Process channel IAPP
[2025-04-15 03:27:02] [3991797] [INFO] Write image channel
[2025-04-15 03:27:21] [3991797] [INFO] - progress: 100%
[2025-04-15 03:27:21] [3991797] [INFO] Process channel KRT19
[2025-04-15 03:27:21] [3991797] [INFO] Write image channel
[2025-04-15 03:27:37] [3991797] [INFO] - progress: 100%
[2025-04-15 03:27:37] [3991797] [INFO] Process channel PECAM-1
[2025-04-15 03:27:37] [3991797] [INFO] Write image channel
[2025-04-15 03:27:59] [3991797] [INFO] - progress: 100%
[2025-04-15 03:27:59] [3991797] [INFO] Process channel SST
[2025-04-15 03:27:59] [3991797] [INFO] Write image channel
[2025-04-15 03:28:17] [3991797] [INFO] - progress: 100%
[2025-04-15 03:28:17] [3991797] [INFO] Process channel VIM
[2025-04-15 03:28:17] [3991797] [INFO] Write image channel
[2025-04-15 03:28:39] [3991797] [INFO] - progress: 100%
[2025-04-15 03:28:39] [3991797] [INFO] SUBMISSION DONE: {
protein_images: {
59656ac1f77c445286ac2b4b3bbef5d2: human_pancreas_codex_protein_images
}
}.
[2025-04-15 03:28:39] [3991797] [INFO] DONE.
{
study_id: ST-01JRVQMFQVB2JYPM27C331SXKB
sample_id: SP-01JRVQMG6SD4EGW2VNYJ6RVP0E
sample_data: [
{
data_id: DA-01JRVQMG70A1G37MZBB6E538C8
submit_id: SB-01JRVQMG6SD4EGW2VNYHDJ6GRG
submit_name: human_pancreas_codex
}
]
submit_id: SB-01JRVQMG6SD4EGW2VNYHDJ6GRG
job_id: 15
err_message:
}
- Retrieving Data ID: Obtain the
data_idfor adding extended elements and running analyses.
STUDY_ID = proteomics_submission_results[ConnectorKeys.STUDY_ID.value]
SAMPLE_ID = proteomics_submission_results[ConnectorKeys.SAMPLE_ID.value]
DATA_ID = proteomics_submission_results[ConnectorKeys.SAMPLE_DATA.value][0][ConnectorKeys.DATA_ID.value]
- Adding New Segmentation: Add a new segmentation layer to the dataset.
add_segmentation_result = connector.add_sample_data_element(
title="Proteomics Segmentation",
data_id=DATA_ID,
adding_types=[SegmentationSubmission.PARQUET.value],
paths={
SubmissionElementKeys.SEGMENTATION.value: os.path.join(
connector.s3["bioturingpublic"],
"mount/examples/spatialx/human_pancreas_codex/human_pancreas_segmentation.parquet",
)
},
tracking=True,
)
spatialx_connector.format_print(add_segmentation_result)
[2025-04-15 03:28:39] [3992386] [INFO] SUBMISSION JOB IS WAITING AT ID 16.
[2025-04-15 03:28:39] [3991788] [INFO] RUN SUBMISSION JOB ID 16: {
study_path: <USER_PATH>/study/ST-01JRVQMFQVB2JYPM27C331SXKB
sample_id: SP-01JRVQMG6SD4EGW2VNYJ6RVP0E
submit_id: SB-01JRVQVSE1WBZN6FC94MZPY7KD
data_id: DA-01JRVQMG70A1G37MZBB6E538C8
submission_name: Proteomics Segmentation
submission_info: {
submission_type: SUBMIT_SPATIAL_PROTEOMICS
technology: PROTEIN_QPTIFF
files: [
{
key: segmentation
value: <USER_PATH>/public_cloud/bioturingpublic/mount/examples/spatialx/human_pancreas_codex/human_pancreas_segmentation.parquet
}
]
folders: []
args: [
{
key: ignore_technology_elements
value: True
}
]
kwargs: []
identities: [
SEGMENTATION_PARQUET
]
}
}.
[2025-04-15 03:28:40] [3991788] [INFO] SUBMIT: segmentation
[2025-04-15 03:28:41] [3991788] [INFO] Write segmentation.
[2025-04-15 03:28:49] [3991788] [INFO] SUBMIT: cell_centers
[2025-04-15 03:28:52] [3991788] [INFO] SUBMISSION DONE: {
segmentation: {
a84b2978677c49c18395825a16ce9738: Proteomics Segmentation_segmentation
}
cell_centers: {
589255595512465c9b49d882385b6c0a: Proteomics Segmentation_cell_centers
}
}.
[2025-04-15 03:28:52] [3991788] [INFO] DONE.
{
study_id: ST-01JRVQMFQVB2JYPM27C331SXKB
sample_id: SP-01JRVQMG6SD4EGW2VNYJ6RVP0E
sample_data: None
submit_id: SB-01JRVQVSE1WBZN6FC94MZPY7KD
job_id: 16
err_message:
}
- Retrieving Existing Elements: List the existing elements associated with the dataset.
sample_data_elements = connector.get_sample_data_elements(DATA_ID)
spatialx_connector.format_print(sample_data_elements)
{
protein_images: [
human_pancreas_codex_protein_images
]
cell_centers: [
Proteomics Segmentation_cell_centers
]
segmentation: [
Proteomics Segmentation_segmentation
]
}
- Adding New Expression Matrix: Add a new expression matrix to the dataset.
add_expression_result = connector.add_sample_data_element(
title="Proteomics Expression",
data_id=DATA_ID,
adding_types=[ExpressionSubmission.IMPORT_ANNDATA.value],
paths={
SubmissionElementKeys.EXPRESSION.value: os.path.join(
connector.s3["bioturingpublic"],
"mount/examples/spatialx/human_pancreas_codex/human_pancreas_protein.h5ad",
),
},
args={
SubmissionElementKeys.SPATIAL_ID.value: sample_data_elements[SubmissionElementKeys.CELL_CENTERS.value][0],
},
tracking=True,
)
spatialx_connector.format_print(add_expression_result)
[2025-04-15 03:28:52] [3992386] [INFO] SUBMISSION JOB IS WAITING AT ID 17.
[2025-04-15 03:28:52] [3991797] [INFO] RUN SUBMISSION JOB ID 17: {
study_path: <USER_PATH>/study/ST-01JRVQMFQVB2JYPM27C331SXKB
sample_id: SP-01JRVQMG6SD4EGW2VNYJ6RVP0E
submit_id: SB-01JRVQW63HR24Q8NBCPAWFGWPZ
data_id: DA-01JRVQMG70A1G37MZBB6E538C8
submission_name: Proteomics Expression
submission_info: {
submission_type: SUBMIT_SPATIAL_PROTEOMICS
technology: PROTEIN_QPTIFF
files: [
{
key: annotated_data
value: <USER_PATH>/public_cloud/bioturingpublic/mount/examples/spatialx/human_pancreas_codex/human_pancreas_protein.h5ad
}
]
folders: []
args: [
{
key: spatial_id
value: Proteomics Segmentation_cell_centers
}
{
key: ignore_technology_elements
value: True
}
]
kwargs: []
identities: [
IMPORT_ANNDATA
]
}
}.
