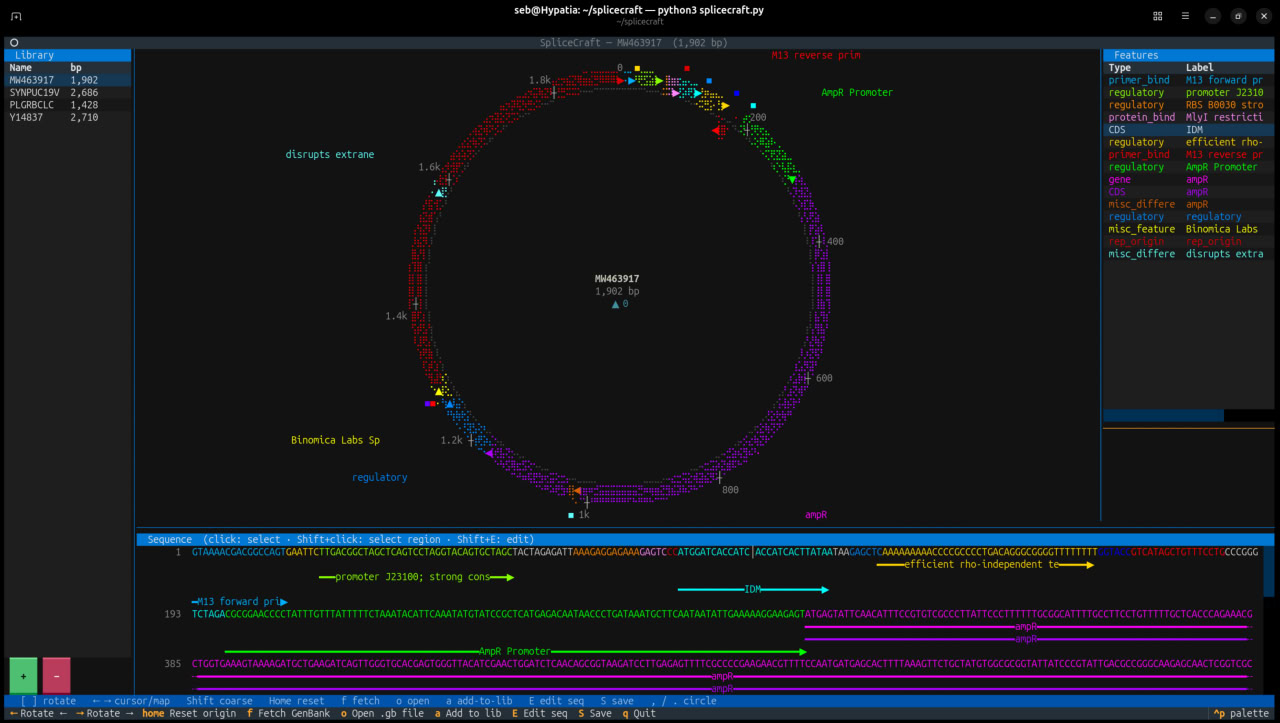
Terminal-based circular plasmid map viewer, sequence editor, and Primer3/Golden Braid primer design workbench
Project description
SpliceCraft
╔═══════════════════════════════════════════════════════════════════════════════╗
║ ⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶ ║
║ ║
║ ________ __________ _________ ____________ ║
║ __ ___/__________ /__(_)____________ ____/____________ ___ __/_ /_ ║
║ _____ \___ __ \_ /__ /_ ___/ _ \ / __ ___/ __ `/_ /_ _ __/ ║
║ ____/ /__ /_/ / / _ / / /__ / __/ /___ _ / / /_/ /_ __/ / /_ ║
║ /____/ _ .___//_/ /_/ \___/ \___/\____/ /_/ \__,_/ /_/ \__/ ║
║ /_/ ║
║ ║
║ · I n - T e r m i n a l P l a s m i d W o r k b e n c h · ║
║ ║
║ ⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶⠶ ║
╚═══════════════════════════════════════════════════════════════════════════════╝
A terminal-based circular plasmid map viewer, sequence editor, primer design workbench, and Golden Braid parts domesticator — rendered entirely in your shell. Fetch any GenBank record by accession, load local files, annotate features with pLannotate, design diagnostic / cloning / Golden Braid primers with Primer3, and edit sequences — without ever leaving the terminal.
