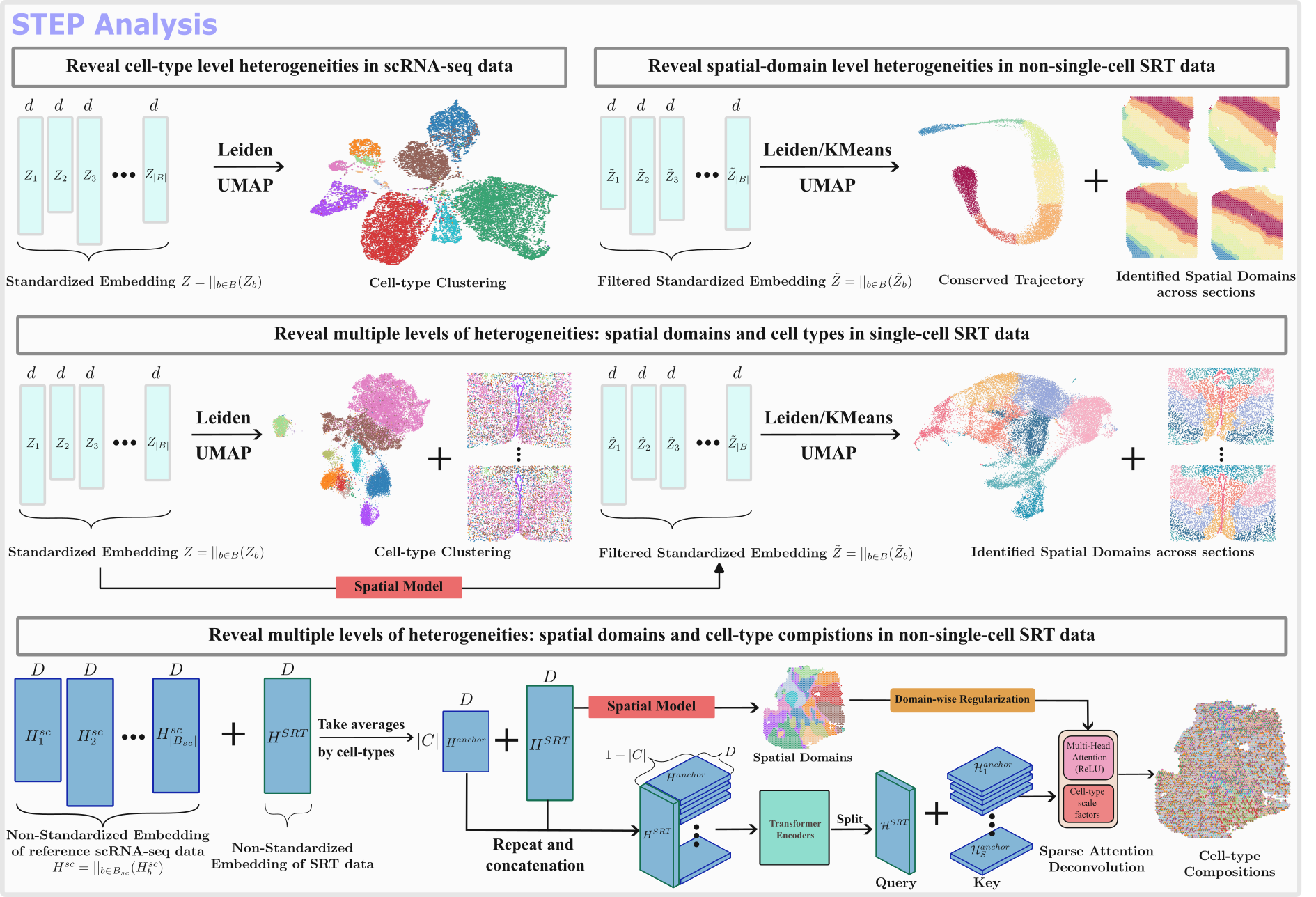
STEP, an acronym for Spatial Transcriptomics Embedding Procedure, is a deep learning-based tool for the analysis of single-cell RNA (scRNA-seq) and spatially resolved transcriptomics (SRT) data. STEP introduces a unified approach to process and analyze multiple samples of scRNA-seq data as well as align several sections of SRT data, disregarding location relationships. Furthermore, STEP conducts integrative analysis across different modalities like scRNA-seq and SRT.