[2025-04-15 03:28:52] [3991797] [INFO] SUBMIT: annotated_data
[2025-04-15 03:28:53] [3991797] [INFO] Start importing
[2025-04-15 03:28:53] [3991797] [INFO] Merging expression
[2025-04-15 03:28:53] [3991797] [INFO] Processing obsm
[2025-04-15 03:28:53] [3991797] [INFO] Processing obs
[2025-04-15 03:28:54] [3991797] [INFO] Add to table
[2025-04-15 03:28:54] [3991797] [INFO] Run scVI.
[2025-04-15 03:28:55] [3991797] [INFO] Parameters: {
batch_key: None
n_latents: 20
encode_covariates: True
n_layers: 2
train_size: 0.9
dropout_rate: 0.2
n_top_genes: 10
}.
[2025-04-15 03:28:55] [3991797] [INFO] Parameters: {
batch_key: None
n_latents: 20
encode_covariates: True
n_layers: 2
train_size: 0.9
dropout_rate: 0.2
n_top_genes: 10
}.
[2025-04-15 03:28:56] [3991797] [INFO] Run highly_variable_genes.
[2025-04-15 03:28:56] [3991797] [INFO] Parameters: {
n_top_genes: 10
flavor: seurat
normalize_method: raw
}.
[2025-04-15 03:28:59] [3991797] [INFO] Run highly_variable_genes successfully.
[2025-04-15 03:29:00] [3991797] [INFO] Training scVI...
[2025-04-15 03:29:00] [3991797] [WARNING] /home/nhat/BioTuring/spatialx/runtimes/pyapps/lib/python3.10/site-packages/scvi/data/fields/_base_field.py:64: UserWarning: adata.X does not contain unnormalized count data. Are you sure this is what you want?
self.validate_field(adata)
[2025-04-15 03:29:00] [3991797] [WARNING] /home/nhat/BioTuring/spatialx/runtimes/pyapps/lib/python3.10/site-packages/lightning/pytorch/trainer/connectors/data_connector.py:441: The 'train_dataloader' does not have many workers which may be a bottleneck. Consider increasing the value of the `num_workers` argument` to `num_workers=47` in the `DataLoader` to improve performance.
[2025-04-15 03:29:00] [3991797] [WARNING] /home/nhat/BioTuring/spatialx/runtimes/pyapps/lib/python3.10/site-packages/lightning/pytorch/trainer/connectors/data_connector.py:441: The 'val_dataloader' does not have many workers which may be a bottleneck. Consider increasing the value of the `num_workers` argument` to `num_workers=47` in the `DataLoader` to improve performance.
[2025-04-15 03:29:51] [3991797] [INFO] Inferring scVI...
[2025-04-15 03:29:52] [3991797] [INFO] Run scVI successfully.
[2025-04-15 03:29:52] [3991797] [INFO] Run neighbors_graph.
[2025-04-15 03:29:52] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:29:53] [3991797] [INFO] Run nearest_neighbors.
[2025-04-15 03:29:53] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
}.
[2025-04-15 03:29:53] [3991797] [INFO] Run nearest_neighbors successfully.
[2025-04-15 03:29:54] [3991797] [INFO] Run neighbors_graph successfully.
[2025-04-15 03:29:54] [3991797] [INFO] Run t-SNE.
[2025-04-15 03:29:54] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
perplexity: 30
init: pca
}.
[2025-04-15 03:29:55] [3991797] [INFO] Run nearest_neighbors.
[2025-04-15 03:29:55] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
}.
[2025-04-15 03:29:55] [3991797] [INFO] Found analytic results for nearest_neighbors with same parameters.
[2025-04-15 03:29:57] [3991797] [INFO] Run t-SNE successfully.
[2025-04-15 03:29:57] [3991797] [INFO] Run UMAP.
[2025-04-15 03:29:57] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 15
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
init: pca
deterministic: False
}.
[2025-04-15 03:29:57] [3991797] [INFO] Run neighbors_graph.
[2025-04-15 03:29:57] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 15
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:29:58] [3991797] [INFO] Run nearest_neighbors.
[2025-04-15 03:29:58] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 15
}.
[2025-04-15 03:29:58] [3991797] [INFO] Run nearest_neighbors successfully.
[2025-04-15 03:29:58] [3991797] [INFO] Run neighbors_graph successfully.
[2025-04-15 03:29:59] [3991797] [INFO] Run UMAP successfully.
[2025-04-15 03:29:59] [3991797] [INFO] Run Louvain.
[2025-04-15 03:29:59] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
resolution: 0.1
max_clusters: 100
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:29:59] [3991797] [INFO] Run neighbors_graph.
[2025-04-15 03:29:59] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:29:59] [3991797] [INFO] Found analytic results for neighbors_graph with same parameters.
[2025-04-15 03:29:59] [3991797] [INFO] Run Louvain successfully.
[2025-04-15 03:30:00] [3991797] [INFO] Run Louvain.
[2025-04-15 03:30:00] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
resolution: 0.5
max_clusters: 100
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:30:00] [3991797] [INFO] Run neighbors_graph.
[2025-04-15 03:30:00] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:30:00] [3991797] [INFO] Found analytic results for neighbors_graph with same parameters.
[2025-04-15 03:30:00] [3991797] [INFO] Run Louvain successfully.
[2025-04-15 03:30:01] [3991797] [INFO] Run Louvain.
[2025-04-15 03:30:01] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
resolution: 1
max_clusters: 100
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:30:01] [3991797] [INFO] Run neighbors_graph.
[2025-04-15 03:30:01] [3991797] [INFO] Parameters: {
embedding_key: f7078b35e2834e389a0ed2cbd9726135
is_global: True
n_neighbors: 90
local_connectivity: 1
bandwidth: 1
mix_ratio: 1
}.
[2025-04-15 03:30:01] [3991797] [INFO] Found analytic results for neighbors_graph with same parameters.
[2025-04-15 03:30:01] [3991797] [INFO] Run Louvain successfully.
[2025-04-15 03:30:01] [3991797] [INFO] SUBMISSION DONE: {
annotated_data: {
4a90b06c28e94f75998ded7e03e43a1c: Proteomics Expression_annotated_data
}
cell_centers: {
589255595512465c9b49d882385b6c0a: Proteomics Segmentation_cell_centers
}
}.
[2025-04-15 03:30:01] [3991797] [INFO] DONE.
