No project description provided
Project description
Squidly
Squidly, is a tool that employs a biologically informed contrastive learning approach to accurately predict catalytic residues from enzyme sequences. We offer Squidly as ensembled with Blast to achieve high accuracy at low and high sequence homology settings.
If you use squidly in your work please cite our preprint: https://www.biorxiv.org/content/10.1101/2025.06.13.659624v1
Also if you have any issues installing, please post an issue! We have tested this on ubuntu.
Requirements
Squidly is dependant on the ESM2 3B or 15B protein language model. Running Suidly will automatically attempt to download each model. The Smaller 3B model is lighter, runs faster and requires less VRAM. The 3B and 15B models expect roughly 25GB and 74GB of VRAM, respectively. Our tests on 100 sequences with an average length of 400+ took about 40 seconds and 3 minutes for the 3B and 15B model.
Squidly can only predict sequences of less than 1024 residues!
Currently we expect GPU access but if you require a CPU only version please let us know and we can update this!
Installation
conda create --name squidly python=3.10
conda activate squidly
pip install squidly
squidly install
Running squidly install should automatically download all models from huggingface. Now you can run squidly (see Usage below).
Note if you get the below error:
ValueError: numpy.dtype size changed, may indicate binary incompatibility. Expected 96 from C header, got 88 from PyObject
you may need to update numpy and pandas.
Usage
By default squidly runs the ensembled squidly model. It's optimised to run as fast as the single model.
For example to run the 3B model with a fasta file
squidly run example.fasta esm2_t36_3B_UR50D
Squidly can also be further ensembled with BLAST (you need to pass the database as well)
squidly run example.fasta esm2_t36_3B_UR50D output_folder/ --database reviewed_sprot_08042025.csv
Where reviewed_sprot_08042025.csv is the example database (i.e. a csv file with the following columns)
You can see ours which is zipped in the data folder..
| Entry | Sequence | Residue |
|---|---|---|
| A0A009IHW8 | MSLEQKKGADIIS | 207 |
| A0A023I7E1 | MRFQVIVAAATITMIY | 499|577|581 |
| A0A024B7W1 | MKNPKKKSGGFRIV | 1552|1576|1636|2580|2665|2701|2737 |
| A0A024RXP8 | MYRKLAVISAFL | 228|233 |
A threshold can be selected for sequence identity between the query and the blast database, such that Squidly will be used under a certain threshold. This is because BLAST tends to outperform all currently available ML models at high sequence identities
Running a single Squidly model (non ensembled)
squidly run example.fasta esm2_t36_3B_UR50D output_folder/ --single-model --cr-model-as squidly/models/3B/CataloDB_esm2_t36_3B_UR50D_CR_1.pt --lstm-model-as squidly/models/3B/CataloDB_esm2_t36_3B_UR50D_LSTM_1.pth
Squidly args
squidly --help
Usage: squidly [OPTIONS] COMMAND [ARGS]...
╭─ Options ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╮
│ --install-completion Install completion for the current shell. │
│ --show-completion Show completion for the current shell, to copy it or customize the installation. │
│ --help Show this message and exit. │
╰──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╯
╭─ Commands ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╮
│ install Install the models for the package. │
│ run Find catalytic residues using Squidly and BLAST. │
╰──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╯
squidly run --help
Usage: squidly run [OPTIONS] FASTA_FILE ESM2_MODEL [OUTPUT_FOLDER] [RUN_NAME]
Find catalytic residues using Squidly and BLAST.
