SPECTROView: A Tool for Spectroscopic Data Processing and Visualization
Project description

SPECTROview : A Tool for Spectroscopic Data Processing and Visualization.
Spectroscopy techniques (such as Raman, Photoluminescences, XRD, XPS…) are widely used in various fields, including materials science, chemistry, biology, and geology. In recent years, these techniques have increasingly found their place in cleanroom environments, particularly within the microelectronics industry, where they serve as critical metrology tools for wafer-scale measurements. The data collected from these in-line measurements (wafer data) require specific processing, but existing software solutions are often not optimized for this type of data and typically lack advanced plotting and visualization capabilities. Additionally, the licensing requirements of these software solutions can restrict access for a broader community of users.
SPECTROview addresses these gap by offering free, open-source software that is compatible with both in-line data (wafer-map) as well as standard spectroscopic data (discret spectra, 2D maps). It also features a built-in visualization tool, enabling users to streamline both data processing and visualization in a single application, making the workflow more efficient.
Features:
- Cross-platform compatibility (Windows, macOS, Linux).
- Supports processing of spectral data (1D) and hyperspectral data (2D maps or wafer maps)*.
- Ability to fit multiple spectra or 2Dmaps using predefined models or by creating custom fit models*.
- Collect all best-fit results with one click.
- Optimized user inferface for easy and quick inspection and comparison of spectra.
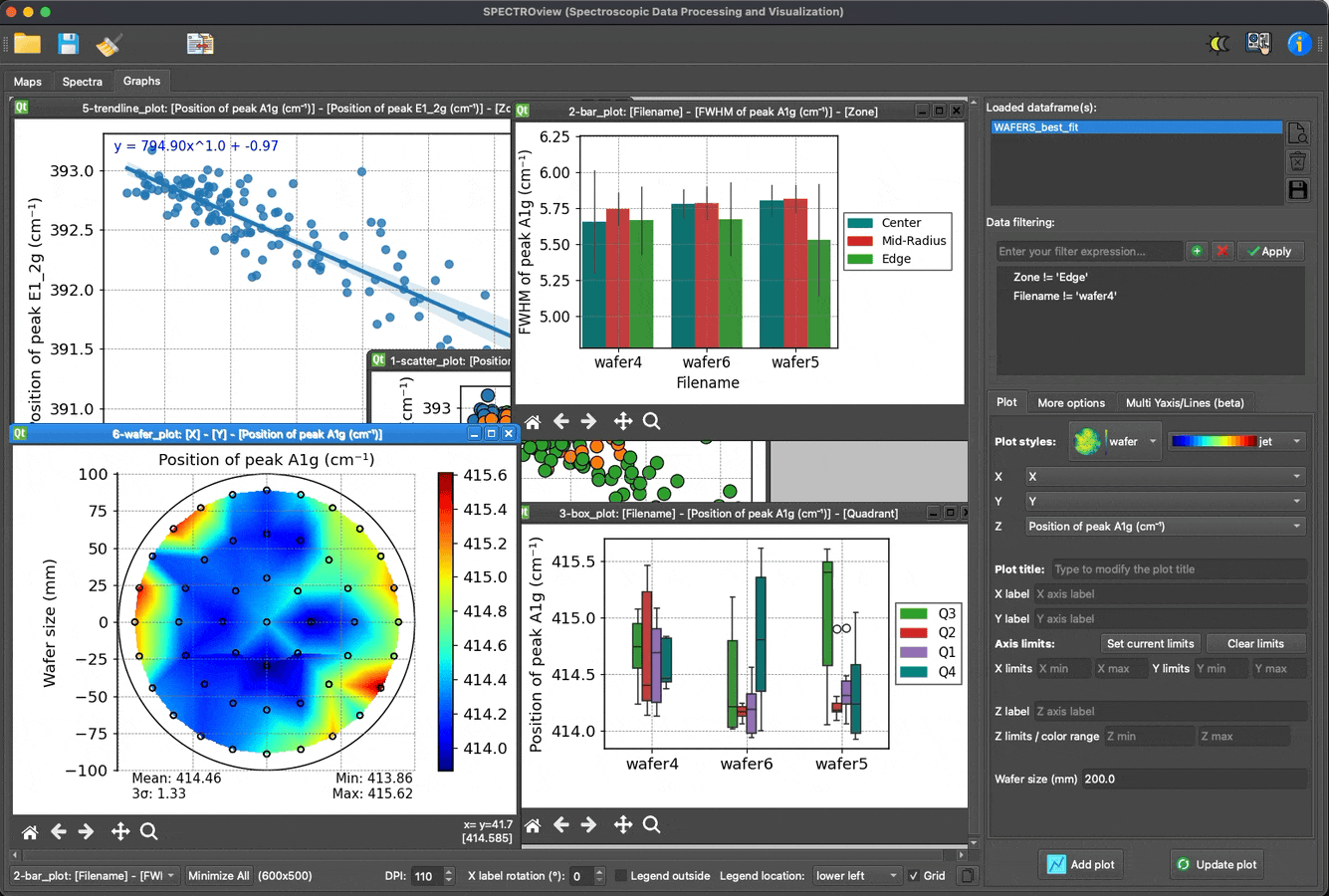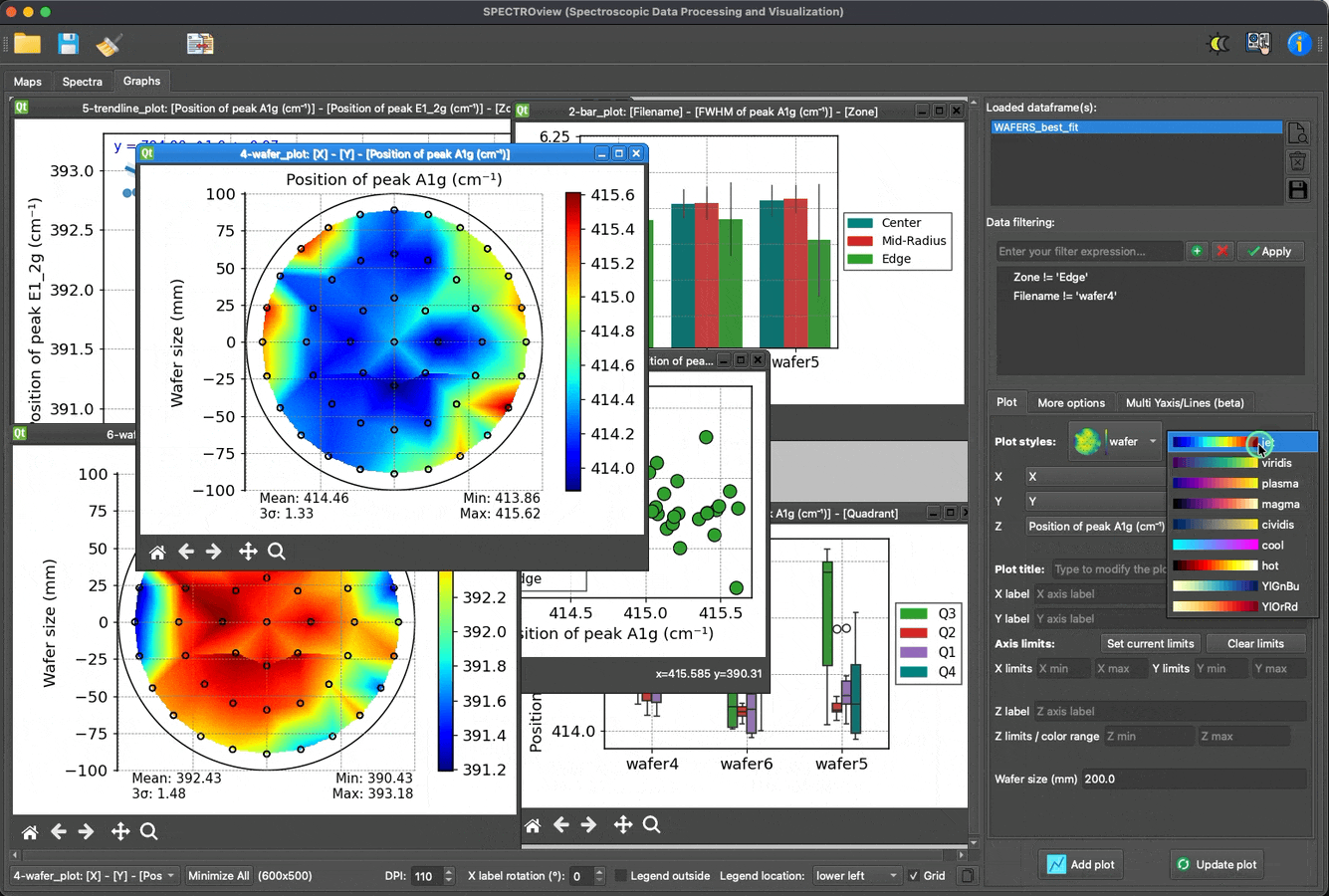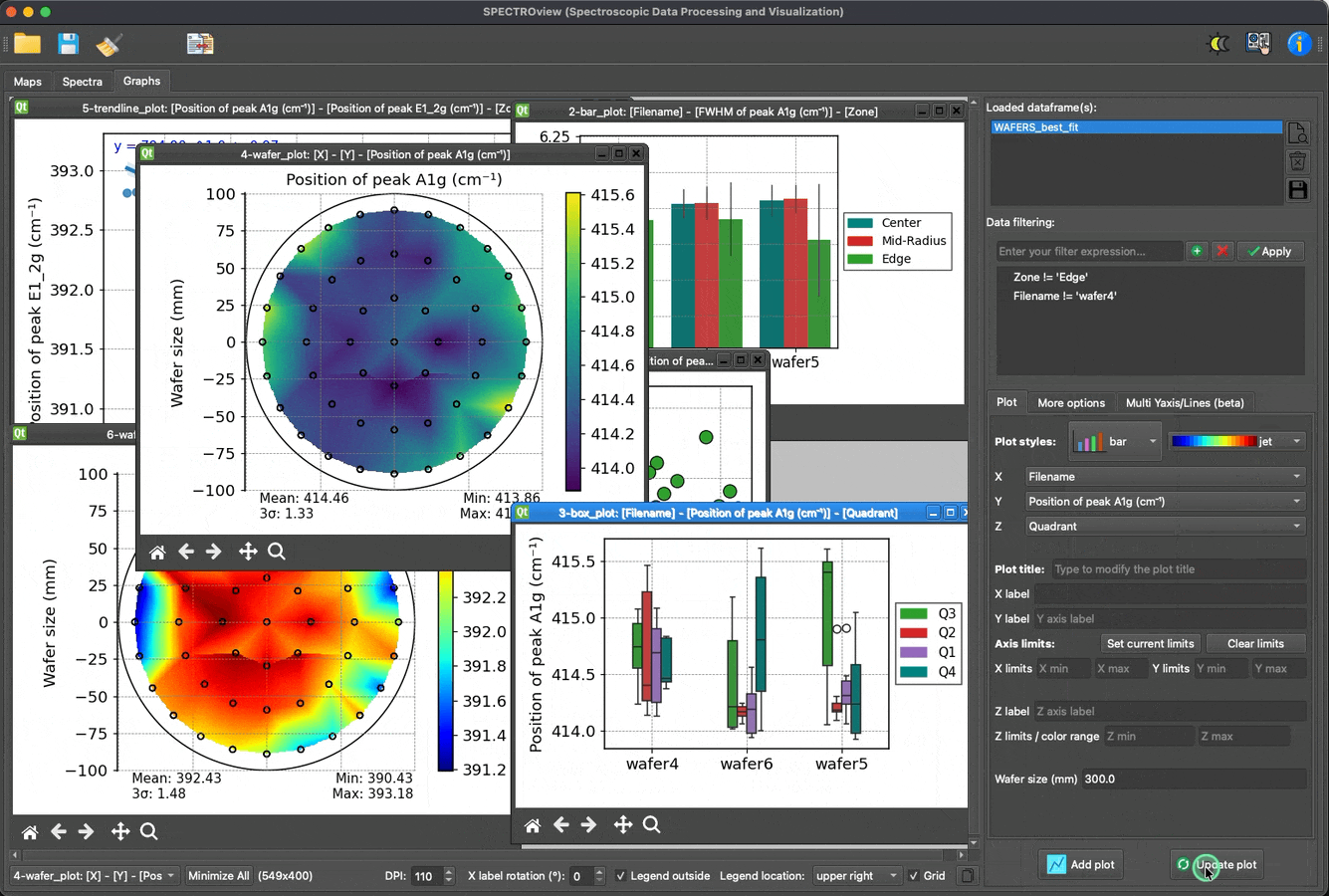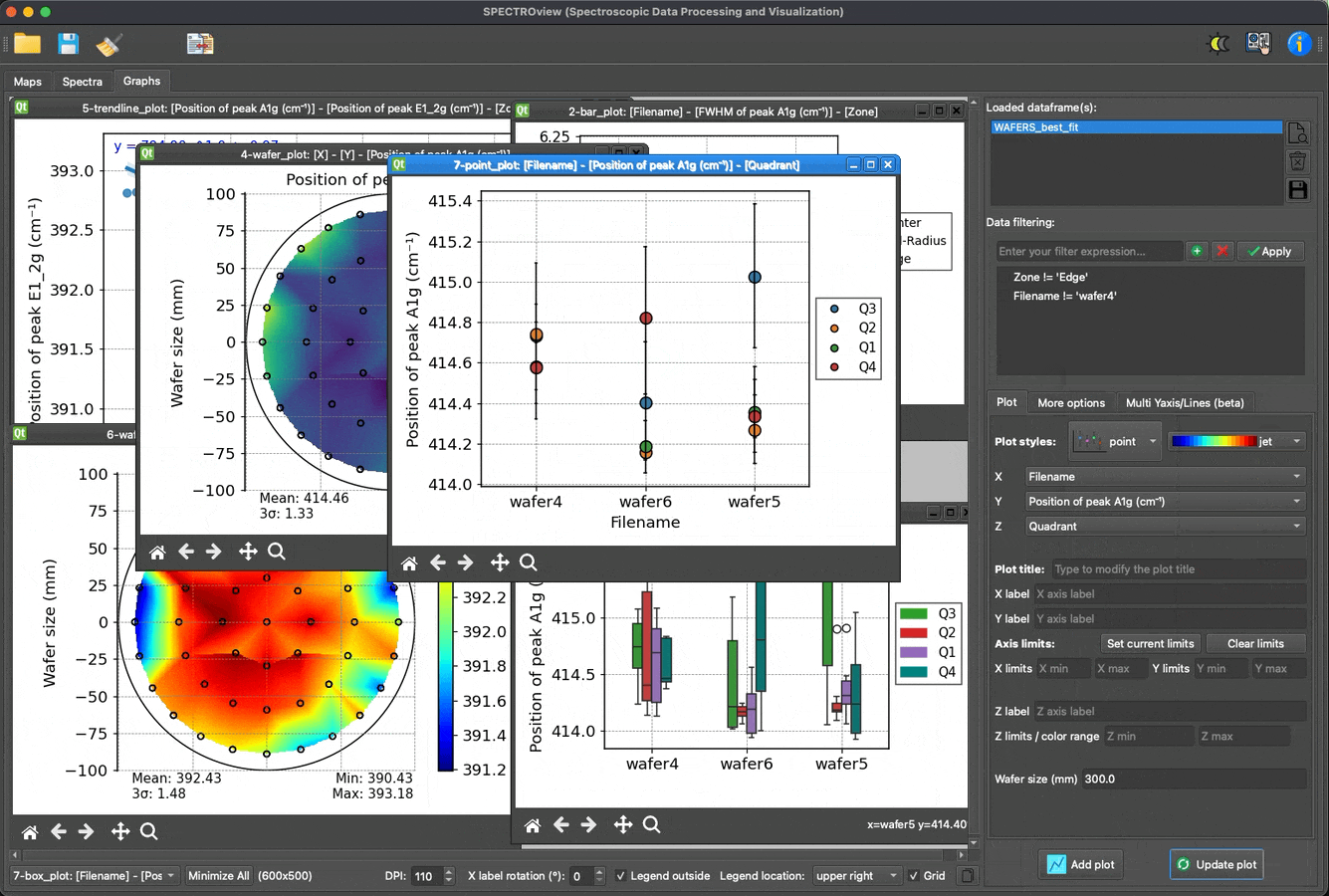
- Dedicated module for effortless, fast, and easy data visualization.
*Fitting features are powered by the fitspy and lmfit open-source packages.