{
study_id: ST-01JRVQMFQVB2JYPM27C331SXKB
sample_id: SP-01JRVQMG6SD4EGW2VNYJ6RVP0E
sample_data: None
submit_id: SB-01JRVQW63HR24Q8NBCPAWFGWPZ
job_id: 17
err_message:
}
- Get AnnData: Get recently submitted AnnData
sample_data_elements = connector.get_sample_data_elements(DATA_ID)
adata = connector.get_anndata(
data_id=DATA_ID,
anndata_id=sample_data_elements[SpatialAttrs.ANNOTATED_DATA.value][0]
)
adata
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
protein_names = adata.var_names.values
protein_names
array(['ACTA2', 'BETA-CATENIN', 'CGC', 'CHGA', 'CPEP', 'E-CADHERIN',
'HOECHST', 'IAPP', 'KRT19', 'PECAM-1', 'SST', 'VIM'], dtype=object)
- Import Metadata: Upload new metadata to server
- Create metadata to upload (we can use
pd.read_csvto read a dataframe instead)
new_dataframe = pd.DataFrame(
{
"ACTA2": adata.X[:, 0].toarray().reshape(-1) * adata.obs["area"].values
},
index=adata.obs_names
)
new_dataframe["Sample"] = "pancreas"
new_dataframe.head(5)
.dataframe tbody tr th {
vertical-align: top;
}
.dataframe thead th {
text-align: right;
}
| ACTA2 | Sample | |
|---|---|---|
| Barcodes | ||
| 0 | 8.081808 | pancreas |
| 1 | 1.339335 | pancreas |
| 2 | 2.115501 | pancreas |
| 3 | 1.514645 | pancreas |
| 4 | 2.824838 | pancreas |
- Upload metadata to server
adata.upload_metadata(new_dataframe)
adata
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'ACTA2', 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'Sample', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
- Import Embeddings: Upload new embeddings to server. To make sure that the embeddings match with cell barcodes, you need to assign it to
adata.obsm["count_intensity"] = adata.obsm["counts_per_pixel"] * adata.obs["area"].values.reshape((-1, 1))
adata.upload_embeddings("count_intensity") # Or ["count_intensity"], for multiple embeddings
adata
[2025-04-11 17:19:35] [2164367] [INFO] ANALYSIS JOB IS WAITING AT ID 14.
[2025-04-11 17:19:35] [2163580] [INFO] ANALYZE: import_embeddings.
[2025-04-11 17:19:35] [2163580] [INFO] import_anndata_embeddings
[2025-04-11 17:19:36] [2163580] [INFO] Run import_anndata_embeddings successfully.
[2025-04-11 17:19:36] [2163580] [INFO] ANALYSIS DONE: {}.
[2025-04-11 17:19:36] [2163580] [INFO] DONE.
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'ACTA2', 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'Sample', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'count_intensity', 'count_intensity (1)', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
1.7.7. Customing Data Structure of Submission
- Create an empty study
This section enables users to submit data with customized data structures, facilitating the upload of diverse spatial omics datasets, such as spatial proteomics data from Akoya CODEX.
study_id = connector.create_study(
group=DefaultGroup.PERSONAL_WORKSPACE,
species=Species.HUMAN,
title="Human Pancreas - CODEX"
)
study_id
- Custom you data structure with multiple types of elements and paths, arguments
connector.add_custom_sample(
study_id=study_id,
sample_name="human_pancreas_codex",
data_name="human_pancreas_codex",
technology=Technologies.PROTEIN_QPTIFF,
adding_types=[
ImagesSubmission.PROTEIN_QPTIFF,
SegmentationSubmission.PARQUET,
ExpressionSubmission.IMPORT_ANNDATA,
],
paths={
SubmissionElementKeys.PROTEIN_IMAGES: os.path.join(
connector.s3["bioturingpublic"],
"mount/examples/spatialx/human_pancreas_codex/human_pancreas_codex.qptiff",
),
SubmissionElementKeys.SEGMENTATION: os.path.join(
connector.s3["bioturingpublic"],
"mount/examples/spatialx/human_pancreas_codex/human_pancreas_segmentation.parquet",
),
SubmissionElementKeys.EXPRESSION: os.path.join(
connector.s3["bioturingpublic"],
"mount/examples/spatialx/human_pancreas_codex/human_pancreas_protein.h5ad",
),
},
args={
SubmissionElementKeys.MPP: 1,
},
)
1.8. Accessing study information
Use these functions to get detailed information about your studies in different workspaces.
1.8.1. Listing studies:
The following code retrieves a list of your studies within your personal workspace. To list studies in a different group, replace DefaultGroup.PERSONAL_WORKSPACE.value with the desired group name (e.g., "Demo").
studies = connector.list_study(
group=DefaultGroup.PERSONAL_WORKSPACE.value,
species=Species.HUMAN.value,
)
studies
│ Study ID │ Title │ Total Sample │ Total Data │
───┼───────────────────────────────┼─────────────────────────────────────────┼──────────────┼────────────┤
0 │ ST-01JRVQMFQVB2JYPM27C331SXKB │ Human Pancreas CODEX │ 1 │ 1 │
1 │ ST-01JRJ7SVR6VVDF5A1BCBCM815R │ Human Pancreas CODEX │ 1 │ 1 │
2 │ ST-01JRFJE3H5XD1T3QAX56Y97WSF │ Human Ovary Cancer │ 1 │ 1 │
3 │ ST-01JPWFDSNSY8DZA18V00Z7FKSA │ Human Lung Cancer │ 10 │ 10 │
4 │ ST-01JQ0XFZSZ5FX32QKG0ZECRRR0 │ Xenium Human Colon │ 6 │ 6 │
5 │ ST-01JQ5SR20KTGYFYRHHYM9NTVVV │ Human Colon Colorectal Cancer Patient 2 │ 3 │ 5 │
6 │ ST-01JQ87VD8RE1G56X8DVDEAPYTB │ Human Ovarian Cancer │ 2 │ 2 │
7 │ ST-01JQNJQDJNYWSSECFGGNZSJ4RJ │ LiverCancerFiles │ 1 │ 2 │
8 │ ST-01JRAPB8K59AGY7T4ZZ1E68C7G │ Human Pancreas CODEX │ 1 │ 1 │
9 │ ST-01JQE0QCY9EHZ7FWXPR065SNRG │ Lung6 │ 1 │ 1 │
10 │ ST-01JQNQD3RX5007ADGVZ1P7CDZV │ CosMx Proteomics │ 1 │ 1 │
11 │ ST-01JQ65XM31WEB5CE8QDHKZ9JG1 │ B01207E3G4 │ 1 │ 2 │
1.8.2. Select a study to analyze:
To select a study to analyze, select with the desired study's identifier or the number.
