Fast pairwise cosine distance calculation and numba accelerated evolutionary matrix subset extraction
Project description

/sʌb.pɛɹ/
SubPair 
"All you need is love and evolutionary matrix subset extraction." - J. Lennon
Pairwise cosine distance is great to easily compare many vectors. However, you can end up with a very sizeable distance matrix. What if you would like to find a small subset of that matrix? Let's search it by evolution.
Given N elements and their (N,N) pairwise distance matrix we would like to get the subset of S elements such that the sum of elements in the corresponding (S,S) submatrix is minimal. See example below.
[0 1 2 3 4] indeces
i j k
│ │ │ i j k = [1, 2, 4]
0 1 6 4 1
i──1 0 3 1 7 i 0 3 7
j──6 3 0 2 3 --> j 3 0 3 --> 7 + 3 + 3 = 13 👎
4 1 2 0 1 k 7 3 0
k──1 7 3 1 0
i j k
│ │ │ i j k = [2, 3, 4]
0 1 6 4 1
1 0 3 1 7 i 0 2 3
i──6 3 0 2 3 --> j 2 0 1 --> 2 + 1 + 3 = 6 👍
j──4 1 2 0 1 k 3 1 0
k──1 7 3 1 0
All the possible subsets are ${N}\choose{S}$ and for N = 1024, S = 20 (like in the tests) we would have to check ${1024}\choose{20}$ $= 5.479 \times 10^{41}$ of them.
A few too many. Instead we are going to use an evolutionary approach to search for it.
Example usage
The usage is quite straight forward since there are only a couple of functions exported pairwise_cosine and extract.
>>> import matplotlib.pyplot as plt
>>> from subpair import pairwise_cosine
>>>
>>> X = np.random.rand(N, K).astype(np.float32)
>>> distances = pairwise_cosine(X) # (N,N)
>>> ...
>>> best, stats = extract(distances, P=200, S=S, K=50, M=3, O=2, its=3_000)
100%|█████████████████████████████████| 3000/3000 [00:03<00:00, 817.42it/s]
>>> plt.plot(stats["fits"]); plt.show()
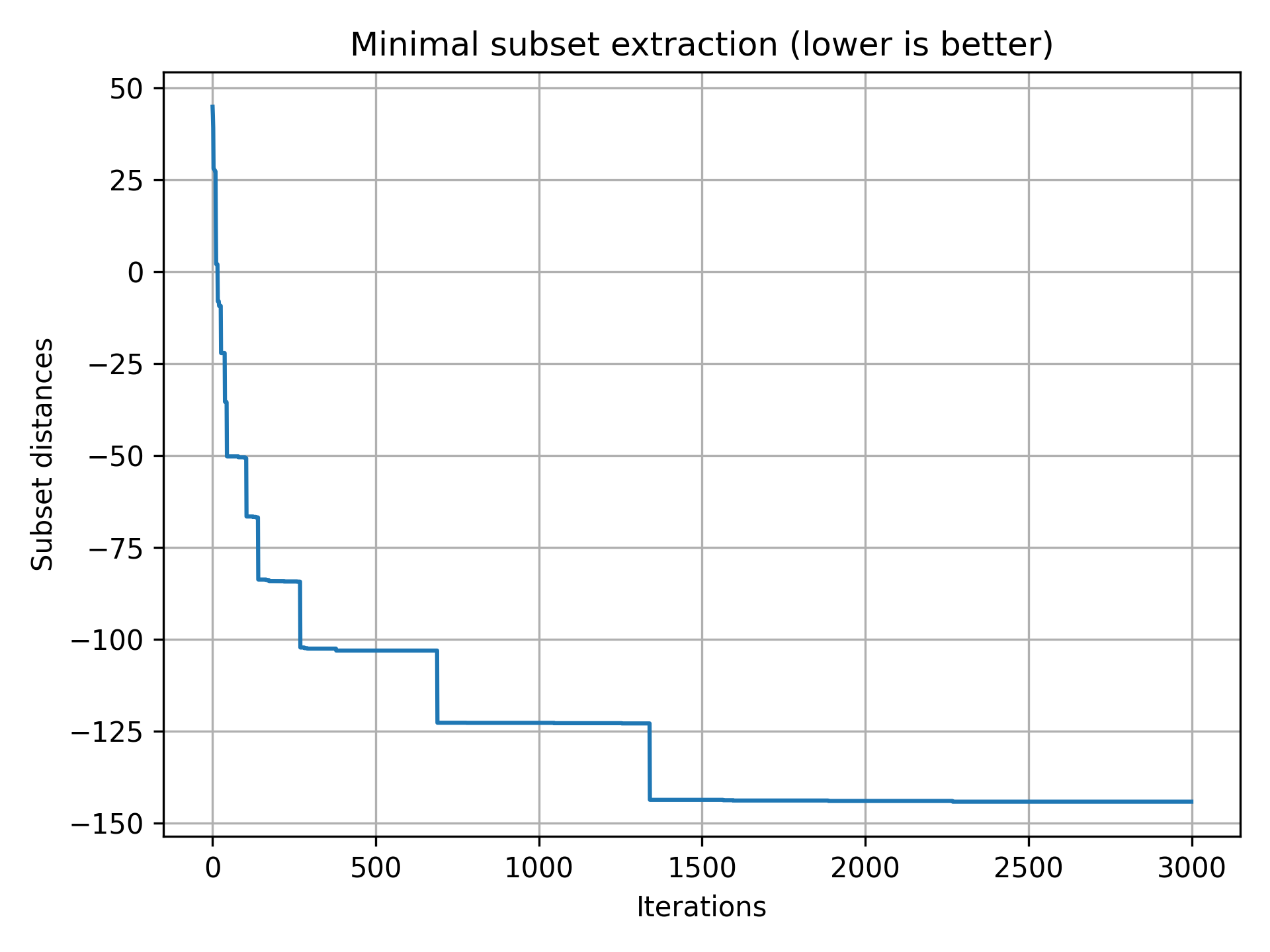
(We have sprinkled a few negative numbers to see if the algorithm can find them)
Where the options of extract are parameters for the evolutionary algorithm:distances (int, int) : N vectors of length L
P (int) : population size
S (int) : desired subset size <- determines size of output