Features
Core visualization & editing
- Braille dot-matrix circular map — plasmids rendered as crisp Unicode braille rings with per-strand feature arcs, directional arrowheads, and proximity-placed labels
- Linear map view — toggle with
vfor a horizontal strip layout - Dithered sequence panel — per-base DNA viewer with feature bars, restriction site overlays, and double-stranded display
- Live NCBI fetch — pull any GenBank record by accession number on demand
- Local file support — open
.gb/.gbkfiles directly from disk - Free rotation — spin the origin left or right with
[/] - Restriction enzyme overlay — 200+ NEB enzymes including Type IIS (BsaI, BsmBI, BbsI, SapI) with visible recognition arcs + cut markers
Libraries (all persist to JSON)
- Plasmid library — SnapGene-style collection, auto-saves on import, survives restarts, supports rename and handslip-protected delete
- Parts Bin — Golden Braid L0 parts catalog with user-domesticated parts including sequences and primer pairs
- Primer library — all designed primers with Tm, length, date, status (Designed / Ordered / Validated), multi-select for batch operations
Primer design (Primer3)
- Detection primers — diagnostic PCR; Primer3 picks the ideal pair within a selected region, 450-550 bp product by default (configurable)
- Cloning primers — RE-site tails + GCGC padding; 30+ common enzymes or type a custom recognition sequence
- Golden Braid primers — BsaI domestication for all L0 positions (Promoter, 5' UTR, CDS, CDS-NS, C-tag, Terminator)
- Generic primers — simple binding primers, no tails
- Primers can be added to the plasmid map as
primer_bindfeatures - Scrollable
TextAreafor custom sequence input; highlighted text = target
Annotation
- pLannotate integration — press
Shift+A(or use the◈library button) to auto-annotate a plasmid against pLannotate's curated feature database. Optional — see install notes below
Feature operations
- Feature sidebar — click a row to highlight on map; click the map to select the feature under the cursor
- Undo/redo — 50-deep snapshot stack for all sequence edits
- Delete protection — focus-aware Delete key; confirmation modal (default focus = No) for library entries
- Clipboard — OSC-52 copy works in Windows Terminal, iTerm2, modern WSL
Data safety
- Atomic saves — all JSON files written via tempfile +
os.replace - Automatic backups — every save writes
*.json.bakbefore overwriting - Corrupt-file recovery — missing files don't crash; corrupt files auto-
restore from
.bakwith a warning notification on startup
Installation
Requires Python 3.10+.
From PyPI (recommended)
pip install splicecraft
splicecraft # empty canvas
splicecraft L09137 # fetch pUC19 from NCBI
splicecraft myplasmid.gb # open a local GenBank file
All required dependencies (textual, biopython, primer3-py, platformdirs)
are pulled in automatically. User data (library, parts bin, primers) lives in
the platform-appropriate data directory:
| Platform | Path |
|---|---|
| Linux | ~/.local/share/splicecraft/ |
| macOS | ~/Library/Application Support/splicecraft/ |
| Windows | %APPDATA%\splicecraft\ |
Override with SPLICECRAFT_DATA_DIR=/path/to/dir splicecraft.
From source
git clone https://github.com/Binomica-Labs/SpliceCraft.git
cd SpliceCraft
pip install -e .
Optional dependencies
pLannotate (for automatic plasmid annotation via Shift+A) — requires conda:
conda create -n plannotate -c conda-forge -c bioconda plannotate
conda activate plannotate
plannotate setupdb # one-time ~500 MB BLAST database download
# then run SpliceCraft from the same conda env
SpliceCraft runs fine without pLannotate — the annotation feature just notifies the user how to install it if pressed.
Usage
# After pip install:
splicecraft # empty canvas
splicecraft L09137 # fetch pUC19 from NCBI on launch
splicecraft myplasmid.gb # open a local GenBank file
splicecraft --version # print version
splicecraft --help # quick usage hint
If running from a git clone (pip install -e .), the same commands work;
you can also still run python3 splicecraft.py directly.