study = studies[0] # Or: study = studies[STUDY_ID]
study
Human Pancreas CODEX (Species: human)
0. [Sample - SP-01JRVQMG6SD4EGW2VNYJ6RVP0E] Human Pancreas CODEX (Status: Ready to use)
0. [Data - DA-01JRVQMG70A1G37MZBB6E538C8] human_pancreas_codex (Technology: PROTEIN_QPTIFF - Status: Ready to use)
1.8.3. Select a Sample within a Study:
sample = study[0] # Or: sample = study[SAMPLE_ID]
sample
[Sample - SP-01JRVQMG6SD4EGW2VNYJ6RVP0E] Human Pancreas CODEX (Status: Ready to use)
0. [Data - DA-01JRVQMG70A1G37MZBB6E538C8] human_pancreas_codex (Technology: PROTEIN_QPTIFF - Status: Ready to use)
1.8.4. Select a data for seeing elements:
data = sample[0] # Or: data = sample[DATA_ID]
data.spatialdata
SpatialData with elements:
Images: human_pancreas_codex_protein_images
Shapes: Proteomics Segmentation_segmentation
Points: Proteomics Segmentation_cell_centers
Tables: Proteomics Expression_annotated_data
1.8.5. Interact with elements:
- Select an image
images = data.spatialdata.images["human_pancreas_codex_protein_images"]
images["0"].shape
(12, 18440, 25816)
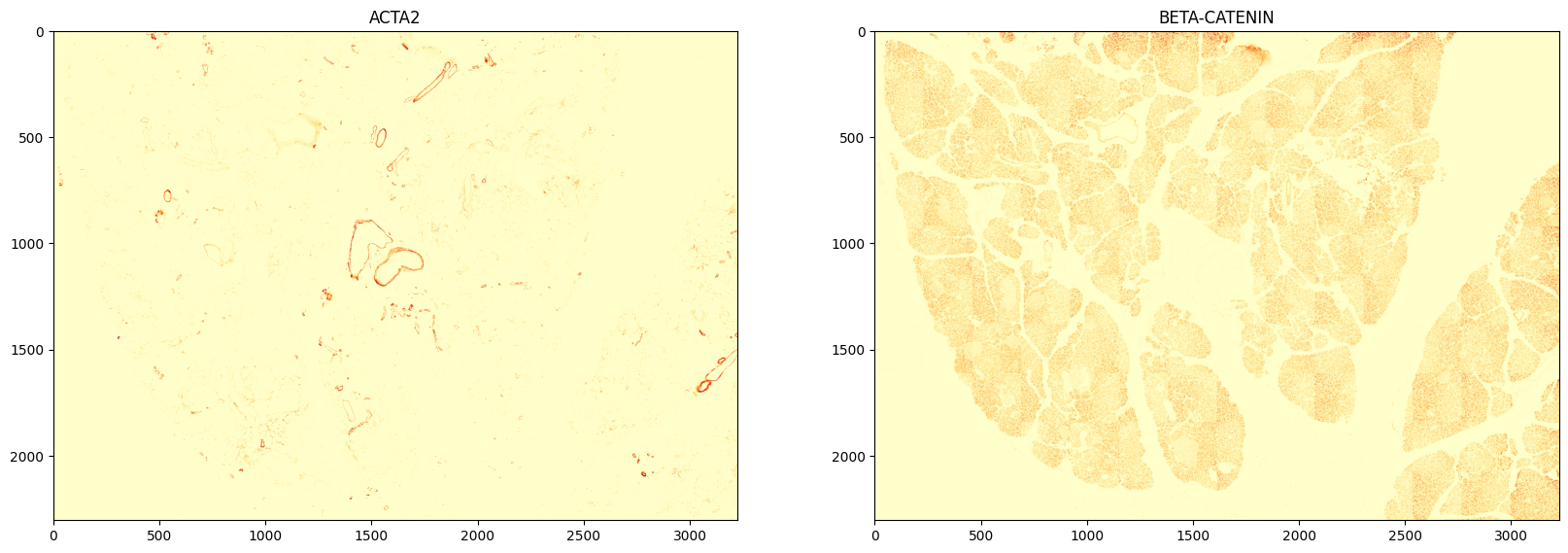
- Show image in low resolution
pyramid_level = 3
channel_names = data.spatialdata.images.get_channel_names(images)
plt.figure(1, (20, 10))
plt.subplot(1, 2, 1)
plt.imshow(images[pyramid_level][0], cmap="YlOrRd")
plt.title(channel_names[0])
plt.subplot(1, 2, 2)
plt.imshow(images[pyramid_level][1], cmap="YlOrRd")
plt.title(channel_names[1])
plt.show()
- Simplify segmentation
data.spatialdata.shapes.simplify("Proteomics Segmentation_segmentation", 1)
1.0
- Load segmentation
data.spatialdata.shapes["Proteomics Segmentation_segmentation"]
.dataframe tbody tr th {
vertical-align: top;
}
.dataframe thead th {
text-align: right;
}
| geometry | |
|---|---|
| 0 | POLYGON ((609 282, 603 285, 601 289, 610 291, ... |
| 1 | POLYGON ((557 349, 553 352, 553 354, 561 353, ... |
| 2 | POLYGON ((653 347, 646 351, 646 357, 650 362, ... |
| 3 | POLYGON ((623 360, 621 365, 625 372, 643 370, ... |
| 4 | POLYGON ((512 367, 506 369, 509 376, 528 372, ... |
| ... | ... |
| 269892 | POLYGON ((25546 18310, 25541 18320, 25548 1832... |
| 269893 | POLYGON ((25525 18310, 25516 18317, 25516 1833... |
| 269894 | POLYGON ((25546 18335, 25540 18338, 25536 1834... |
| 269895 | POLYGON ((25527 18357, 25523 18360, 25525 1837... |
| 269896 | POLYGON ((25535 18429, 25536 18438, 25546 1843... |
269897 rows × 1 columns
- Load cell centers
points = data.spatialdata.points["Proteomics Segmentation_cell_centers"]
points.compute()
.dataframe tbody tr th {
vertical-align: top;
}
.dataframe thead th {
text-align: right;
}
| x | y | |
|---|---|---|
| Barcodes | ||
| 0 | 608.344543 | 286.925079 |
| 1 | 557.166687 | 352.210541 |
| 2 | 654.912659 | 354.925751 |
| 3 | 631.174866 | 366.474548 |
| 4 | 516.161316 | 371.587189 |
| ... | ... | ... |
| 269892 | 25549.826172 | 18318.613281 |
| 269893 | 25524.333984 | 18321.414062 |
| 269894 | 25545.021484 | 18346.152344 |
| 269895 | 25527.964844 | 18363.166016 |
| 269896 | 25539.058594 | 18434.892578 |
269897 rows × 2 columns
1.8.4. Select a Annotated Data for more analysis:
adata = data.spatialdata.tables["Proteomics Expression_annotated_data"]
adata
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - tSNE', 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
1.9. Analysis
You can now run analyses and see the results directly in the SpatialX connector! This is our first version, and we're planning to add features like analysis logs and result export soon. Stay tuned for updates! Also, we'd love to hear your feedback! If you have any function requests, please reach out to us at support@bioturing.com.
from spatialx_connector import analysis
1.9.1. Embeddings
analysis.pp.pca(adata, title="Connector - PCA")
adata
[2025-04-15 03:50:50] [3992189] [INFO] ANALYSIS JOB IS WAITING AT ID 19.