╭─ Arguments ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╮
│ * fasta_file TEXT Full path to query fasta (note have simple IDs otherwise we'll remove all funky characters.) [required] │
│ * esm2_model TEXT Name of the esm2_model, esm2_t36_3B_UR50D or esm2_t48_15B_UR50D [required] │
│ output_folder [OUTPUT_FOLDER] Where to store results (full path!) [default: Current Directory] │
│ run_name [RUN_NAME] Name of the run [default: squidly] │
╰────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╯
╭─ Options ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╮
│ --single-model --no-single-model Whether or not to use single model instead of the ensemble. We recommend the ensemble. It is faster than the single model version. [default: no-single-model] │
│ --model-folder TEXT Full path to the model folder. │
│ --database TEXT Full path to database csv (if you want to do the ensemble), needs 3 columns: 'Entry', 'Sequence', 'Residue' where residue is a | separated list of residues. See default DB provided by Squidly. [default: None] │
│ --cr-model-as TEXT Contrastive learning model for the catalytic residue prediction when not using the ensemble. Ensure it matches the esm model. │
│ --lstm-model-as TEXT LSTM model for the catalytic residue prediction when not using the ensemble. Ensure it matches the esm model. │
│ --as-threshold FLOAT When using the single squidly models, you must specify a prediction threshold. We found >0.9 to work best in practice, depending on the model. [default: 0.95] │
│ --blast-threshold FLOAT Sequence identity with which to use Squidly over BLAST defualt 0.3 (meaning for seqs with < 0.3 identity in the DB use Squidly). [default: 0.3] │
│ --chunk INTEGER Max chunk size for the dataset. This is useful for when running Squidly on >50000 sequences as memory is storing intermediate results during inference. [default: 0] │
│ --mean-prob FLOAT Mean probability threshold used in the ensemble. [default: 0.6] │
│ --mean-var FLOAT Mean variance cutoff used in the ensemble. [default: 0.225] │
│ --filter-blast --no-filter-blast Only run on the ones that didn't have a BLAST residue. [default: filter-blast] │
│ --help Show this message and exit. │
╰────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────╯
Threshold selection:
The below figure showcases Squidly's performance on the CataloDB benchmark at varying thresholds for probability and variance in the ensemble model.