Project description
STEP: Spatial Transcriptomics Embedding Procedure
Introduction
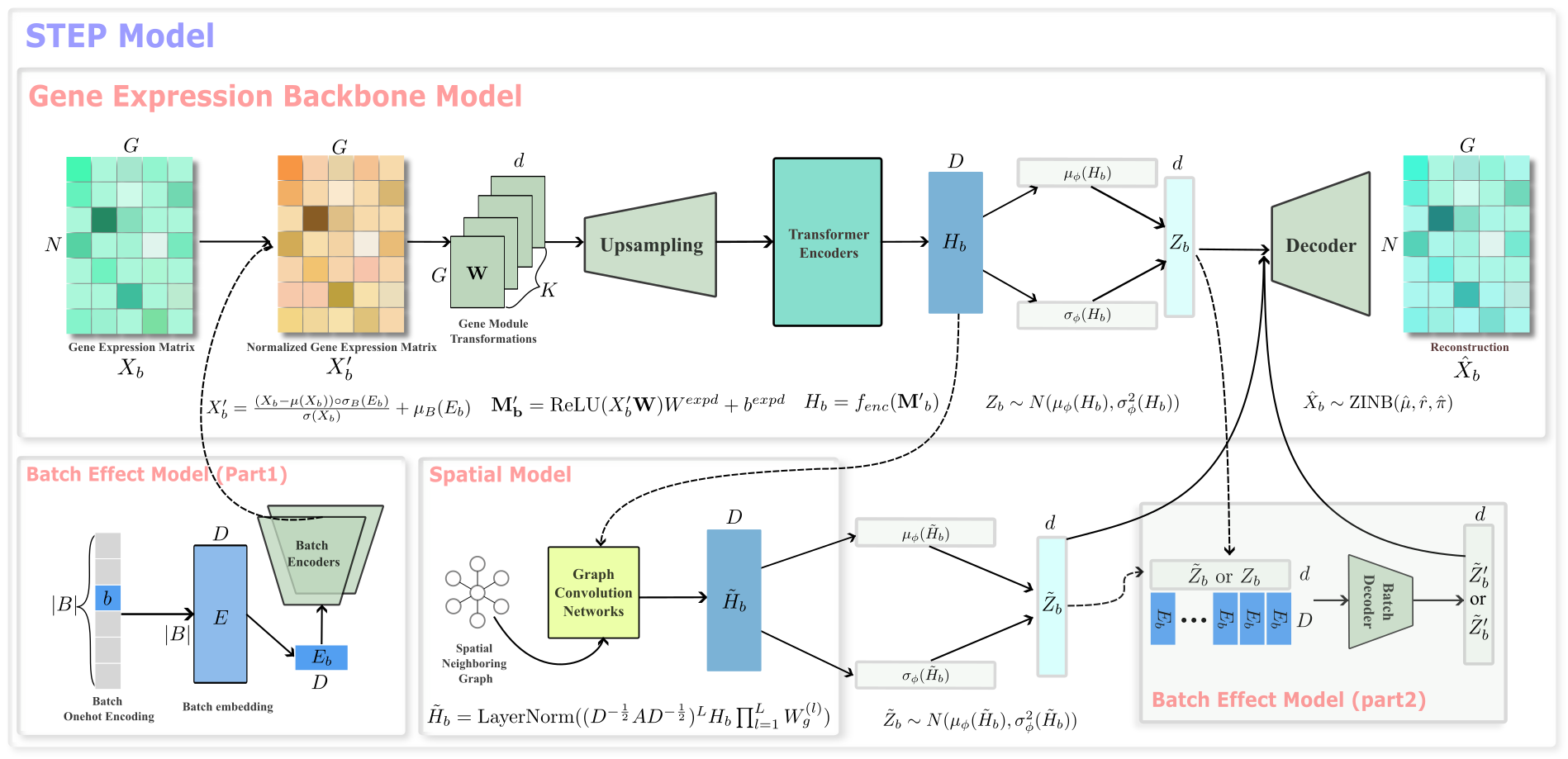
STEP, an acronym for Spatial Transcriptomics Embedding Procedure, is a foundation deep learning/AI architecture for the analysis of spatially resolved transcriptomics (SRT) data, and is also compatible with scRNA-seq data. STEP roots on the precise captures of three major varitions occured in the SRT (and scRNA-seq) data: Transcriptional Variations, Batch Variations and Spatial Variations with the correponding modular designs: Backbone model: a Transformer based model togther with gene module seqeunce mapping; Batch-effect model: A pair of inverse transformations utilizing the batch-embedding conception for the decoupled batch-effect elimination; Spatial model: a GCN-based spatial filter/smoother working on the extracted embedding from the Backbone model, different from the usage of GCN in other methods as a feature extractor. Thus, with the proper combinations of these models, STEP introduces a unified approach to systematically process and analyze single or multiple samples of SRT data, disregarding location relationships between sections (meaning both contiguous and non-contiguous sections), to reveal multi-scale bilogical heterogeneities (cell types and spatial domains) in multi-resolution SRT data. Furthermore, STEP can also conduct integrative analysis on scRNA-seq and SRT data.
Key Features
- Integration of multiple scRNA-seq and single-cell resolution SRT samples to reveal cell-type level heterogeneities.