[2025-04-15 03:50:50] [3991783] [INFO] ANALYZE: PCA.
[2025-04-15 03:50:50] [3991783] [INFO] Run PCA.
[2025-04-15 03:50:50] [3991783] [INFO] Parameters: {
n_dims: 50
normalize_method: log1p-normalized
batch_key: None
}.
[2025-04-15 03:50:51] [3991783] [WARNING] 50 is to many dimensions with genes expression has 269897 cells and 12 genes.
[2025-04-15 03:50:51] [3991783] [WARNING] Set number of dimensions to 11
[2025-04-15 03:50:51] [3991783] [INFO] Run PCA successfully.
[2025-04-15 03:50:51] [3991783] [INFO] ANALYSIS DONE: {"obsm": "ff6770f42ba04e6eb7c21dfdafe18973"}.
[2025-04-15 03:50:51] [3991783] [INFO] DONE.
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
analysis.pp.scvi(adata, title="Connector - scVI", n_top_genes=2000)
adata
[2025-04-15 04:08:43] [3992386] [INFO] ANALYSIS JOB IS WAITING AT ID 21.
[2025-04-15 04:08:43] [3991790] [INFO] ANALYZE: scVI.
[2025-04-15 04:08:43] [3991790] [INFO] Run scVI.
[2025-04-15 04:08:45] [3991790] [INFO] Parameters: {
batch_key: None
n_latents: 20
encode_covariates: True
n_layers: 2
train_size: 0.9
dropout_rate: 0.2
n_top_genes: 2000
}.
[2025-04-15 04:08:45] [3991790] [INFO] Parameters: {
batch_key: None
n_latents: 20
encode_covariates: True
n_layers: 2
train_size: 0.9
dropout_rate: 0.2
n_top_genes: 2000
}.
[2025-04-15 04:08:45] [3991790] [INFO] Training scVI...
[2025-04-15 04:08:45] [3991790] [WARNING] /home/nhat/BioTuring/spatialx/runtimes/pyapps/lib/python3.10/site-packages/scvi/data/fields/_base_field.py:64: UserWarning: adata.X does not contain unnormalized count data. Are you sure this is what you want?
self.validate_field(adata)
[2025-04-15 04:08:45] [3991790] [WARNING] /home/nhat/BioTuring/spatialx/runtimes/pyapps/lib/python3.10/site-packages/lightning/pytorch/trainer/connectors/data_connector.py:441: The 'train_dataloader' does not have many workers which may be a bottleneck. Consider increasing the value of the `num_workers` argument` to `num_workers=47` in the `DataLoader` to improve performance.
[2025-04-15 04:08:45] [3991790] [WARNING] /home/nhat/BioTuring/spatialx/runtimes/pyapps/lib/python3.10/site-packages/lightning/pytorch/trainer/connectors/data_connector.py:441: The 'val_dataloader' does not have many workers which may be a bottleneck. Consider increasing the value of the `num_workers` argument` to `num_workers=47` in the `DataLoader` to improve performance.
[2025-04-15 04:09:36] [3991790] [INFO] Monitored metric elbo_validation did not improve in the last 45 records. Best score: 87.137. Signaling Trainer to stop.
[2025-04-15 04:09:36] [3991790] [INFO] Inferring scVI...
[2025-04-15 04:09:38] [3991790] [INFO] Run scVI successfully.
[2025-04-15 04:09:38] [3991790] [INFO] ANALYSIS DONE: {"obsm": "a90833bf89dd49baba3ed7d47b04492b"}.
[2025-04-15 04:09:38] [3991790] [INFO] DONE.
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - scVI', 'Connector - tSNE', 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
analysis.tl.umap(adata, embedding_key="Connector - PCA", title="Connector - UMAP")
adata
[2025-04-04 17:21:26] [3762126] [INFO] ANALYSIS JOB IS WAITING AT ID 529.
[2025-04-04 17:21:26] [3780787] [INFO] ANALYZE: UMAP
[2025-04-04 17:21:27] [3780787] [INFO] Run UMAP.
[2025-04-04 17:21:27] [3780787] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_neighbors: 15
local_connectivity: 1.0
bandwidth: 1
mix_ratio: 1
init: pca
deterministic: True
}.
[2025-04-04 17:21:27] [3780787] [INFO] Run neighbors_graph.
[2025-04-04 17:21:27] [3780787] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_neighbors: 15
local_connectivity: 1.0
bandwidth: 1
mix_ratio: 1
}.
[2025-04-04 17:21:27] [3780787] [INFO] Run nearest_neighbors.
[2025-04-04 17:21:27] [3780787] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_neighbors: 15
}.
[2025-04-04 17:21:27] [3780787] [INFO] Run nearest_neighbors successfully
[2025-04-04 17:21:28] [3780787] [INFO] Run neighbors_graph successfully.
[2025-04-04 17:21:28] [3780787] [INFO] Run UMAP successfully.
[2025-04-04 17:21:28] [3780787] [INFO] ANALYSIS DONE: {"obsm": "2f62a8a1997741268622e5cf7d66cb24"}.
[2025-04-04 17:21:28] [3780787] [INFO] DONE
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - PCA (1)', 'Connector - UMAP', 'Spatial Cell centers', 'counts_per_pixel', 'spatial'
analysis.tl.tsne(adata, embedding_key="Connector - PCA", title="Connector - tSNE")
adata
[2025-04-15 03:51:00] [3992386] [INFO] ANALYSIS JOB IS WAITING AT ID 20.
[2025-04-15 03:51:00] [3991792] [INFO] ANALYZE: t-SNE.
[2025-04-15 03:51:00] [3991792] [INFO] Run t-SNE.
[2025-04-15 03:51:00] [3991792] [INFO] Parameters: {
embedding_key: ff6770f42ba04e6eb7c21dfdafe18973
is_global: True
n_neighbors: 90
perplexity: 30.0
init: pca
}.
[2025-04-15 03:51:00] [3991792] [INFO] Run nearest_neighbors.
[2025-04-15 03:51:00] [3991792] [INFO] Parameters: {
embedding_key: ff6770f42ba04e6eb7c21dfdafe18973
is_global: True
n_neighbors: 90
}.