Three separate tabs for processing discrete spectra, hyperspectral data, and data visualization:

Fit a spectrum; create/save and apply a fit models; collect fitted data with single click:

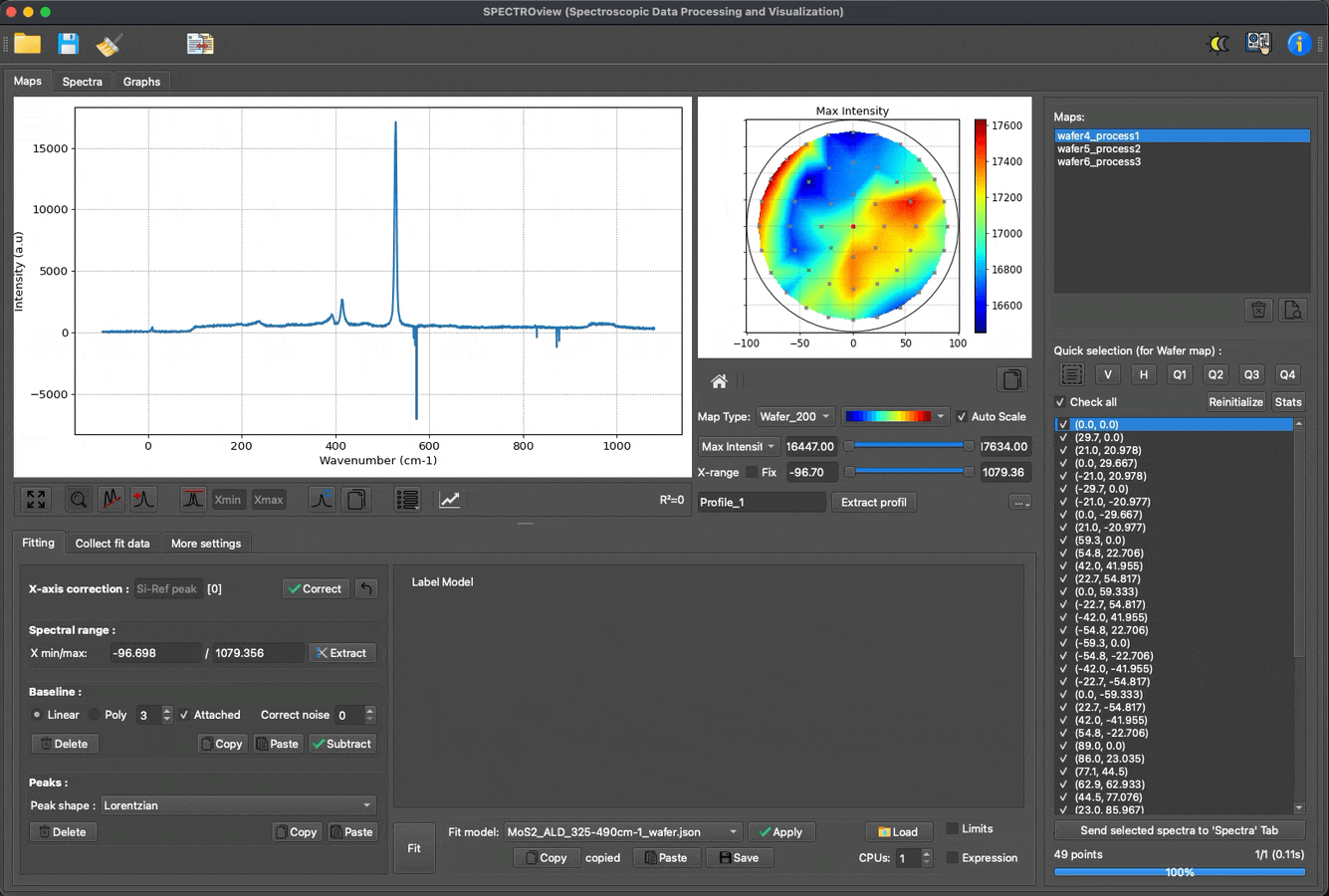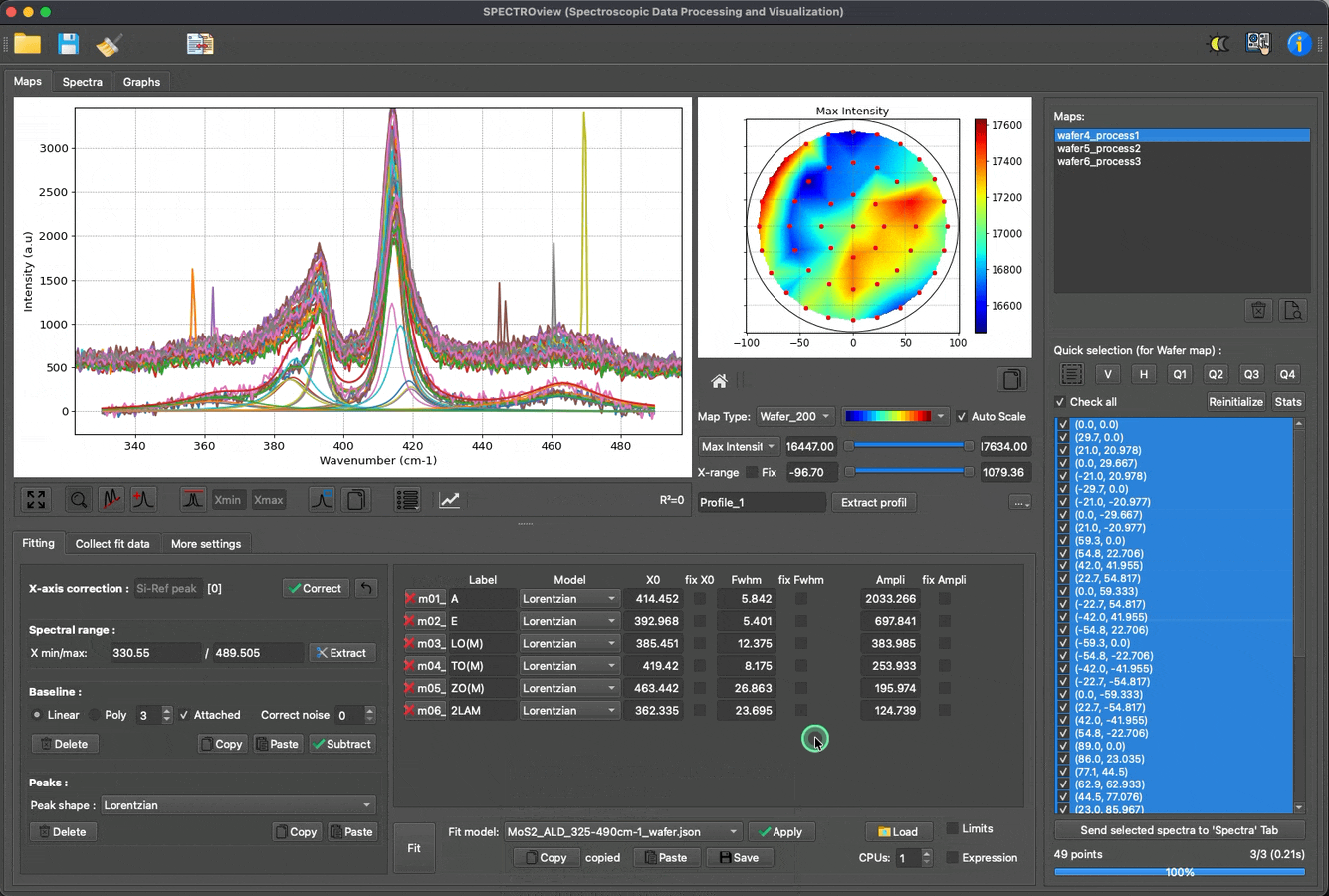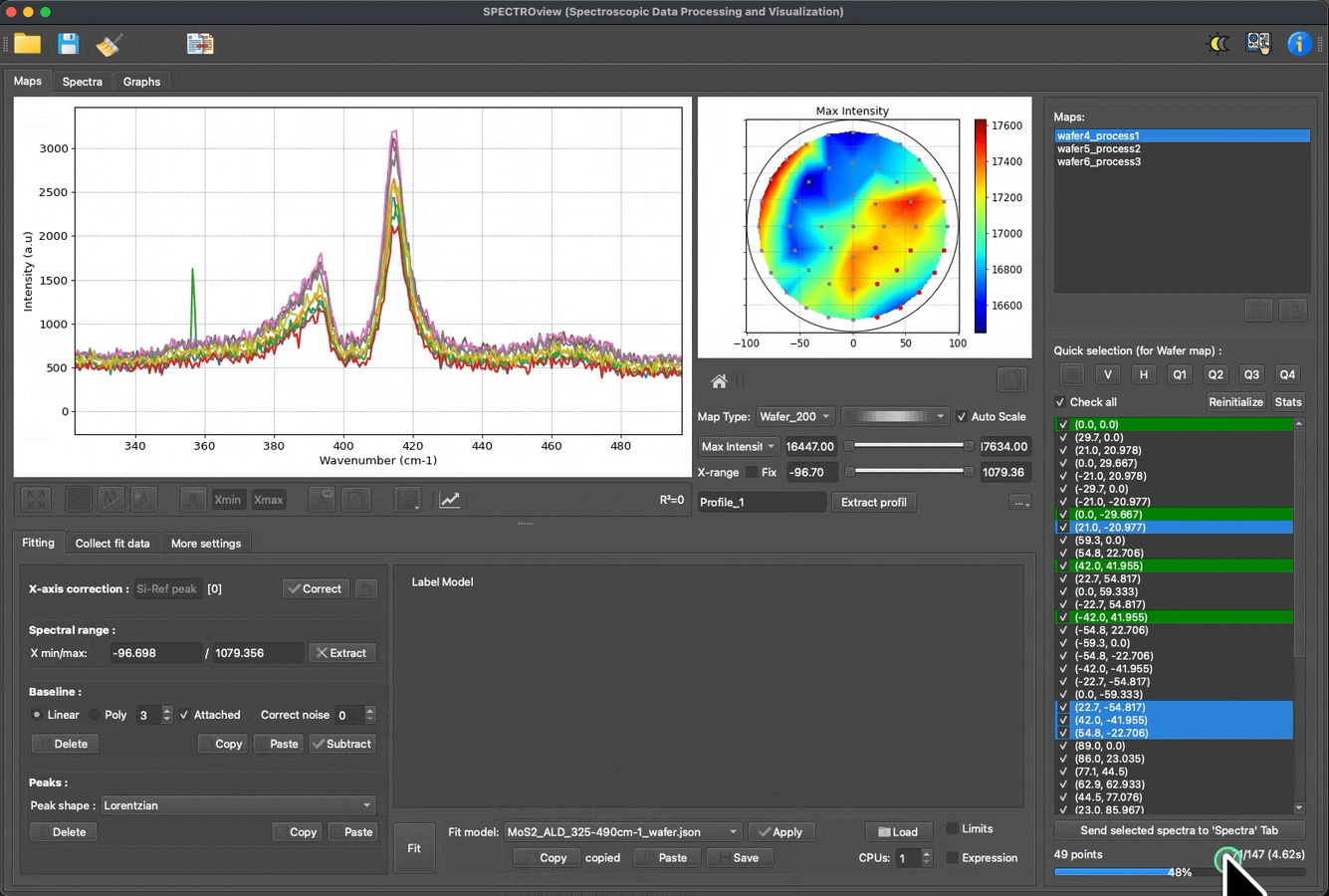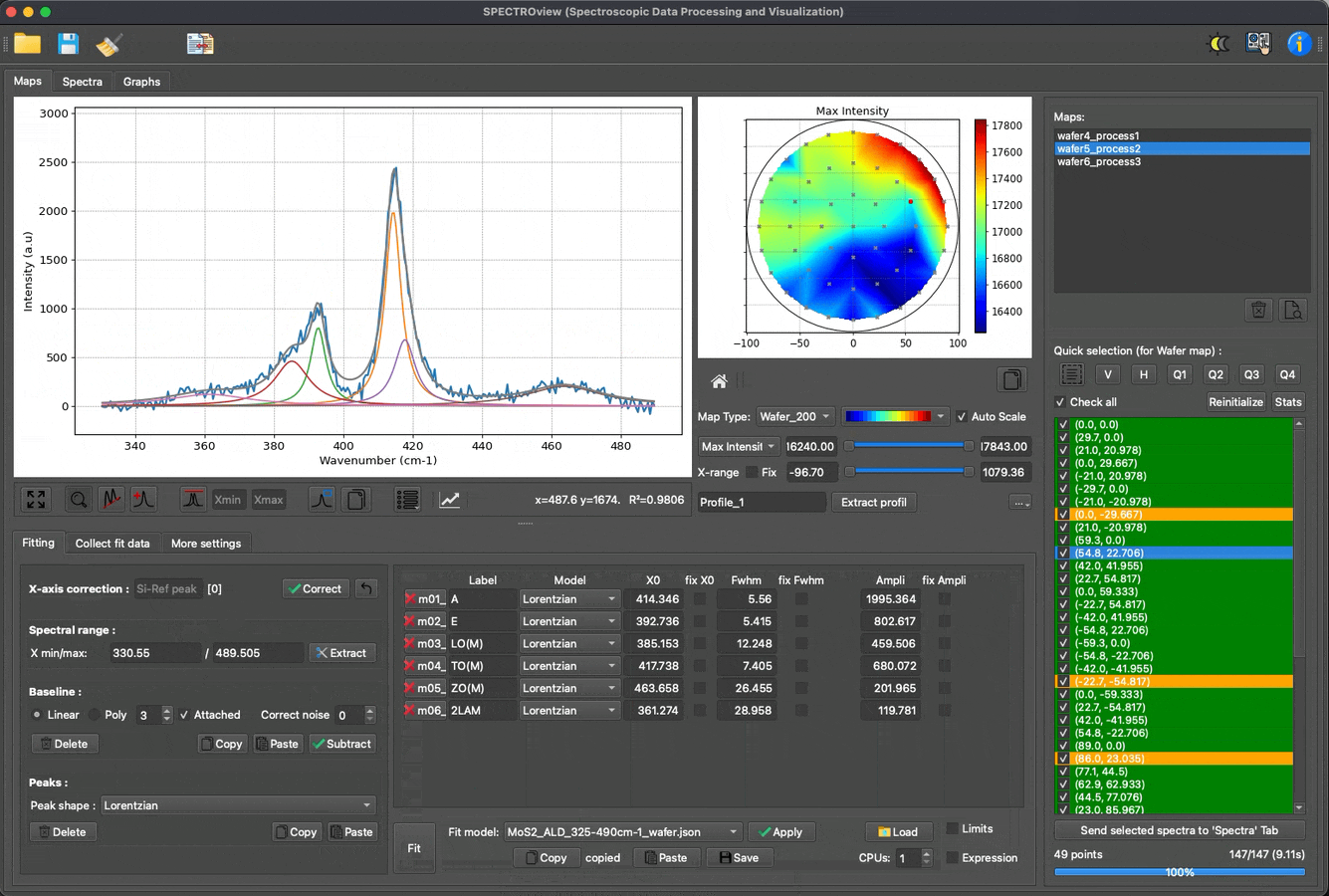
Fit multiple spectra, wafer data, and 2D maps with predefined models:
Plot and visualize data with ease:
Installation from PyPI:
