A structural variant caller for genome-genome alignments.
Project description

SVIM-asm (pronounced SWIM-assem) is a structural variant caller for haploid or diploid genome-genome alignments. It analyzes a given sorted BAM file (preferably from minimap2) and detects five different variant classes between the query assembly and the reference: deletions, insertions, tandem and interspersed duplications and inversions.
Note! To analyze raw long sequencing reads please use our other method SVIM.
Background
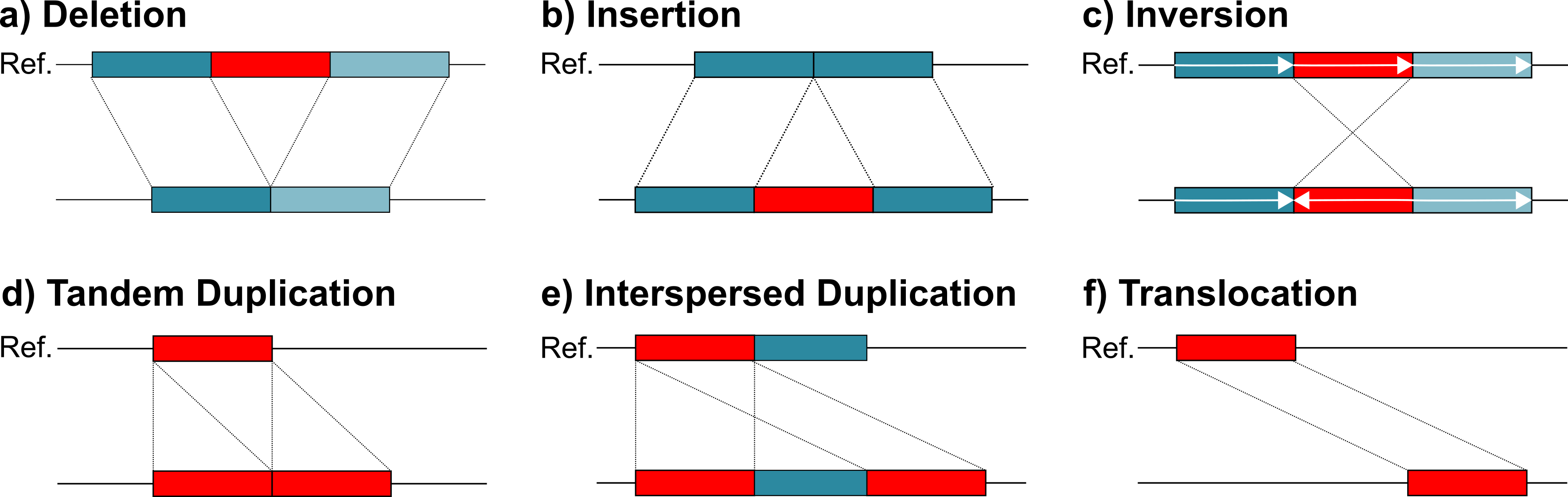
Structural variants (SVs) are typically defined as genomic variants larger than 50bps (e.g. deletions, duplications, inversions). Studies have shown that they affect more bases in an average genome than all SNPs or all small Indels together. Consequently, they have a large impact on genes and regulatory regions. This is reflected in the large number of genetic disorders and other disease that are associated to SVs.
Nowadays, SVs are usually detected using data from second-generation sequencing (Illumina) or third-generation sequencing (PacBio and Oxford Nanopore). Typically, the reads from a sequencing experiment are first aligned to a reference genome before the alignments are analyzed for characteristic signatures of SVs. Recently, substantial advances in sequencing technology and software development have made the de novo assembly of large mammalian genomes more efficient than ever. Accurate assemblies of the human genome can now be generated in a few days and at a fraction of its former cost. [Shafin et al.]
Similarly to raw sequencing reads, the genome assemblies can be aligned to another genome to uncover genomic rearrangements and structural variants. Our tool, SVIM-asm, detects structural variants between different assemblies or reference genomes from given genome-genome alignments. It is fast (<5 min for a human genome-genome alignment), easy to use and detects all major variant types.