[2025-04-15 03:51:01] [3991792] [INFO] Run nearest_neighbors successfully.
[2025-04-15 03:51:03] [3991792] [INFO] Run t-SNE successfully.
[2025-04-15 03:51:03] [3991792] [INFO] ANALYSIS DONE: {"obsm": "a6fd9bb9b8d345bcb96b24a1ca6c91e3"}.
[2025-04-15 03:51:03] [3991792] [INFO] DONE.
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Louvain clustering - resolution=0.1', 'Louvain clustering - resolution=0.5', 'Louvain clustering - resolution=1', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - tSNE', 'Spatial Cell centers', 'UMAP - n_neighbors=15', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 't-SNE - perplexity=30'
1.9.2. Clustering
analysis.tl.louvain(adata, embedding_key="Connector - PCA", resolution=0.2, title="Connector - Louvain")
adata
[2025-04-04 17:22:22] [3762126] [INFO] ANALYSIS JOB IS WAITING AT ID 531.
[2025-04-04 17:22:22] [3780791] [INFO] ANALYZE: Louvain
[2025-04-04 17:22:22] [3780791] [INFO] Run Louvain.
[2025-04-04 17:22:22] [3780791] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
resolution: 0.2
max_clusters: 100
n_neighbors: 90
local_connectivity: 1.0
bandwidth: 1
mix_ratio: 1
}.
[2025-04-04 17:22:22] [3780791] [INFO] Run neighbors_graph.
[2025-04-04 17:22:22] [3780791] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_neighbors: 90
local_connectivity: 1.0
bandwidth: 1
mix_ratio: 1
}.
[2025-04-04 17:22:22] [3780791] [INFO] Run nearest_neighbors.
[2025-04-04 17:22:22] [3780791] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_neighbors: 90
}.
[2025-04-04 17:22:23] [3780791] [INFO] Run nearest_neighbors successfully
[2025-04-04 17:22:24] [3780791] [INFO] Run neighbors_graph successfully.
[2025-04-04 17:22:24] [3780791] [INFO] Run Louvain successfully.
[2025-04-04 17:22:24] [3780791] [INFO] ANALYSIS DONE: {"obs": "29d2411d4a10405da31d6fe2ee6eebb9"}.
[2025-04-04 17:22:24] [3780791] [INFO] DONE
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Connector - Louvain', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - PCA (1)', 'Connector - UMAP', 'Connector - tSNE', 'Spatial Cell centers', 'counts_per_pixel', 'spatial'
analysis.tl.leiden(adata, embedding_key="Connector - PCA", resolution=0.2, title="Connector - Leiden")
adata
[2025-04-04 17:22:45] [3762108] [INFO] ANALYSIS JOB IS WAITING AT ID 532.
[2025-04-04 17:22:45] [3780791] [INFO] ANALYZE: Leiden
[2025-04-04 17:22:45] [3780791] [INFO] Run Leiden.
[2025-04-04 17:22:45] [3780791] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
resolution: 0.2
max_clusters: 100
n_neighbors: 90
local_connectivity: 1.0
bandwidth: 1
mix_ratio: 1
}.
[2025-04-04 17:22:45] [3780791] [INFO] Run neighbors_graph.
[2025-04-04 17:22:45] [3780791] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_neighbors: 90
local_connectivity: 1.0
bandwidth: 1
mix_ratio: 1
}.
[2025-04-04 17:22:45] [3780791] [INFO] Found analytic results for neighbors_graph with same parameters.
[2025-04-04 17:22:46] [3780791] [INFO] Run Leiden successfully.
[2025-04-04 17:22:46] [3780791] [INFO] ANALYSIS DONE: {"obs": "4a3d5b92bd4c4129848983e8a3770570"}.
[2025-04-04 17:22:46] [3780791] [INFO] DONE
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Connector - Leiden', 'Connector - Louvain', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - PCA (1)', 'Connector - UMAP', 'Connector - tSNE', 'Spatial Cell centers', 'counts_per_pixel', 'spatial'
analysis.tl.kmeans(adata, embedding_key="Connector - PCA", n_clusters=8, title="Connector - kmeans")
adata
[2025-04-04 17:23:19] [3762126] [INFO] ANALYSIS JOB IS WAITING AT ID 533.
[2025-04-04 17:23:19] [3780787] [INFO] ANALYZE: k-means
[2025-04-04 17:23:19] [3780787] [INFO] Run k-means.
[2025-04-04 17:23:19] [3780787] [INFO] Parameters: {
embedding_key: 65e59ec140644952861ee2fcdcb069ca
is_global: True
n_clusters: 8
}.
[2025-04-04 17:23:19] [3780787] [INFO] Run k-means successfully.
[2025-04-04 17:23:19] [3780787] [INFO] ANALYSIS DONE: {"obs": "88bbe06c086e46ddb0870fbc1deaf243"}.
[2025-04-04 17:23:19] [3780787] [INFO] DONE
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Connector - Leiden', 'Connector - Louvain', 'Connector - kmeans', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - PCA (1)', 'Connector - UMAP', 'Connector - tSNE', 'Spatial Cell centers', 'counts_per_pixel', 'spatial'
1.9.3. Cell Type Prediction
analysis.tl.metadata_reference(
adata,
cluster_key="Connector - Leiden",
species=Species.HUMAN.value,
annotation_type="major",
title="Connector - Cell Types",
)
adata
[2025-04-04 17:25:35] [3762108] [INFO] ANALYSIS JOB IS WAITING AT ID 534.
[2025-04-04 17:25:35] [3780787] [INFO] ANALYZE: celltype_prediction
[2025-04-04 17:25:35] [3780787] [INFO] Run celltype_prediction.
[2025-04-04 17:25:35] [3780787] [INFO] Metareference Version: 0.1.4
[2025-04-04 17:25:35] [3780787] [INFO] Metareference Server: https://talk2data.bioturing.com/meta_reference
[2025-04-04 17:25:35] [3780787] [INFO] Parameters: {
cluster_key: Connector - Leiden
anno_type: major
custom_gene_sets: {}
species: human
log2fc_threshold: 0.5
pct_diff_threshold: 0.1
name: Connector - Cell Types
}.
[2025-04-04 17:25:35] [3780787] [INFO] Computing marker genes for clusters...
[2025-04-04 17:25:36] [3780787] [INFO] Predicting cell types using MetaReference..
[2025-04-04 17:25:42] [3780787] [INFO] Preparing output...
[2025-04-04 17:25:42] [3780787] [INFO] Writing output files..
[2025-04-04 17:25:44] [3780787] [INFO] Writing metadata...
[2025-04-04 17:25:44] [3780787] [INFO] Run celltype_prediction successfully.
[2025-04-04 17:25:44] [3780787] [INFO] ANALYSIS DONE: {"obs": "18ebec09cd7f4877934a202640828e5b", "analysis": "celltype_prediction.zarr"}.