K (int) : number of parents (P-K children)
M (int) : number of mutations
O (int) : fraction of crossovers e.g. O=2 -> 1/2, O=10 -> 1/10, (bigger=faster)
Note
This repo contains both numpy and numba/CUDA versions of the pairwise cosine distance matrix calculation. But numpy is already blazingly fast so the cuda version is provided mostly for inspiration. Our numpy version is very similar to sklearn's metrics.pairwise.cosine_distances but slightly faster. Sklearn's one has some extra nicities that our simplified version does not have.
> python flops.py # On Macbook pro M1 Max
N=513 K=2304 GOPs=1
sklearn: 0.01s - 109.4 GFLOPS
numpy: 0.00s - 162.4 GFLOPS
N=1027 K=2304 GOPs=2
sklearn: 0.02s - 135.9 GFLOPS
numpy: 0.01s - 192.4 GFLOPS
N=2055 K=2304 GOPs=10
sklearn: 0.07s - 142.9 GFLOPS
numpy: 0.06s - 166.0 GFLOPS
N=4111 K=2304 GOPs=39
sklearn: 0.20s - 195.8 GFLOPS
numpy: 0.16s - 248.6 GFLOPS
N=8223 K=2304 GOPs=156
sklearn: 0.61s - 255.3 GFLOPS
numpy: 0.54s - 289.5 GFLOPS
N=16447 K=2304 GOPs=623
sklearn: 2.11s - 295.4 GFLOPS
numpy: 1.79s - 347.9 GFLOPS
Todo
- Add type info to minimize.py to allow for AOT compilation.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file subpair-0.1.1.tar.gz.
File metadata
- Download URL: subpair-0.1.1.tar.gz
- Upload date:
- Size: 7.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.10.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e3b51a8b9423358bfac561301b96e4ef167c1c74bd65cb7dfd0a69bb737a8600
|
|
| MD5 |
f5e065442b2690dc08e9a3c6ef269172
|
|
| BLAKE2b-256 |
09c6fbb4394423f7cadddc9b78d5ef6804bd071f3635a7d5af0469c23f128a29
|
File details
Details for the file subpair-0.1.1-py3-none-any.whl.
File metadata
- Download URL: subpair-0.1.1-py3-none-any.whl
- Upload date:
- Size: 7.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.10.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a55ccb182fa3841589d9eed3edd3a746c42f107957898e55f84bf51abdf9f8ea
|
|
| MD5 |
2794e4965a311286f3ea3f446d96aa23
|
|
| BLAKE2b-256 |
fbaedd312e3233ac0137115fe0cf0372da35b43eac8db9fd60df5f1167e66aa2
|