Installation
SVIM-asm can be installed most easily using conda:
#Recommended: Install via conda into a new environment
conda create -n svimasm_env --channel bioconda svim-asm
#Alternatively: Install via conda into existing (active) environment
conda install --channel bioconda svim-asmAlternatively, SVIM-asm can be installed using pip:
#Install from github (requires Python 3)
git clone https://github.com/eldariont/svim-asm.git
cd svim-asm
pip install .Changelog
v1.0.1: reduce memory consumption substantially
v1.0.0: add genotyping of translocation breakpoints (BNDs), bugfixes
v0.1.1: improve breakend detection, add FORMAT:CN tag for tandem duplications, add two new command-line options to output duplications as INS records in VCF, bugfixes
v0.1.0: initial beta release
Execution
SVIM-asm analyzes alignments between a query assembly and a reference assembly in SAM/BAM format. We recommend to produce the alignments using minimap2. See this example for a haploid query assembly:
minimap2 --paf-no-hit -a -x asm5 --cs -r2k -t <num_threads> <reference.fa> <assembly.fasta> > <alignments.sam>
samtools sort -m4G -@4 -o <alignments.sorted.bam> <alignments.sam>
samtools index <alignments.sorted.bam>
svim-asm haploid <working_dir> <alignments.sorted.bam> <reference.fa>To analyze a diploid assembly consisting of two haplotypes, you need to align both to the reference assembly:
minimap2 --paf-no-hit -a -x asm5 --cs -r2k -t <num_threads> <reference.fa> <haplotype1.fasta> > <alignments_hap1.sam>
minimap2 --paf-no-hit -a -x asm5 --cs -r2k -t <num_threads> <reference.fa> <haplotype2.fasta> > <alignments_hap2.sam>
samtools sort -m4G -@4 -o <alignments_hap1.sorted.bam> <alignments_hap1.sam>
samtools sort -m4G -@4 -o <alignments_hap2.sorted.bam> <alignments_hap2.sam>
samtools index <alignments_hap1.sorted.bam
samtools index <alignments_hap2.sorted.bam
svim-asm diploid <working_dir> <alignments_hap1.sorted.bam> <alignments_hap2.sorted.bam> <reference.fa>Output
SVIM-asm creates all output files in the given working directory. The following files are produced:
variants.vcf contains the detected SVs in VCF format (see http://samtools.github.io/hts-specs/VCFv4.2.pdf)
sv-lengths.png contains a histogram of SV sizes
SVIM_<day>_<time>.log contains the same logging output as the command line
Contact
If you experience problems or have suggestions please create an issue or a pull request or contact heller_d@molgen.mpg.de.
Citation
SVIM-asm is a fork of our long-read caller SVIM. Feel free to read and cite our paper in Bioinformatics: https://doi.org/10.1093/bioinformatics/btz041
License
The project is licensed under the GNU General Public License.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file svim-asm-1.0.1.tar.gz.
File metadata
- Download URL: svim-asm-1.0.1.tar.gz
- Upload date:
- Size: 234.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.13.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.4.0 requests-toolbelt/0.9.1 tqdm/4.32.2 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7aaf6d588ac9b1115c6d2b9a4a28669e5aa4799d4e61c4a0e60de6d226bf2010
|
|
| MD5 |
d205af720b38382e5ad8e9fbde052621
|
|
| BLAKE2b-256 |
35bb0a1f71ca0eae62b55237814ca4036964b9ad354e720dc693f4e8c90de415
|
File details
Details for the file svim_asm-1.0.1-py3-none-any.whl.
File metadata
- Download URL: svim_asm-1.0.1-py3-none-any.whl
- Upload date:
- Size: 53.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/1.13.0 pkginfo/1.5.0.1 requests/2.22.0 setuptools/41.4.0 requests-toolbelt/0.9.1 tqdm/4.32.2 CPython/3.7.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a2f9ff82fd15695f61f42390e067a026cf0efe1af7a26de5c661082e9b3da3b8
|
|
| MD5 |
f20a33067e4739a189ea011b0f7987d4
|
|
| BLAKE2b-256 |
bd4cefe1a4df89ef82c98330f86b259541ee0402aa3646eeaf9495b875734dc6
|