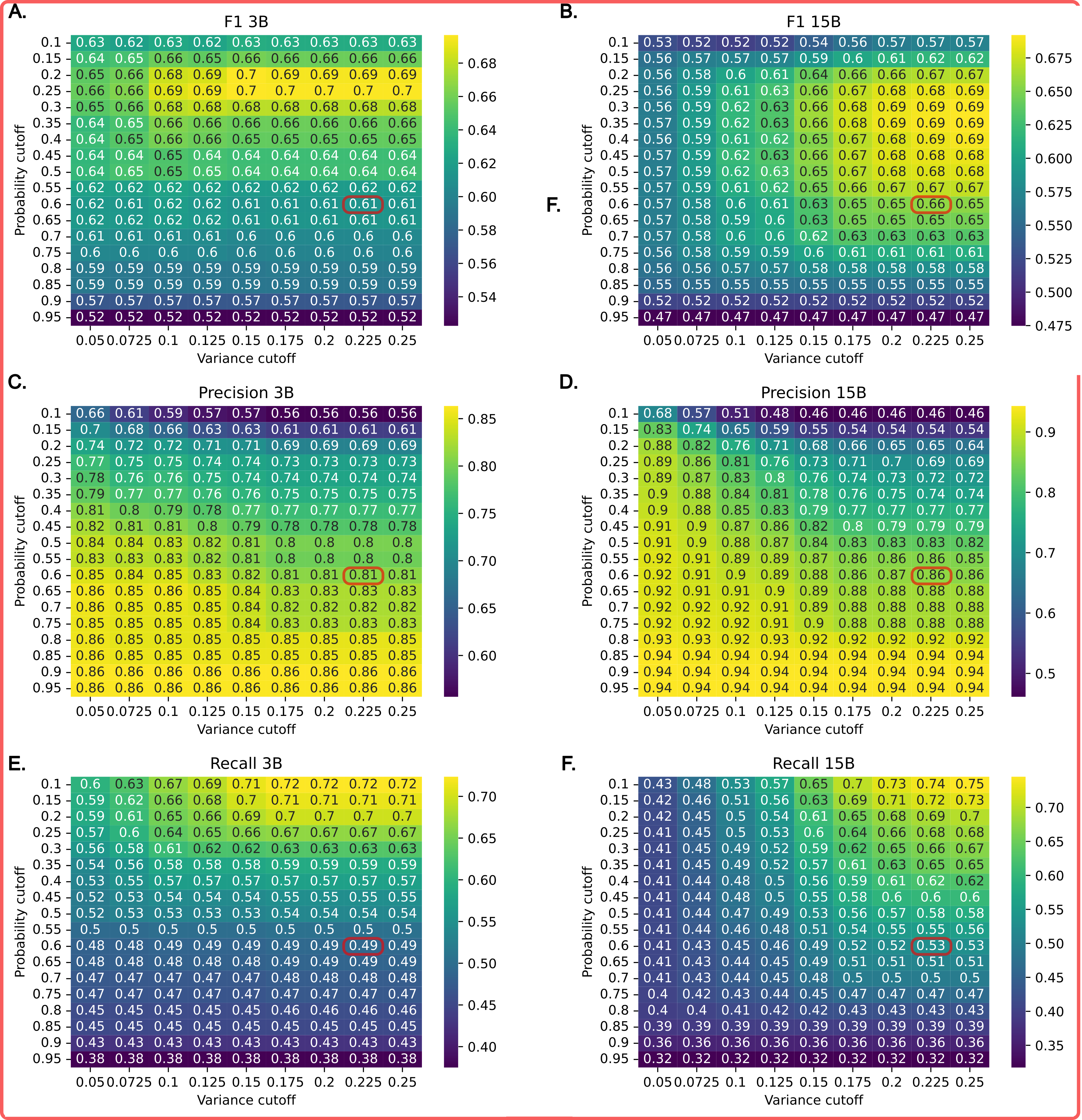
Optimal thresholds for specific uses may vary. Lower thresholds have been found to work in practice when preicting certain EC numbers... figure coming soon.
Data Availability
All datasets used in the paper are available here https://zenodo.org/records/15541320.
Reproducing Squidly
We developed reproduction scripts for each benchmark training/test scenario.
- AEGAN and Common Benchmarks: Trained on Uni14230 (AEGAN), and tested on Uni3175 (AEGAN), HA_superfamily, NN, PC, and EF datasets.
- CataloDB: Trained on a curated training and test set with structural/sequence ID filtering to less than 30% identity.
The corresponding scripts can be found in the reproduction_run directory.
Before running them, download the datasets.zip file from zenodo and place them and unzip it in the base directory of Squidly.
Datasets: https://zenodo.org/records/15541320
Model weights: https://huggingface.co/WillRieger/Squidly
python reproduction_scripts/reproduce_squidly_CataloDB.py --scheme 2 --sample_limit 16000 --esm2_model esm2_t36_3B_UR50D --reruns 1
You must choose the pair scheme for the Squidly models:
Scheme 2 and 3 had the sample limit parameter set to 16000, and scheme 1 at 4000000.
You must also correctly specify the ESM2 model used. You can either use esm2_t36_3B_UR50D or esm2_t48_15B_UR50D. The scripts will automatically download these if specified like so. You may also instead provide your own path to the models if you have them downloaded somewhere.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file squidly-0.0.7.tar.gz.
File metadata
- Download URL: squidly-0.0.7.tar.gz
- Upload date:
- Size: 22.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
472188ec41f36db0bd6b36ca790af018d819d78c7c8de68f232a9e0744f7db4d
|
|
| MD5 |
6fb934d560fb37bd4589aa9284c5e664
|
|
| BLAKE2b-256 |
e3b6814a1b8b4b21c03077b4065d5385679b34c7df3b12c737d8e720085f2ee6
|
File details
Details for the file squidly-0.0.7-py3-none-any.whl.
File metadata
- Download URL: squidly-0.0.7-py3-none-any.whl
- Upload date:
- Size: 24.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e72ad06b9110697f181edd1fcedf45a144e3f8f1e5f6112bf355c6a5334418a5
|
|
| MD5 |
c94baaa991dab2d7d7b872ce635a4ac1
|
|
| BLAKE2b-256 |
d3340ff44d62f0de67126e0ef15fe5eedaf69a8e82185c61146c1d9957a88fb2
|