[2025-04-04 17:25:44] [3780787] [INFO] DONE
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'Connector - Cell Types', 'Connector - Leiden', 'Connector - Louvain', 'Connector - kmeans', 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Connector - PCA', 'Connector - PCA (1)', 'Connector - UMAP', 'Connector - tSNE', 'Spatial Cell centers', 'counts_per_pixel', 'spatial'
1.9.4. Differential Expression
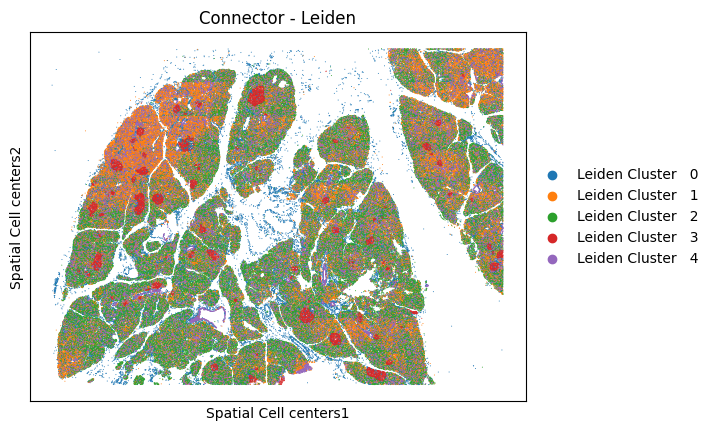
analysis.pl.embedding(adata, "Spatial Cell centers", color="Connector - Leiden", s=2)
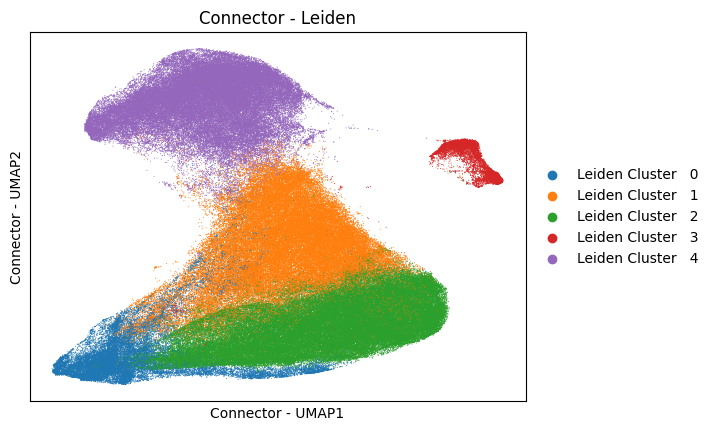
analysis.pl.embedding(adata, "Connector - UMAP", color="Connector - Leiden", s=2)
[2025-04-04 17:34:36] [3788266] [WARNING] [SKIP] Not support `_sanitize` function.
[2025-04-04 17:34:37] [3788266] [WARNING] [SKIP] Not support `_sanitize` function.
adata.obs["Connector - Leiden"].value_counts()
Connector - Leiden
Leiden Cluster 2 93573
Leiden Cluster 1 71343
Leiden Cluster 4 70129
Leiden Cluster 0 25493
Leiden Cluster 3 9359
Name: count, dtype: int64
groups, refs = sorted(list(adata.obs["Connector - Leiden"].value_counts().index))[-2:]
groups, refs
('Leiden Cluster 3', 'Leiden Cluster 4')
analysis.tl.rank_genes_groups(adata, groupby="Connector - Leiden", groups=groups, reference=refs)
[2025-04-04 17:44:23] [3762126] [INFO] ANALYSIS JOB IS WAITING AT ID 535.
[2025-04-04 17:44:23] [3780791] [INFO] ANALYZE: de_genes
[2025-04-04 17:44:23] [3780791] [INFO] Run de_genes.
[2025-04-04 17:44:23] [3780791] [INFO] Parameters: {
method: venice
sample_id_1: SP-01JR0TTWZ278NPYQXWPGCA3CX1
sample_id_2: SP-01JR0TTWZ278NPYQXWPGCA3CX1
table_id_1: 837faa22893e4df2a83e2e1885955e0f
table_id_2: 837faa22893e4df2a83e2e1885955e0f
n_cells_1: 9359
n_cells_2: 70129
group_1_name: ['Leiden Cluster 3']
group_2_name: ['Leiden Cluster 4']
include: []
exclude: []
coverage: 10
normalize_method: log1p-normalized
}
[2025-04-04 17:44:24] [3780791] [INFO] Run de_genes successfully.
[2025-04-04 17:44:24] [3780791] [INFO] ANALYSIS DONE: {"analysis": "de_genes.msgpack.gz"}.
[2025-04-04 17:44:24] [3780791] [INFO] DONE
1.9.5. Spatial Analysis - Region Segmentation
mpp = data.spatialdata.images.read_attrs("human_pancreas_codex_protein_images")[ConnectorKeys.MPP.value]
mpp
1
analysis.tl.region_segmentation(adata, radius=50, mpp=mpp, species=Species.HUMAN.value)
1.9.6. Download data to local to run analysis and upload to server
import pandas as pd
import scanpy as sc
memory_adata = adata.to_memory()
# Or slicing data with obs and var first: memory_adata = adata[obs["obs field"], var["var field"]].copy()
memory_adata
[2025-04-08 10:03:06] [2758802] [WARNING] group not found at path None
[2025-04-08 10:03:06] [2758802] [WARNING] Create `layers` in memory.
[2025-04-08 10:03:06] [2758802] [WARNING] Create `uns` in memory.
[2025-04-08 10:03:06] [2758802] [WARNING] group not found at path None
[2025-04-08 10:03:06] [2758802] [WARNING] group not found at path None
AnnData object with n_obs × n_vars = 269897 × 12
obs: 'area'
var: 'Top 10 Highly Variable Genes'
obsm: 'Spatial Cell centers', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial'
sc.pp.normalize_total(memory_adata)
sc.pp.log1p(memory_adata)
sc.pp.pca(memory_adata, 10)
memory_adata
WARNING: adata.X seems to be already log-transformed.