Make sure that Python (> 3.8) is already installed.
pip install spectroview
Installation from Github:
pip install git+https://github.com/CEA-MetroCarac/SPECTROview.git
To launch SPECTROview:
spectroview
Acknowledgements
This work, carried out on the CEA - Platform for Nanocharacterisation (PFNC), was supported by the “Recherche Technologique de Base” program of the French National Research Agency (ANR).
Citation
Le, V.-H., & Quéméré, P. (2025). SPECTROview : A Tool for Spectroscopic Data Processing and Visualization. Zenodo. https://doi.org/10.5281/zenodo.14147172
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file spectroview-26.3.1.tar.gz.
File metadata
- Download URL: spectroview-26.3.1.tar.gz
- Upload date:
- Size: 5.6 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
63be7d5a78685f54a5f3526a60d56eccbaec7ec56b1c1984e415660b8a230b41
|
|
| MD5 |
0c241e68175f288c26d443770ae7ba4d
|
|
| BLAKE2b-256 |
405340af9fbf1511617ff5ac2d9f628547555c39d204d0952f21ace22639e489
|
File details
Details for the file spectroview-26.3.1-py3-none-any.whl.
File metadata
- Download URL: spectroview-26.3.1-py3-none-any.whl
- Upload date:
- Size: 5.6 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b00fb42269f89a227f2af41463af735b81e167aeba000472d79fa0274d24621e
|
|
| MD5 |
2ad69cf744d09139ae692ce3a5f5017c
|
|
| BLAKE2b-256 |
a3bab8c999cdfbb017fb650e11bbc85693fcfa4051a591d675d84e0bff625d27
|