- Alignment of various SRT data sections contiguous or non-contiguous to identify spatial domains across sections.
- Scalable to different data resolutions, i.e., wild range of technologies and platforms of SRT data, including Visium HD, Visum, MERFISH, STARmap, Stereo-seq, ST, etc.
- Scalable to large datasets with a high number of cells and spatial locations.
- Performance of integrative analysis across modalities (scRNA-seq and SRT) and cell-type deconvolution for the non-single-cell resolution SRT data.
Other Capabilities
- Capability to produce not only the batch-corrected embeddings but also batch-corrected gene expression profiles for scRNA-seq data.
- Capability to perform spatial mapping of reference scRNA-seq data points to the spatial locations of SRT data based on the learned co-embeddings and kNN.
- Comprehensive
adataprocessing by specifically desingedBaseDatasetclass and its view versionMaskedDatasetclass for the SRT data, which can be easily integrated with the PyTorchDataLoaderclass for training and validation. - Low computational cost and high efficiency in processing large-scale SRT data: 4-8 GB GPU memory is sufficient for processing a dataset with 100,000 spatial locations with 2000+ sample-size in fast mode, and consumes less than 6 minutes for training in 2000 iterations (tested on NVIDIA RTX 3090).
System Requiremtns
- Software Requirements: Python 3.10+, CUDA 11.6+, PyTorch 1.13.1+, and dgl 1.1.3+.
- Hardware Requirements: NVIDIA GPU with CUDA support (recommended), 8GB+ GPU memory. As possible as high RAM and CPU cores for storing and processing large-scale data (this is not required by STEP, but by the data itself).
Installation
Quick Installation (Recommended for Python 3.10, Linux, CUDA 11.7)
pip install step-kit[cu117]
This command installs STEP along with compatible versions of PyTorch and DGL.
Basic Installation
pip install step-kit
⚠️ Critical Note:
- This basic installation does not include PyTorch and DGL.
- STEP will not function without PyTorch and DGL properly installed.
- You must install these dependencies separately before using STEP.
Manual Dependency Installation
If you need to install specific versions of PyTorch and DGL:
-
Install DGL (example for latest DGL compatible with PyTorch 2.3.x and CUDA 11.8):
pip install dgl -f https://data.dgl.ai/wheels/torch-2.3/cu118/repo.html
-
Install PyTorch (example for PyTorch 2.3.1 with CUDA 11.8):
pip install torch==2.3.1+cu118 -f https://download.pytorch.org/whl/cu118/torch_stable.html
For more information on installing PyTorch and DGL, visit:
- PyTorch: https://pytorch.org/
- DGL: https://www.dgl.ai/
Remember to choose versions compatible with your system configuration and CUDA version (if applicable).
Contribution
We welcome contributions! Please see CONTRIBUTING.md for more details!
License
step is licensed under LICENSE
Contact
If you have any questions, please feel free to contact us at here, or feel free to open an issue on this repository.
Citation
The preprint of STEP is available at bioRxiv. If you use STEP in your research, please cite:
@article{Li2024.04.15.589470,
title = {{{STEP}}: {{Spatial}} Transcriptomics Embedding Procedure for Multi-Scale Biological Heterogeneities Revelation in Multiple Samples},
author = {Li, Lounan and Li, Zhong and Li, Yuanyuan and Yin, Xiao-ming and Xu, Xiaojiang},
year = {2024},
journal = {bioRxiv : the preprint server for biology},
eprint = {https://www.biorxiv.org/content/early/2024/04/20/2024.04.15.589470.full.pdf},
publisher = {Cold Spring Harbor Laboratory},
doi = {10.1101/2024.04.15.589470},
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file step_kit-0.3.tar.gz.
File metadata
- Download URL: step_kit-0.3.tar.gz
- Upload date:
- Size: 92.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.3 CPython/3.12.5 Linux/6.6.51
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
785ac90df2cefbf470976d9efe632406a76e3eb847005d758f7e3f95ffba997d
|
|
| MD5 |
7c433cda22efd14fb895bb9fea66aaab
|
|
| BLAKE2b-256 |
f35185d375ecf2c8491f9f49357edfc6f93658fbc445a6506eb73076fb86a621
|
File details
Details for the file step_kit-0.3-py3-none-any.whl.
File metadata
- Download URL: step_kit-0.3-py3-none-any.whl
- Upload date:
- Size: 106.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.3 CPython/3.12.5 Linux/6.6.51
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
880fe277f134a45184b31009ec4d1f9096c32409cbec95a3d32756fc4cc1a68a
|
|
| MD5 |
07a6ccb8e30df5e4a4003d6429545868
|
|
| BLAKE2b-256 |
788fbeb589fc0f645086d1f36173bcebc3bdb67e1115fbd94c06764c67cf3dc2
|