AnnData object with n_obs × n_vars = 269897 × 12
obs: 'area'
var: 'Top 10 Highly Variable Genes'
uns: 'log1p', 'pca'
obsm: 'Spatial Cell centers', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 'X_pca'
varm: 'PCs'
sc.pp.neighbors(memory_adata)
sc.tl.umap(memory_adata, init_pos="random")
memory_adata
AnnData object with n_obs × n_vars = 269897 × 12
obs: 'area'
var: 'Top 10 Highly Variable Genes'
uns: 'log1p', 'pca', 'neighbors', 'umap'
obsm: 'Spatial Cell centers', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 'X_pca', 'X_umap'
varm: 'PCs'
obsp: 'distances', 'connectivities'
sc.tl.louvain(memory_adata, 0.1, key_added="louvain_0.1")
sc.tl.louvain(memory_adata, 0.5, key_added="louvain_0.5")
memory_adata
AnnData object with n_obs × n_vars = 269897 × 12
obs: 'area', 'louvain_0.1', 'louvain_0.5'
var: 'Top 10 Highly Variable Genes'
uns: 'log1p', 'pca', 'neighbors', 'umap', 'louvain_0.1', 'louvain_0.5'
obsm: 'Spatial Cell centers', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial', 'X_pca', 'X_umap'
varm: 'PCs'
obsp: 'distances', 'connectivities'
adata.obsm["X_pca"] = memory_adata.obsm["X_pca"]
adata.obsm["X_umap"] = memory_adata.obsm["X_umap"]
adata.upload_embeddings(["X_pca", "X_umap"])
[2025-04-08 10:30:56] [2730684] [INFO] ANALYSIS JOB IS WAITING AT ID 569.
[2025-04-08 10:30:56] [2730024] [INFO] ANALYZE: import_embeddings.
[2025-04-08 10:30:57] [2730024] [INFO] import_anndata_embeddings
[2025-04-08 10:30:57] [2730024] [INFO] Run import_anndata_embeddings successfully.
[2025-04-08 10:30:57] [2730024] [INFO] ANALYSIS DONE: {}.
[2025-04-08 10:30:57] [2730024] [INFO] DONE.
adata.obs["louvain_0.1"] = pd.Categorical([f"Louvain {i}" for i in memory_adata.obs["louvain_0.1"]])
adata.obs["louvain_0.5"] = pd.Categorical([f"Louvain {i}" for i in memory_adata.obs["louvain_0.5"]])
adata.upload_metadata(adata.obs[["louvain_0.1", "louvain_0.5"]]) # Or: adata.upload_metadata(["louvain_0.1", "louvain_0.5"])
adata.update()
adata
ConnectorAnnData object with n_obs × n_vars = 269897 × 12
obs: 'area', 'louvain_0.1', 'louvain_0.5'
var: 'Top 10 Highly Variable Genes'
obsm: 'Spatial Cell centers', 'X_pca', 'X_umap', 'counts_per_pixel', 'scVI - 20 latents - 10 top genes', 'spatial'
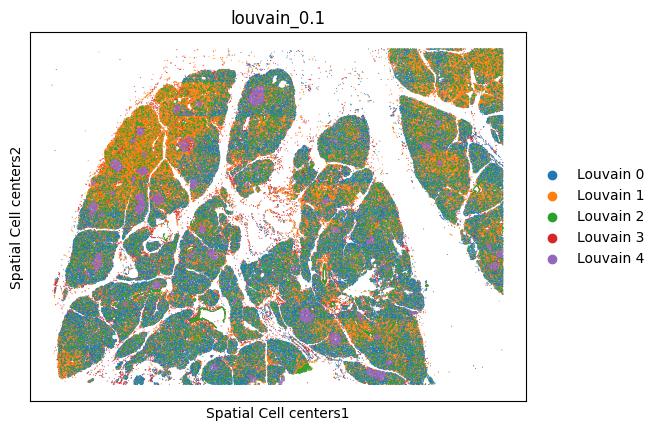
analysis.pl.embedding(adata, "Spatial Cell centers", color="louvain_0.1", s=2)
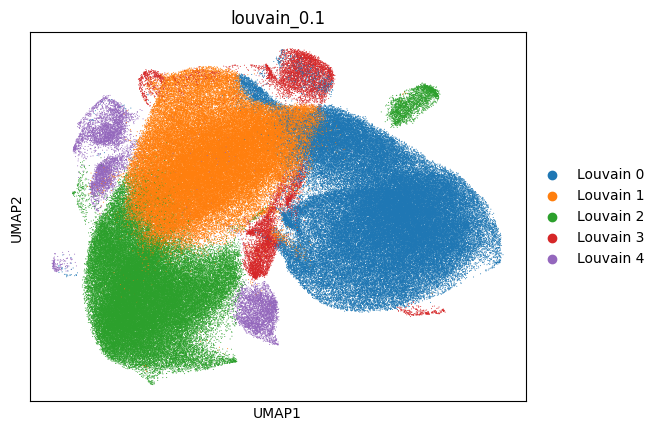
analysis.pl.umap(adata, color="louvain_0.1", s=2)
[2025-04-08 10:42:17] [2758802] [WARNING] [SKIP] Not support `_sanitize` function.
[2025-04-08 10:42:17] [2758802] [WARNING] [SKIP] Not support `_sanitize` function.
1.10. Convert Data from Lens
This section guides existing BioTuring Lens users on how to migrate their data to SpatialX.
1.10.1. Install BioTuring Lens Connector
Before proceeding, ensure you have installed the BioTuring Lens connector in addition to the SpatialX connector.
!pip install bioturing_connector
1.10.2. Input Domain and Token
To obtain your domain URL and personal token, navigate to "BioTuring Lens SDK" in the left panel of your BioTuring Lens interface. Then, enter the information in the fields below. For example:
DOMAIN = "https://example.bioturing.com/lens_sc/"
TOKEN = "000000000000000000000000NM"
- Example of BioTuring Lens SC (Single cell)
LENS_SC_HOST: str = ""
LENS_SC_TOKEN: str = ""
lens_sc_studies = connector.list_lens_sc_studies(
host=LENS_SC_HOST, token=LENS_SC_TOKEN,
group=DefaultGroup.PERSONAL_WORKSPACE,
species=Species.HUMAN.value,
)
spatialx_connector.format_print(lens_sc_studies)
# Convert a study
connector.convert_data_from_lens(lens_sc_studies[0])
- Example of BioTuring Lens Bulk
LENS_BULK_HOST: str = ""
LENS_BULK_TOKEN: str = ""
lens_bulk_studies = connector.list_lens_bulk_studies(
host=LENS_BULK_HOST, token=LENS_BULK_TOKEN,
group=DefaultGroup.PERSONAL_WORKSPACE,
species=Species.HUMAN.value,
)
spatialx_connector.format_print(lens_bulk_studies)
# Convert multiple studies
connector.convert_data_from_lens(lens_bulk_studies)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
File details
Details for the file spatialx_connector-0.2.12.tar.gz.
File metadata
- Download URL: spatialx_connector-0.2.12.tar.gz
- Upload date:
- Size: 130.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a9fc61cf5eb472d66b784880fc43938423aa088b33721e74782a081034f27653
|
|
| MD5 |
7643cc7465f207372957b3c462df068d
|
|
| BLAKE2b-256 |
02c3fd3e88eb8d43bcc1baa6c06c8d3a43f6ca4d1e01e0b6e948c3f8209d3121
|