Key Bindings
Main screen
| Key | Description |
|---|---|
[ / ] |
Rotate map origin left / right |
Shift+[/] |
Rotate coarse (10× step) |
Home |
Reset origin to 0 |
, / . |
Circular map aspect wider / taller |
v |
Toggle circular ↔ linear map |
l |
Toggle feature label connector lines |
r |
Toggle restriction-site overlay |
f |
Fetch a record from NCBI by accession |
o |
Open a .gb file from disk |
a |
Add current plasmid to the library |
Shift+A |
Annotate plasmid with pLannotate |
Shift+E |
Enter sequence editor mode |
Shift+S |
Save edits to file |
Delete |
Context-aware delete (feature or library entry) |
Ctrl+Z |
Undo |
Ctrl+Shift+Z |
Redo |
Ctrl+C |
Copy selection to clipboard |
q |
Quit |
Primer Design screen
| Key | Description |
|---|---|
esc |
Close primer screen |
m |
Mark / unmark primer under cursor (★) |
M |
Mark / unmark all primers |
S |
Cycle status: Designed → Ordered → Validated |
Tab |
Cycle focus between fields |
Mouse
| Action | Description |
|---|---|
| Click | Place cursor / select feature under pointer |
| Double-click | Select full feature span |
| Drag | Select a sequence range |
| Scroll wheel | Rotate map (when over map panel) |
Menus
| Menu | Items |
|---|---|
| File | Open .gb file · Fetch from NCBI · Add to Library · Save · Quit |
| Edit | Edit Sequence · Undo · Redo · Delete Feature |
| Enzymes | Show RE sites · Unique cutters · 6+/4+ bp sites · Connectors |
| Features | Add Feature · Delete Feature · Annotate with pLannotate |
| Primers | Opens the full-screen Primer Design workbench |
| Parts | Opens the Parts Bin (Golden Braid L0 parts catalog) |
| Constructor | Opens the Assembly Constructor for TU building |
Requirements
| Package | Version | Purpose |
|---|---|---|
| Python | ≥ 3.10 | Runtime |
| Textual | ≥ 8.2.3 | TUI framework and rendering engine |
| Rich | ≥ 14.0 | Terminal rendering (Textual dependency) |
| Biopython | ≥ 1.87 | GenBank parsing and NCBI Entrez fetch |
| primer3-py | ≥ 2.3.0 | Primer design (Tm, thermodynamic screening) |
| pytest | ≥ 9.0 | Test suite (dev only) |
| pytest-asyncio | ≥ 1.3 | Async test support (dev only) |
| pLannotate | optional, conda | Automatic plasmid annotation (Shift+A) |
| BLAST+ | optional, conda | Required by pLannotate |
| Primer3 CLI | optional, apt |
Not used directly — primer3-py bundles it |
Data files
All user data persists as human-readable JSON in the repo directory:
| File | Purpose |
|---|---|
plasmid_library.json |
Saved plasmid collection (GenBank + metadata) |
parts_bin.json |
User-domesticated Golden Braid parts |
primers.json |
Designed primer library |
*.json.bak |
Automatic backup — written before each save |
These files are in .gitignore — they're user-local data, not repo content.
A manual backup rotation happens on every save so accidental data loss is
always recoverable via the .bak file.
Tests
python3 -m pytest -q # full suite (246 tests, ~45 s)
python3 -m pytest tests/test_dna_sanity.py # biology correctness only (< 1s)
License
MIT
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file splicecraft-0.2.0.tar.gz.
File metadata
- Download URL: splicecraft-0.2.0.tar.gz
- Upload date:
- Size: 236.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
06986a8392cad40120d14c571f1ed9447f3bec5419f02b6800a2dbe524be994f
|
|
| MD5 |
4b15022b7f5ac6c6cf01978cacccf759
|
|
| BLAKE2b-256 |
eb3c9c193d274093b6c02c347b5d45a731a7506c09f618a120729e413cab141e
|
File details
Details for the file splicecraft-0.2.0-py3-none-any.whl.
File metadata
- Download URL: splicecraft-0.2.0-py3-none-any.whl
- Upload date:
- Size: 80.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2e5ac8f97cae832df83b6712969a7ad3ec5ef0b453a46d1d76a9ffd790eb8377
|
|
| MD5 |
881c4e367fddd420415f04f8138e81ca
|
|
| BLAKE2b-256 |
e054633ff3108772f14289863fb095ac2eeffccb86d5066c5ce94a5d9f0cd392
|